* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Nitrogen Assimilation
Survey
Document related concepts
Transcript
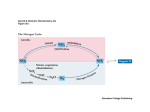
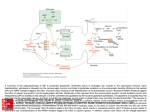
Nitrogen Assimilation • How is NH3 incorporated into organic molecules? • Glutamate Dehydrogenase vs Glutamate Synthase • Properties of Glutamine Synthetase • Regulation of Glutamine Synthetase • Glutamine as a major nitrogen donor 3 Gateways to Biological Molecules CO2 ATP Carbamoyl phosphate Arginine Urea NH3 Aspartate Asparagine None The only inorganic nitrogen source for mammals is NH3 a-Ketoglutarate Glutamate Other amino acids Of the 3, the most versatile is glutamine Glutamate Glutamine Purine nucleotides, Cytidine nucleotides Amino sugars, Tryptophan, Histidine Pyrimidine nucleotides Glutamate Dehydrogenase a-Kg + NH3 + NAD(P)H + 2H+ Glutamate + NAD(P)+ + H2O Bacteria and Plants Animals Make glutamate, assimilate NH3 Glutamate is major solute in the bacteria Provide NH3 to urea cycle, provide a-Kg to Krebs High Km for NH3 limits forward Mitochondrial location (-) by ATP, GTP (+) by ADP, GDP Glutamate Synthase (Bacteria Only) a-Kg + glutamine + NADPH + H+ 2 glutamates + NADP+ Glutamine is the nitrogen donor Reaction is a reductive amination Glutamine Synthetase is a Primary Regulatory Point in Nitrogen Metabolism Properties of Bacterial Enzyme 12 identical 50,000 mwt subunits Combined Mwt of 600,000 6 6 Hexagonal stacked rings SIDE VIEW Each subunits subject to allosteric regulation 8 allosteric sites on each subunit One covalent site (Tyr 397) Regulation is cumulative TOP VIEW Glutamine Synthetase Glutamate + NH4+ + ATP Glutamine + ADP + Pi (Biosynthesis - anabolic) a-Ketoglutarate (Degradation – catabolic) Take Home: Shutting down the enzyme favors using glutamate as an energy substrate. Not shutting down the enzyme keeps the cell in a biosynthetic mode. Covalent Modification by Adenylylation Adenylylated tyrosine residue Enzyme O Tyr H COO CH2 CH2 + C NH3 COO + NH3 + ATP O P O O H CH2 O Adenine OH OH O C NH2 CH2 CH2 + C NH3 + ADP + Pi COO Allosteric Effectors Each inhibits Glutamine Synthetase (Favors boosting cell energy or shutting down a pathway requiring glutamine ) AMP (Low energy state exists, oxidize a-Kg) CTP (End product of pyrimidine synthesis) Histidine Glycine Tryptophan Alanine Amino acids that are allosteric effectors Glucosamine (Glutamine sufficient for amino sugar synthesis) Carbamoyl-PO4 (Glutamine sufficient for pyrimidine synthesis) Covalent Regulation Adenylylation of Tyrosine 397 on EACH of the 12 subunits Adenyl = group attached Adenylylation = group attached and process For example: An acyl group attached via an acylation reaction is ACYLACYLATION OR ACYLYLATION (PRONOUNCED ACIL-LIL-ATION) Covalent Regulation of Glutamine Synthetase 2 Transferases Each puts on and takes off groups AT (Adenylyltransferase - adenylylates GS) UT (Uridylyltransferase - uridylylates PII) One Regulatory Protein PII Two States AT-PII Adenylates AT-PII-UMP Deadenylylates Rules of Engagement • Transferases catalyze adenylylation (uridylylation) and deadenylylation (deuridylylation reactions • Adenylylation shuts GS down cumulatively Deadenylylation turns GS back on cumulatively • AT requires PII to adenylylate • AT requires PII-UMP to deadenylylate Active GS* ATP AT PII AMP AT PII UMP AMP GS Less Active UTP PPi PII UMP AT = Useless PII UMP UT UR AT PII = Adenylylates AT PII UMP = Deadenylylates Uridylate removing Enzyme UT = Uridylylates (2 activities on same enzyme) UR = Deuridylylates Activates GS ATP (-) glutamine (+) a-Kg UTP UR UT PII Inactivates GS PII Deadenylylates PPi PII UMP UT UR PII UMP UMP Leads to adenylylation Responsive to cell’s nitrogen requirements