* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Information Extraction in Biology
Biochemistry wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Gene regulatory network wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Paracrine signalling wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Proteolysis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Signal transduction wikipedia , lookup
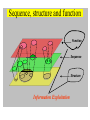
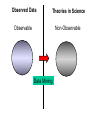
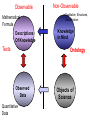
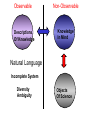
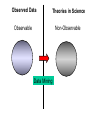
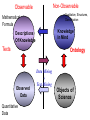
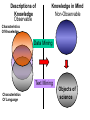
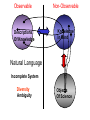
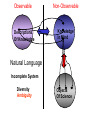
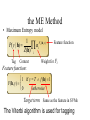
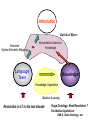
NLP for Biomedicine - Ontology building and Text Mining Junichi Tsujii GENIA Project (http://www-tsujii.is.s.u-tokyo.ac.jp/GENIA/) Computer Science Graduate School of Information Science and Technology University of Tokyo JAPAN My Talk 1. Background : Why NLP in Biomedicines 2. Examples of NLP in Biomedicines 3. Text Mining and NLP 4. Our current Work 4.1 Terms and NE 4.2 Resource Building 4.3 Event Recognition 5. Concluding Remarks My Talk 1. Background : Why NLP in Biomedicines 2. Examples of NLP in Biomedicines 3. Text Mining and NLP 4. Our current Work 4.1 Terms and NE 4.2 Resource Building 4.3 Event Recognition 5. Concluding Remarks Why NLP in Biomedicine ? From Biology and Medical Sciences From Natural Language Processing Why NLP in Biomedicine ? From Biology and Medical Sciences From Natural Language Processing Genome sequencing. by D. Devos Sequence, structure and function Function Sequence Structure Information Exploitation Scientists in areas such as molecular biology and biochemistry aim to discover new biological entities and their functions. Typical cases could be discoveries of the implications of new proteins and genes in an already known process, or implication of proteins with previously characterized functions in a separate process. The use of available information (published papers, etc.) is a key step for the discovery process, since in many cases weak or indirect evidences about possible relations hidden in the literature are used to substantiate working hypothesis that are experimentally explored. [C.Blaschke, A.Valencia: 2001] Scientists in areas such as molecular biology and biochemistry aim to discover new biological entities and their functions. Typical cases could be discoveries of the implications of new proteins and genes in an already known process, or implication of proteins with previously characterized functions in a separate process. The use of available information (published papers, etc.) is a key step for the discovery process, since in many cases weak or indirect evidences about possible relations hidden in the literature are used to substantiate working hypothesis that are experimentally explored. [C.Blaschke, A.Valencia: 2001] Scientists in areas such as molecular biology and biochemistry aim to discover new biological entities and their functions. Typical cases could be discoveries of the implications of new proteins and genes in an already known process, or implication of proteins with previously characterized functions in a separate process. The use of available information (published papers, etc.) is a key step for the discovery process, since in many cases weak or indirect evidences about possible relations hidden in the literature are used to substantiate working hypothesis that are experimentally explored. [C.Blaschke, A.Valencia: 2001] Why NLP in Biomedicine ? From Biology and Medical Sciences From Natural Language Processing Information Statistical Biases Grammar Syntax-Semantic Mapping Interpretation based on Knowledge Language Texts Knowledge Knowledge Acquisition Machine Learning Revolution in LT in the last decade Huge Ontology: Next Revolution ? Bio-Medical Application: UMLS, Gene Ontology, etc. My Talk 1. Background : Why NLP in Biomedicines 2. Examples of NLP in Biomedicines 3. Text Mining and NLP 4. Our current Work 4.1 Terms and NE 4.2 Resource Building 4.3 Event Recognition 5. Concluding Remarks What can we do in Biomedical domains by NLP ? Examples Protein-Protein Interaction extracted from texts by C. Blaschke Organized Knowledge through terms by C. Blaschke From Data to Understanding: Interpretation by Language Oliveros, Blaschke et al., GIW 2000 Information Extraction from Texts QA Answering Systems Characteristics of Signal Pathway (1) • Granularity of Knowledge Units Different types of entities which are interrelated with each other Cells, Sub-locations of cells Proteins, substructures of proteins, Subclasses of proteins Ions, other chemical substances Genes, RNA, DNA G-protein coupled receptor pathway model figure from TRANSPATH CSNDB (National Institute of Health Sciences) • A data- and knowledge- base for signaling pathways of human cells. – It compiles the information on biological molecules, sequences, structures, functions, and biological reactions which transfer the cellular signals. – Signaling pathways are compiled as binary relationships of biomolecules and represented by graphs drawn automatically. – CSNDB is constructed on ACEDB and inference engine CLIPS , and has a linkage to TRANSFAC. – Final goal is to make a computerized model for various biological phenomena. Example. 1 • A Standard Reaction Signal_Reaction: “EGF receptor Grb2” From_molecule “EGF receptor” To_molecule “Grb2” Tissue “liver” Effect “activation” Interaction “SH2+phosphorylated Tyr” Reference [Yamauchi_1997] Excerpted @[Takai98] Example. 3 • A Polymerization Reaction Signal_Reaction: “Ah receptor + HSP90 ” Component “Ah receptor” “HSP90” Effect “activation dissociation” Interaction “PAS domain of Ah receptor” Activity “inactivation of Ah receptor” Reference [Powell-Coffman_1998] Excerpted @[Takai98] My Talk 1. Background : Why NLP in Biomedicines 2. Examples of NLP in Biomedicines 3. Text Mining and NLP 4. Our current Work 4.1 Terms and NE 4.2 Resource Building 4.3 Event Recognition 5. Concluding Remarks Observed Data Theories in Science Observable Non-Observable Data Mining Observable Mathematical Formula Descriptions Of Knowledge Texts Non-Observable Qualitative, Structures, Classification Knowledge In Mind Ontology Observed Data Quantitative Data Objects of Science Observable Descriptions Of Knowledge Non-Observable Knowledge In Mind Natural Language Incomplete System Diversity Ambiguity Objects Of Science Observed Data Theories in Science Observable Non-Observable Data Mining Non-Observable Observable Qualitative, Structures, Classification Mathematical Formula Knowledge In Mind Descriptions Of Knowledge Texts Ontology Observed Data Quantitative Data Data Mining + Text Mining Objects of Science Descriptions of Knowledge Observable Knowledge in Mind Non-Observable Characteristics Of Knowledge Data Mining Text Mining Characteristics Of Language Objects of science Observable Descriptions Of Knowledge Non-Observable Knowledge In Mind Natural Language Incomplete System Diversity Ambiguity Objects Of Science Observable Descriptions Of Knowledge Non-Observable Knowledge In Mind Natural Language Incomplete System Diversity Ambiguity Objects Of Science My Talk 1. Background : Why NLP in Biomedicines 2. Examples of NLP in Biomedicines 3. Text Mining and NLP 4. Our current Work 4.1 Terms and NE 4.2 Resource Building 4.3 Event Recognition 5. Concluding Remarks Terms are the basic units of knowledge Classification, Features NE recognition Event Recognition Semantic Disambiguation Task difficulties in molecular-biology •Inconsistent naming conventions e.g. IL-2, IL2, Interleukin 2, Interleukin-2, Il-2 NF kappa B, NF-kappa B, (NF)-kappa B, NF-Kappa B, … •Wide-spread synonymy Many synonyms in wide usage, e.g. PKB and Akt cycline-dependent kinase inhibitor p27, p27kip1 <cdc25, cdc25a>, <p52shc, p52(Shc)> •Open, growing vocabulary for many classes Linking Problem Diversity Lexicon Static Processing Term Recognition •Cross-over of names between classes depending on context Ambiguity Context Dependent •Protein vs DNA Dynamic Processing •Frequent uses of coordination inside term formations Ambiguity • Abbreviation Extraction (Schwartz 2003) – Extracts short and long form pairs Short form Long form AA Alcoholic Anonymous American Americans Arachidonic acid arachidonic acid amino acid amino acids anaemia anemia : Experiment [Tsuruoka, et.al. 03 SIGIR] • Corpus – MEDLINE: the largest collection of abstracts in the biomedical domain • Rule learning – 83,142 abstracts – Obtained rules: 14,158 • Evaluation – 18,930 abstracts – Count the occurrences of each generated variant. Results: “NF-kappa B” Generation Probability Generated Variants Frequency 1.0 (Input) NF-kappa B 857 0.417 NF-kappaB 692 0.417 nF-kappa B 0 0.337 Nf-kappa B 0 0.275 NF kappa B 25 0.226 NF-kappa b 0 : : : Results: “antiinflammatory effect” Generation Probability Generated Variants Frequency 1.0 (input) antiinflammatory effect 7 0.462 anti-inflammatory effect 33 0.393 antiinflammatory effects 6 0.356 Antiinflammatory effect 0 0.286 antiinflammatory-effect 0 0.181 anti-inflammatory effects 23 : : : Results: “tumour necrosis factor alpha” Generation Probability Generated Variants Frequency 1.0 (Input) tumour necrosis factor alpha 15 0.492 tumor necrosis factor alpha 126 0.356 tumour necrosis factor-alpha 30 0.235 Tumour necrosis factor alpha 2 0.175 tumor necrosis factor alpha 182 0.115 Tumor necrosis factor alpha 8 : : : Task difficulties in molecular-biology •Inconsistent naming conventions e.g. IL-2, IL2, Interleukin 2, Interleukin-2, Il-2 NF kappa B, NF-kappa B, (NF)-kappa B, NF-Kappa B, … •Wide-spread synonymy Many synonyms in wide usage, e.g. PKB and Akt cycline-dependent kinase inhibitor p27, p27kip1 <cdc25, cdc25a>, <p52shc, p52(Shc)> •Open, growing vocabulary for many classes Linking Problem Diversity Lexicon Static Ptocessing Term Recognition •Cross-over of names between classes depending on context Ambiguity Context Dependent •Protein vs DNA Dynamic Processing •Frequent uses of coordination inside term formations Genia Ontology Substance +substance-+-compound-+-organic-+-nucleic_acid-+-poly_nucleotides | | | | +-nucleotide | | | | +-DNA | | | | +-RNA | | | | | | | +-amino_acid_monomer | | | | +-protein | | | +-lipid | | | +-carbohydrate | | | +-other_organic_compounds | | +-inorganic | +-atom +-amino_acid-+-peptide Genia Ontology: Source +-source-+-natural-+-organism-+-multi_cell | | | +-mono_cell | | | +-virus | | +-body_part | | +-tissue | | +-cell_type | +-artificial-+-cell_line | +-other_artificial_sources Number of Tagged Objects • Texts: 2,500 MEDLINE Abstracts – Papers on Transcription Factors in Human blood cells – 550,000 words, 20,000 sentences • Tagged objects: 147,000 – – – – – Protein: DNA: RNA: Source: Other: ~ 77,000 ~ 24,000 ~ 2,400 ~ 27,000 ~ 37,000 Distributions of Semantic Classes organism cell type tissue others cell component cell line atom artificial source inorganic compound protein other organic compound carbohydrate peptide lipid RNA nucleotides polynucleotides amino acid monomer DNA Extension of GENIA Ontology • Small classes (to be embedded in UMLS) – 5242 terms labelled with ‘other_names’ class • Events, Biological reactions 3800 • Disease 636 – – – – – Names of Diseases Treatments 61 Diagnoses 52 Pathology 3 Others 39 Classification of "other_names" 501 Event or Reaction Disease Experiment Other • Experiments 578 – Methods 493 – Materials 25 – Others 60 Sub-classification of "Disease" Sub-classification of "Experiment" • Others 228 Disease name Diagnosis Other Treatment method Pathology Method Material Other Biomedical NE Task (Collier Coling00,Kazama ACL02, Kim ISMB02) • Recognize “names” in the text – Technical terms expressing proteins, genes, cells, etc. Thus, CIITA not only activates the expression of class II genes PROTEIN DNA but recruits another B cell-specific coactivator to increase transcriptional activity of class II promoters in B cells . CELLTYPE DNA Identify and classify NE Task as Classification • To a class (tag) representing the semantic class and the position in the term – The task is reduced to a tagging task • We can use methods developed for tagging – The structure is encoded in a tag • BIO (Begin, Inside, and Other) tagging Term of class X Words: BIO tags: Term of class Y … o B-X I-X I-X (OTHER) o o B-Y o o NE Tagging Illustrated • Classify a word depending on the context Words: activity of POS tags: N P class II promoters in N Sym Ns P conversion to features context BIO tags: classifier O O B-DNA I-DNA Deterministic tagging: - Only the most probable tag at each word (SVM) The Viterbi tagging: - The most probable sequence among all (probabilistic models) The GENIA Corpus [Tateishi HLT02., Ohta PSB00, ISMB02] Annotated MEDLINE abstracts # of abstracts: # of sentences: # of tokens (words): # of named entities: # of semantic classes: 670 5,109 152,216 23,793 24 Big enough to: make SVM usage nontrivial Small enough to: make sparseness serious - 2,000-abstract version soon A gold standard for biomedical NLP tasks http://www-tsujii.is.s.u-tokyo.ac.jp/GENIA/ the ME Method • Maximum Entropy model 1 Fi (h,y ) P(y | h) i Z(h) i Tag Context Feature function Weight for Fi Feature function: 1 if y = T f (h) 1 F(h, y) otherwise 0 Target term Same as the feature in SVMs The Viterbi algorithm is used for tagging SOHMM modeling (J.KIM, et.al. ACL03) • SOHMM modeling l W arg max Pti | ci Pwi | ti t1,l ci ci i 1 A set of contextual feature values which are visible at the moment oftpredicting . i A classification function from sets of contextual feature values to context patterns grouped appropriately. – No assumption is made arbitrarily. – Instead, a context classification function is induced from a corpus. • SOHMM learning – Inducing the context classification function – Estimating parameters Experimental Results • Biological source recognition Matching method precision recall F-score hard matching 59.72 68.92 63.99 soft matching left 63.23 72.97 67.75 soft matching right 61.36 70.81 65.75 soft matching either 64.87 74.86 69.51 • Biological substance recognition Matching method precision recall F-score hard matching 73.76 66.92 70.17 soft matching left 77.64 70.67 73.99 soft matching right 75.19 68.22 71.54 soft matching either 79.07 71.98 75.36 Event Recognition Identity of events in our mind Disambiguation of different events by context Problem: Syntactic Variations ACTIVATOR activate ACTIVATEE RAF6 activates NF-kappaB. Lck is activated by autophosphorylation at Tyr 394. Anandamide induces vasodilation by activating vanilloid receptors. the activation of Rap1 by C3G the GTPase-activating protein rhoGAP the stress-activated group of MAP kinases Verbs Related to Biological Events Frequent Verbs in 100 MEDLINE Abstracts Ver b be induce bind show suggest activate factor demonstrate inhibit have reveal require regulate indicate find result play interact mediate contain C ount 255 56 50 49 42 42 36 35 26 25 21 21 21 21 21 20 19 18 17 17 Ver b C ount involve 16 identify 16 act 15 stimulate 14 provide 14 express 13 affect 13 type 12 report 12 form 12 contribute 12 study 11 observe 11 lead 11 function 11 assay 11 appear 11 occur 10 increase 10 phosphorylate 9 Ver b determine construct associate reduce prevent locate line differ trigger synergize examine block become analyze target signal remain produce present possess C ount 9 9 9 8 8 8 8 8 7 7 7 7 7 7 6 6 6 6 6 6 Ver b explain exert enhance display characterize participate localize investigate imply establish conclude compare use transform transfect test suppress support substitute share C ount 6 6 6 6 6 5 5 5 5 5 5 5 4 4 4 4 4 4 4 4 Argument Frame Extractor 133 argument structures, marked by a domain specialist in 97 sentences among the 180 sentences Extracted Uniquely Extracted with ambiguity Extractable from pp’s Parsing Not extractable Failures Memory limitation,etc 31 32 26 27 17 68% My Talk 1. Background : Why NLP in Biomedicines 2. Examples of NLP in Biomedicines 3. Text Mining and NLP 4. Our current Work 4.1 Terms and NE 4.2 Resource Building 4.3 Event Recognition 5. Concluding Remarks Information Statistical Biases Grammar Syntax-Semantic Mapping Interpretation based on Knowledge Language Texts Knowledge Knowledge Acquisition Machine Learning Revolution in LT in the last decade Huge Ontology: Next Revolution ? Bio-Medical Application: UMLS, Gene Ontology, etc. Genome sequencing. Actual demands in the real world with more homogenous user groups and more concrete criteria for evaluating results by D. Devos http://www-tsujii.is.s.u-tokyo.ac.jp/GENIA/ Resources available Medline Abstracts (4000, about 1 million words) GENIA ontology POS tags Semantic tags Structural tags Co-reference annotations with a Singaporean team Lexical resources mapped to existing ontology