* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Presentation
Restriction enzyme wikipedia , lookup
Proteolysis wikipedia , lookup
Point mutation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Multilocus sequence typing wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Community fingerprinting wikipedia , lookup
Metalloprotein wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Deoxyribozyme wikipedia , lookup
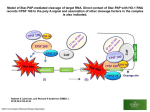
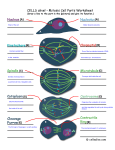
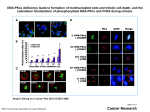
Truncation of Protein Sequences for Fast Profile Alignment with Application to Subcellular Localization Man-Wai MAK and Wei WANG The Hong Kong Polytechnic University Sun-Yuan KUNG Princeton University Contents 1. Introduction – – 2. Speeding Up the Prediction Process – – – 3. 4. Cell Organelles and Proteins Subcellular Localization Signal-Based vs. Homology-Based Methods Predicting Cleaving Site Location Truncating Profiles vs. Truncating Sequences Perturbational Discriminant Analysis Experiments and Results Conclusions 2 Organelles • • • Cells have a set of organelles that are specialized for carrying out one or more vital functions. Proteins must be transported to the correct organelles of a cell to properly perform their functions. Therefore, knowing the subcellular localization is one step towards understanding the functions of proteins. 3 Proteins and Their Subcellular Location 4 Subcellular Localization Prediction Two key methods: 1. Signal-based 2. Homology-based 5 Signal-Based Method Source: S. R. Goodman, Medical Cell Biology, Elsevier, 2008. Cleavage site • The amino acid sequence of a protein contains information about its organelle destination. • Typically, the information can be found within a short segment of 20 to 100 amino acids preceding the cleavage site. • Signal-based methods (e.g. TargetP) can determine the cleavage site location 6 Homology-Based Method N-dim alignment vector Full-length Query Sequence Align with each of the training sequences 1 . . . SVM classifier Subcellular Location N S(1)=KNKA··· S(2)=KAKN··· · · S(N)=KGLL··· Full-length Training sequences Advantage: • Can predict sequences that do not have cleavage sites. Drawback: • Given a query sequence, we need to align it with every training sequence in the training set, causing long computation 7 time. Sequences Length Distribution Cleavage Site Length distribution of Seq. SP Occurrences of Seq. Ext: 21 mTP 820 Cleavage Site Mit: 1050 35 cTP Cleavage Site Chl: Sequence Length 18 760 • Many sequences are fairly long, thus, aligning the whole sequence will take long computation time. • cTP, mTP and SP are under 100 AAs only and contain the most relevant segment. • Computation saving can be achieved by aligning the signal segments only. 8 Proposed Method: Aligning the Segments that Contain the Most Relevant Info. N Amino Acid Sequence truncate … C Signal-based Cleavage Site Predictor (e.g. TargetP) Cleavage Site Homology-based Method Subcellular Location Truncated sequence 9 Aligning Profiles Vs. Aligning Sequences Scheme I : Truncate the profiles Scheme II : Truncate the sequences Scheme I PSIBLAST Long profile Cut short profile Pairwise Alignment Score Vector SVM or KPDA Subcellular Location Cut short sequence PSIBLAST short profile Pairwise Alignment Score Vector SVM or KPDA Subcellular Location Query Sequence Scheme II 10 Perturbational Discriminant Analysis Input and Hilbert Spaces: Input Space Hilbert Space Empirical Space: Empirical Space K ( x1, x) k ( x) N 11 Perturbational Discriminant Analysis • The objective of PDA is to find an optimal discriminant function in the Hilbert space or empirical space: • The optimal solution (see derivation in paper) in the empirical space is • ρ represents the noise (uncertainty) level in the measurement. It also ensures numerical stability of the matrix inverse. • Ρ = 1 in this work. 12 Perturbational Discriminant Analysis Example on 2-D Data 3 classes of 2-dim data in the input space Projection onto the 2-dim PDA space RBF kernal matrix K Decision boundaries in the input space 13 Perturbational Discriminant Analysis Application to Sequence Classification Training sequences Training Profiles PSI-BLAST Test sequence Test Profile PSI-BLAST Pairwise Alignment Align with Training Profiles K Compute PDA Para Compute PDA Score 14 Perturbational Discriminant Analysis Application to Multi-Class Problems 1-vs-Rest PDA Classifier: MAXNET f1 ( x ) fC (x) f 2 ( x) x 15 Perturbational Discriminant Analysis Application to Multi-Class Problems Cascaded PDA-SVM Classifier: Test sequence Project onto (C–1)-dim PDA space 1-vs-rest SVM Classifier Class label 16 Experiments Materials: • Eukaryotic sequences extracted from Swiss-Prot 57.5 • Ext, Mit, and Chl contain experimentally determined cleavage sites • 25% Sequence identity (based on BLASTclust) Performance Evaluation: • 5-Fold cross validation • Prediction accuracy and Matthew’s correlation coefficient (MCC) 17 Comparing Kernel Matrices Kernel matrix (Scheme I) Scheme I PSIBLAST Long profile Cut short profile Pairwise Alignment Score Vector SVM or KPDA Subcellular Location Cut short sequence PSIBLAST short profile Pairwise Alignment Score Vector SVM or KPDA Subcellular Location Query Sequence Scheme II Kernel matrix (Scheme II) 18 Sensitivity Analysis Seq Cut Seq. at p±x Subcellular Localiation Accuracy (%) p: gournd-truth cleave site Subcellular localization (PairProSVM) Cyt/Nuc Ext Overall Subcellular location • The localization performance degrades when the cut-off position drifts away from the ground-truth cleavage site. Mit Chl • mTP and cTP are more sensitive to the error of cleavage site prediction than Ext. Ground-truth cleavage site p-16 p-8 p-2 p p+2 p+16 p+32 p+64 Cut-off Position 19 Csite Prediction ACC(%) Performance of Cleavage Site Prediction • Conditional Random Field (CRF) is better than TargetP(Plant) in terms of predicting the cleavage sites of signal peptide (Ext) but is worse than TargetP(Nonplant). • CRF is slightly inferior to TargetP in predicting the cleavage sites of mitochondria, but it is significantly better than TargetP in predicting the cleavage site of chloroplasts. Category 20 Comparing Profile Creation Time Scheme I PSIBLAST Long profile Cut short profile Pairwise Alignment Score Vector SVM or KPDA Subcellular Location Cut short sequence PSIBLAST short profile Pairwise Alignment Score Vector SVM or KPDA Subcellular Location Query Sequence Scheme II Findings: Profile creation time can be substantially reduced by truncating the protein sequences at the cleavage sites. 21 Training and Classification Time 1-vs-rest SVM Classifier MAXNET f1 ( x ) fC (x) f 2 ( x) x Project onto (C–1)-dim PDA space Findings: The training time of 1-vs-rest PDA and Cascaded PDASVM are substantially shorter than that of SVM. 22 Compare with State-of-the-Art Localization Predictors Query seq. Subcellular localization (SubLoc/TargetP) Subcellular location Cleavage site prediction (TargetP/CRF) Subcellular localization (PairProSVM) Localization Accuracy (%) MCC Conditional Random Fields Findings: In terms of localization accuracy, the proposed “Signal+Homology” method performs slightly better than the signal-based TargetP and is substantially better than the homology-based SubLoc. 23 Conclusion • Fast subcellular-localization-prediction can be achieved by a cascaded fusion of signal-based and homology-based methods. • As far as localization accuracy is concerned, it does not matter whether we truncate the sequences or truncate the profiles. However, truncating the sequence can save the profile creation time by 6 folds. 24 Compare with State-of-the-Art Localization Predictors Query seq. Subcellular localization (SubLoc/TargetP) Subcellular location Cleavage site prediction (TargetP/CRF) Subcellular localization (PairProSVM) 25 Performance of Cascaded Fusion Time (hr.) Time Subcellular localization accuracy Fulllength Seq. Seq. with Csite predicted by TargetP(P) Seq. with Csite predicted by TargetP(N) Acc (%) • The computation time for full-length profile alignment is a striking 116 hours • Our method not only leads to nearly a 20 folds reduction in computation time but also boosts the prediction performance. Seq. with Csite predicted by CRF 26 Fusion of Signal- and Homology-Based Methods 1) Cleavage site detection. The cleavage site (if any) of a query sequence is determined by a signal-based method. 2) Pre-sequence selection. The pre-sequence of the query is obtained by selecting from the N-terminal up to the cleavage site. 3) Pairwise alignment. The pre-sequence is aligned with each of the training pre-sequences to form an N-dim vector, which is fed to a one-vs-rest SVM classifier for prediction. N Amino Acid Sequence truncate … C Signal-based Cleavage Site Predictor Cleavage Site Homology-based Method Subcellular Location Pre-sequence 27 27 Perturbational Discriminant Analysis Spectral Space: Define the kernel matrix K can be factorized via spectral decomposition into Empirical Space K ( x1, x) Spectral Space e ( x ) N 28