* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Spectrophotometry
Survey
Document related concepts
Protein mass spectrometry wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Protein structure prediction wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
List of types of proteins wikipedia , lookup
Transcript
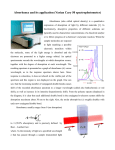
Spectrophotometry August 2011 SLCC/UVU STEP grant workshop Spectrophotometry • How much light of a particular wavelength is absorbed by a sample? Figure © David P. Goldenberg, University of Utah, 2003 3 Absorbance vs. concentration • Absorbance values above 2 tend to be unreliable – best values: 0.1 < A < 1.0 Figure © David P. Goldenberg, University of Utah, 2003 The Beer-Lambert Law: A=C·l· • • • • A = absorbance C = concentration l = cuvette pathlength = extinction coefficient – specific for a particular wavelength – specific for a particular compound The electromagnetic spectrum http://www.rationalizing.us/blog/files/wavelength_figure.jpg Different molecules have different absorption spectra http://dx.doi.org/10.1590/S0103-50532006000300006 UV and visible absorbance often arises from… • Coordinated metal ions – Chlorophyll – Heme • Systems of conjugated double bonds UV absorbance by aromatic amino acids …and nucleic acids! Spectral overlap between proteins and nucleic acids Figure © David P. Goldenberg, University of Utah, 2003 Notes about absorbance • Absorbance is unitless • Absorbance is sometimes also referred to as optical density (OD) – often, the wavelength of light is denoted, e.g. OD600 = absorbance at 600 nm = A600 The Beer-Lambert Law: A=C·l· • For double stranded DNA, – has been found to be 20 L/(g cm) – C = 299 792 458 m / s or 3.0 x 108 m/s – l = A rule of thumb in molecular biology, using standard 1 cm cuvettes: 1 A260 unit for dsDNA = 50 ug/mL • Use Beer’s Law to prove it Practical points • The cuvette must be transparent to light of the wavelength of interest – Glass or standard plastic ok for visible light (≥ 350 nm) – Specialized cuvettes (quartz or TrUView) required for UV light (~200-350 nm) • Absorbances are measured relative to that for a “blank” solution that contains everything except the compound of interest – If your compound is in water, blank with water – If your compound is in a buffer, blank with the buffer • Ensure cuvettes are placed in the proper orientation Two types of specs in our lab • SmartSpec Plus • NanoDrop 1000 SmartSpec Plus • uses cuvettes – must be transparent to wavelength of interest • built in programs estimate concentrations of DNA, protein, cells, etc. • relatively large sample volume (50-500 µL, depending on type of cuvette) NanoDrop • Measures absorbance with sample volume as low as 1 µL Absorbance ratios • Nucleic acid absorbs strongly at 260 nm • Protein absorbs strongly at 280 nm • Ratio of 260/280 allows comparison of amount of nucleic acid vs. protein – ratio of ~1.8 is generally accepted as“pure” for DNA – ratio of ~2.0 is generally accepted as “pure” for RNA – if the ratio is appreciably lower in either case, it may indicate the presence of protein, phenol or other contaminants that absorb strongly at or near 280 nm. Absorbance ratios • Nucleic acid absorbs strongly at 260 nm • Carbohydrates, some reagents used in purification absorb near 230 nm • Expected 260/230 values are commonly in the range of 2.0-2.2. – If the ratio is appreciably lower than expected, it may indicate the presence of contaminants which absorb at 230 nm.