* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Review of Key Microbial Groups
Neisseria meningitidis wikipedia , lookup
Carbapenem-resistant enterobacteriaceae wikipedia , lookup
Quorum sensing wikipedia , lookup
Cyanobacteria wikipedia , lookup
Bacteriophage wikipedia , lookup
Human microbiota wikipedia , lookup
Unique properties of hyperthermophilic archaea wikipedia , lookup
Trimeric autotransporter adhesin wikipedia , lookup
Bacterial morphological plasticity wikipedia , lookup
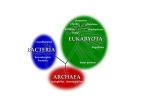
Review of Key Microbial Groups A.Domain Bacteria B.Domain Archaea C.Domain Eucarya Domain Bacteria: General Features ● Prokaryotic cell structure – – ● ● DNA organized in nucleoid; no nuclear membrane, nucleolus, or histones No complex membranous organelles (e.g. endoplasmic reticulum, golgi apparatus) Cell walls containing peptidoglycan found in most groups of bacteria Some features of gene expression (mRNA, tRNA, RNA polymerase, ribosomes) are more similar to archaea; some are more similar to eucarya Domain Bacteria: General features ● Metabolic strategies found in Bacteria: – – – Chemoheterotrophy Chemolithotrophy Photosynthesis Domain Bacteria: General features ● Cell Wall Structures in Bacteria – Gram-negative cell wall ● ● ● – Gram-positive cell wall ● ● – Thick layer of peptidoglycan Teichoic acids “Acid-fast” bacteria ● ● – Outer membrane containing lipopolysaccharide Thin layer of peptidoglycan Periplasmic space Bacteria in Phylum Actinomycetes with high concentrations of mycolic acid Detected by acid-fast staining Mycoplasmas ● Bacteria in Phylum Firmicutes with no cell wall Domain Bacteria: General features ● Other structural features found in Bacteria – – – – – – – – Plasma membrane Capsules Pili or Fimbrae Cytoplasmic inclusions Bacterial DNA Ribosomes Flagella Spores http://science.kennesaw.edu/~jhendrix/bio3340/handouts/microcells.ppt Domain Bacteria: General features ● Identification of the Bacteria – – – – – – – – – – Colony morphology Cell shape & arrangement Cell wall structure (Gram staining) Special cellular structures Biochemical characteristics Serological tests G+C content DNA hybridization DNA fingerprinting Nucleic acid sequencing http://science.kennesaw.edu/~jhendrix/bio3340/handouts/microtaxonomy.ppt Domain Bacteria: Major Groups ● Phylum Proteobacteria – “Gram-negative” type cell wall architecture ● ● ● – Outer membrane with lipopolysaccharide and porin protein Thin layer of peptidoglycan Notable periplasmic space containing transport proteins and hydrolases A very large and metabolically diverse phylum; various groups utilizing chemoheterotrophy (both respiration & fermentation), chemolithotrophy, photosynthesis Domain Bacteria: Major Groups ● Phylum Proteobacteria (cont.) – Major groups of proteobacteria ● ● Enterobacteriacea: “Gram-negative enterics;” common intestinal flora and pathogens; both respiratory and fermentative metabolisms; facultatively anaerobic; oxidase negative; includes genera Escherichia, Klebsiella, Enterobacter, Citrobacter, Serratia, Proteus, Salmonella, Shigella, Yersinia Pseudomonadaceae: Genus Pseudomonas and related genera; common soil and aquatic organism; usually aerobic; oxidase positive; use EntnerDouderoff glycolysis instead of EMP glycolysis; often can metabolize unusual carbon substrates such as aromatic hydrocarbons Domain Bacteria: Major Groups ● Phylum Proteobacteria (cont.) – Major groups of proteobacteria (cont.) ● ● ● ● ● Purple nonsulfur photosynthetic bacteria, e.g. Rhodospirillum Green sulfur photosynthetic bacteria, e.g. Chlorobium Nitifying bacteria: Nitrosomonas (oxidizes ammonium to nitrite) and Nitrobacter (oxidizes nitrite to nitrate) Nitrogen fixing bacteria: Rhizobium (symbiotic in root nodules); Azotobacter (free-living) Various human pathogens in phylum Proteobacteria: Neisseria, Vibrio, Haemophilus, Rickettsia, Coxsiella, Bordetella, Legionella, Camphylobacter, Helicobacter Domain Bacteria: Major Groups ● Phylum Firmicutes – – – “Low G-C” Gram-positive bacteria Cell walls consist of thick layers of extensively crosslinked peptidoglycan (exception: Mycoplasma) Notable genera ● ● ● Clostridium: Strictly anaerobic, spore-forming rods; common in soil; includes botulism & tetanus; significant contaminant in food industry & medicine Bacillus: Facultatively anaerobic, spore-forming rods; common in soil; frequent contaminant; includes Bacillus anthracis Mycoplasma: Have no cell walls; respiratory tract flora & pathogens of humans & other animals Domain Bacteria: Major Groups ● Phylum Firmicutes – Notable genera ● ● ● Lactobacillus: Facultatively anaerobic nonsporeforming rods; oral or intestinal flora; found in sevral dairy products such as yogurt Staphylococcus: Catalase-positive cocci; common skin flora; virulent strains of Staph. aureus are associated with skin infections, food poisoning, and toxic shock syndrome Streptococcus, Enterococcus, Lactococcus: Catalasenegative cocci; diverse group with numerous skin, oral, and intestinal flora as well as several important pathogens such as Streptococcus pyogenes (Group A pyogenic strep) Domain Bacteria: Major Groups ● Phylum Actinomycetes – – – “High G-C” Gram-positive bacteria Cell walls consist of thick layers of extensively crosslinked peptidoglycan Notable genera ● ● ● Corynebacterium: Facultatively anaerobic, irregular rods; coryneform arrangement; common soil & skin flora; common laboratory contaminant Micrococcus: Facultatively anaerobic cocci; tetrads or sarcinae; yellow or pink pigmentation; common soil flora;common laboratory contaminants Actinomyces, Streptomyces; Common soil organism; filamentous growth often mistaken for mold Domain Bacteria: Major Groups ● Phylum Actinomycetes – Notable genera ● Mycobacterium: Acid-fast rods; high concentration of mycolic acid in the cell wall make them difficult to gram stain; certain species are skin and soil flora; includes tuberculosis and leprosy Domain Bacteria: Major Groups ● Phylum Bacteroidetes – – – A group of gram-negative, strictly anaerobic bacteria Most are intestinal and oral flora in humans and animals; some are pathogens Example: genus Bacteroides Domain Bacteria: Major Groups ● Phylum Cyanobacteria – – – The “blue-green algae” Carry out oxygenic photosynthesis Have thylakoid membranes, chlorophyll a, and photosystem II Domain Bacteria: Major Groups ● Phylum Chlamydiae – – ● A group of gram-negative, obligately intracellular parasites Genus Chlamydia Phylum Spirochaetes – – – Characterized by flexible helical-shaped cells Cells covered by an outer sheath and are motile by a modified flagellar structure called an axial filament Example: Treponema pallidum (syphillis) Domain Archaea: General Features ● ● ● ● Prokaryotic cell structure Cell walls have no peptidoglycan; some archaea have pseudomurein or other polymers Some features of gene expression (mRNA, tRNA, RNA polymerase, ribosomes) are more similar to bacteria; some are more similar to eucarya Metabolic strategies found in Archaea: – – Chemoheterotrophy Chemolithotrophy Domain Archaea: Major Groups ● Methanogenic archaea – ● Extremely thermophilic archaea – – ● Sulfate reducers, Archaeoglobus Sulfur reducers, Desulfurococcus, Sulfolobus Extremely halophilic archaea – ● Methanobacterium, Methanococcus Halobacterium, Halococcus Cell wall deficient archaea – Thermoplasma Domain Eucarya: General Features ● ● ● ● Eukaryotic cell structure Cell walls vary; none in “animal-like” cells; cellulose in algae most others, with additional polysaccharides in different groups (e.g., chitin in many fungi) Some features of gene expression (mRNA, tRNA, RNA polymerase, ribosomes) are more similar to bacteria; some are more similar to archaea Metabolic strategies found in Eucarya: – – Chemoheterotrophy (respiration in mitochondria) Photosynthesis (in chloroplasts)