* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Protein Structure Hierarchy
Expression vector wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Point mutation wikipedia , lookup
Catalytic triad wikipedia , lookup
Genetic code wikipedia , lookup
Interactome wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Biochemistry wikipedia , lookup
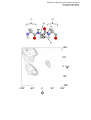
Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 Protein Structure Hierarchy Introduction With great insight - before the first 3D protein structure was determined, the Danish chemist Kaj Linderstrøm-Lang (the father of physical biochemistry), reasoned that there should be at least four levels of structural organization present in protein structure. In Linderstrøm-Lang's hierarchy of protein structure, each level is characterized by a particular type of organizing force and higher levels of organization are composed of elements described by the previous level. We now know that this organization is an Glutaredoxin 3 (E. coli) oversimplification, but the organization of structure into levels is still useful from a pedagogical viewpoint. Altogether, 80-90% of amino acids in globular proteins are in one of the three classical secondary structures: α helix, β sheet, or turn. Here we will begin with a description of primary structure and continue on to the higher levels of protein structure. Objectives 1. 2. 3. 4. 5. Gain experience using different macromolecular representations in RasMol. Measure conformational details (distances, angles) in a protein. Extract information from the coordinate file header. Investigate disulfide bonds in ribonuclease A. Use the molecular-graphics program RasMol to analyze the structural features of a protein. 6. Recognize the main elements of secondary structure within a protein fold. Primary structure of ribonuclease A This computer lab will use the enzyme ribonuclease A to explore secondary structure in proteins. One basic question often asked is what is the primary sequence of a particular structure entry. Exercises: 1. 3-letter code. Last time you downloaded the coordinate file for 5RSA. Search the header for the amino acid sequence. Copy out the sequence and place it in a text editor. 2. 1-letter code. If you do not like the format (3-letter code) of what you find, you can revisit the RCSB homepage and search for the entry 5RSA again. Here you will be able to find the sequence in a number of formats. The FASTA format is a convenient 1-letter code format. Copy this form of the sequence to the same file. 3. Formatting sequences. Try to format your sequence in a useful manner so it will be easy to find a particular residue number. If you are using a word processor as opposed to a text editor, be careful of the choice of font. Proportional spaced fonts (the ones that look nice e.g., Times New Roman) will not allow sequences to be compared as some letters like “W” get more space than others like “I”. The solution is to choose a non-proportional spaced font like Courier. -1- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 4. Take the amino acid challenge. If you did not have time for this during lab 1, try it now! Follow the link for this lab on the course web site to amino acid quiz. Each link represents a coordinate file of one of the 20 naturally-occurring amino acids. It is important to learn to identify the amino acids visually in the different representations. Display a protein structure using different representations Find your copy of the coordinate file for ribonuclease A and open it with RasMol. With this exercise, you will cover most of the important commands and manipulations in RasMol. Exercises: 1. Change the display. Open ribonuclease A in RasMol. By default, all atoms of all structures are displayed by using a “wireframe” representation, atoms are invisible and bonds are drawn as lines. Explore the different display modes (wireframe, backbone, sticks, spacefill, ball & stick, ribbons, strands and cartoons) from the “pull-down” menus. Think about the advantages and disadvantages of these different representations are you see them. Choosing the best way to view a structural detail can only be learned through experience. Disulfide bonds and RasMol scripts Often the primary sequence is defined as the covalent structure of the protein. This then includes the covalent disulfide bonds which form crosslinks within and sometimes between proteins. They are a very important structural detail that can be elucidated through chemical sequencing, or by inspecting the 3D structure if it has been determined at high enough resolution. Exercises: 1. Identification of disulfide bonds. Print out a copy of your sequence for ribonuclease A. Which cysteine residues are paired? One way to find put is to open the coordinate file in a text editor and search for the information in the “HEADER”. Indicate which cysteine residues are in disulfide bonds with lines. 2. Display of Disulfide bonds. Now display the disulfide bonds in ribonuclease A using RasMol. Load the 5RSA structure into RasMol and display the protein using the “backbone” representation from the menu (“Display -> Backbone”). In RasMol, “backbone” display as a menu option means “virtual” bonds connecting Cα positions. Once you have convinced yourself of this, label the cysteine residues at the Cα position. RasMol> select cys.ca RasMol> label %r%s RasMol can identify (usually) disulfide bonds with the terminal command “ssbonds”. This will output the number of disulfide bonds identified in the terminal window and display dotted lines connecting the cysteine sulfur atoms (the default behavior). As these atoms are not displayed, they will appear to float in the structure. For backbone and cartoons it is better to draw the disulfide bonds between the Cα positions. To do this use the commands: -2- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 RasMol> select all RasMol> set ssbonds backbone RasMol> ssbonds 50 RasMol> color ssbonds yellow Notice how the disulfide bonds form crosslinks in the protein structure. Now display the entire cysteine side chain and draw the disulfide bonds between the sulfurs. RasMol> set bondmode or RasMol> select cys and sidechain RasMol> wireframe 50 RasMol> set ssbonds sidechain RasMol> ssbonds 50 From the “Options” menu, the side chain hydrogens can be removed from the display. Disulfides have characteristic values for χ3 +/- 90° yielding right- and left-handed disulfide bonds. What are the values for χ3 in this protein. Use your chemical intuition to rationalize the value of this dihedral, whereas staggered rotamers of carbon are –60, 60, and 180°. 3. RasMol scripts. Combinations of commands can be saved as a text file and read into RasMol to produce the same image as if the commands were entered, one at a time, into the terminal window. Make a RasMol script using the above commands to produce this type of display for a generic protein. Save the file and test is after opening a new PDB file by entering the “script your.filename” command. A working knowledge of RasMol scripts is VERY useful. Higher order structure The description of higher order structure is, in general, less unambiguous than the description of the covalent (primary) structure. Exercises: 2. Describe the secondary/tertiary structure composition of ribonuclease. One way to get an overview of the protein is to display the protein in cartoon or backbone format and to color by structure. Details like disulfide bonds can also be added. RasMol> set ssbonds backbone RasMol> ssbonds 100 RasMol> color ssbonds green Labels containing residue numbers can also be useful. RasMol> select (1,25,50,75,100,124) and *.CA RasMol> label %r Labels can be removed at anytime by the command RasMol> labels off -3- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 Specific residue positions can be identified by clicking the left mouse button on the atom. The atom selected appears in the terminal window. Note: atom identification does not work for cartoons, ribbons, or strands. Prepare a simple 2D plot identifying the secondary structure segment as a function of the amino acid sequence (shown below). Use arrows to indicate strands of a sheet and cylinders to indicate helices. 1 10 20 30 40 50 60 KETAAAKFERQHMDSSTSAASSSNYCNQMMKSRNLTKDRCKPVNTFVHESLADVQAVCSQ 70 80 90 100 110 120 KNVACKNGQTNCYQSYSTMSITDCRETGSSKYPNCAYKTTQANKHIIVACEGNPYVPVHFDASV How many structural domains does the protein have? How does RasMol know where the secondary structures are? Is the beginning and end of the secondary structure unambiguous? Where might the active site be? Identification of H-bonds In this exercise, you will analyze a segment of α helix and a segment of β sheet for the characteristic hydrogen bonding pattern and measure the distances between hydrogen-bonded atoms. Exercises: 1. α helix. Examine one α helix and identify the characteristic hydrogen-bonding pattern. Select a portion of the polypeptide chain (e.g., residues 5-12) that is in an α helical conformation for display. Now we will use the “real” bonds between backbone atoms. RasMol> select all “DisplayMenu->Wireframe” RasMol> restrict backbone and 5-12 RasMol> wireframe 40 RasMol> color cpk RasMol> center 9 It will be helpful to have labels. RasMol> select 5-12 and *.CA RasMol> label %r Note that in most structures determined by diffraction techniques, there are no hydrogen atoms (hydrogen atoms do not diffract X-rays very well). Therefore, identification of hydrogen bonds will require that you recognize the donor (amide nitrogen) and acceptor (carbonyl oxygen). Protein backbone hydrogen bonds normally have a donor-acceptor distance of ~ 3.0 Å. Use RasMol to measure these distances in your helical segment. -4- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 RasMol> set picking distance RasMol> set picking monitor Now use the mouse to select possible hydrogen bond donor-acceptor pairs. The selection will be shown as a dotted line connecting the two atoms and the distance will be displayed. To remove an unwanted selection, simply re-select the two atoms. When you have found the hydrogen bonds in this segment, reset the identification mode of RasMol. RasMol> set picking ident RasMol> monitors off Regular H-bonds. Using the template below, finish drawing the polypeptide chain to represent residues 5-12. Draw line segments connecting the atoms involved in regular α-helical hydrogen bonds: 2. O H N 5 C H H N C H O Typical α-helical hydrogen bonds are between the carbonyl oxygen of residue i and the amide hydrogen of residue i + ___ ? Measure backbone dihedral angles φ, ψ, and ω The fold of the polypeptide backbone is defined by three dihedral angles φ, ψ, and ω for each amino acid in the sequence. Due to partial double bond character, the amide bond (dihedral angle ω) is more-or-less planar. That is to say the atoms Cα(i), C(i), N(i+1), Cα(i+1), lie in a plane. There are two possible planar orientations which can be called cis (ω ≈ 0°) and trans (ω ≈ 180°). The trans conformation is slightly more stable than the cis and is the predominant form in folded proteins. The regular secondary structures can be conveniently identified by a plot of φ versus ψ commonly referred to as a "Ramachandran" plot. Not all values of φ and ψ are possible due to steric hindrance in the polypeptide chain. Thus another use of a Ramachandran plot is as a check of the accuracy of a structural model. Residues with values outside the "allowed" regions may represent inaccuracies. Exercises: 1. Dihedral angles of an α helix. Select a segment of α helix (5-7 residues long). Display only backbone atoms (N, Cα, C, O and perhaps Cβ). Using the definitions provided below, measure the dihedral angles φ, ψ, and ω and record them in the table provided. Enter the values for φ and ψ as points on the Ramachandran plot (page 7). RasMol> restrict (backbone or *.CB) and 5-12 RasMol> wireframe 40 RasMol> select *.CA and 5-12 RasMol> label %r -5- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 The commands for measuring the dihedral angles are: RasMol> set picking torsion Use the mouse to select the following four atoms in succession for each angle: φi: Ci-1, Ni, Cαi, C’i ψi: Ni, Cαi, Ci, Ni+1 ωi: Cαi, Ci, Ni+1, Cαi+1 The values will be displayed in the terminal window. Fill in the numbers on the answer sheet. 2. Dihedral angles of a β strand. Select a piece of β sheet (e.g., residues 42-48, 8086, 98-103). Again, display only backbone atoms, measure the dihedral angles φ, ψ, and ω of the central sheet (80-86) and record them in the table provided. Add the values for φ and ψ as points on the Ramachandran plot (next page). RasMol> restrict backbone and (42-48,80-86,98-103) RasMol> wireframe 40 RasMol> select 80-86 and *.CA RasMol> label %r Fill in these values too on the anwer sheet. Do the values of φ and ψ fall in “allowed” regions? How do they compare to the average values for these secondary structures (see compendium 2.7.1-2.7.3)? What are the relative orientations of the strands in the sheet? A new feature in RasMol 2.7.1 allows for the dihedral angles φ and ψ to be written out directly to an ASCII file suitable for subsequent plotting (Excel) using the command RasMol> write RDF myfile.name -6- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 i+1 i-1 φi ωi O HN C N Cα Hα Cβ ψi 180° 81 90° 0 ψ 6 -90° -180° -90° 0 φ -7- 90° -180° 180° Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 Turns Turns are the third of the three "classical" secondary structures with approximately 1/3 of all residues in globular proteins are contained in turns that serve to reverse the direction of the polypeptide chain. This is perhaps not so surprising as the diameter of the average globular protein domain is roughly 25 Å (an extended polypeptide conformation would require ~7 residues to traverse the domain before having to change directions). Turns are located primarily on the protein surface and accordingly, contain polar and charged residues. Antibody recognition, phosphorylation, glycosylation, hydroxylation, and intron/exon splicing are found frequently at, or adjacent to turns. However it is not clear if this is due to specific recognition or simply the surface location of turns. Type I turns. Note the hydrogen bond between CO of residue i and NH of residue i+3. Many have speculated on the role of turns in the folding of globular proteins. Perhaps the two extremes can be classified as pacifists and activists. The pacifists view turns as a weak link in the polypeptide chain, allowing the other secondary structures (helix and sheet) to determine the conformational outcome. The activists, (encouraged by the recent experimental finding of "turn-like" structures in short peptides in aqueous solution) view turns as structure nucleating segments, formed early in the folding process. Neither is entirely accurate and evidence can be given to support each view. Exercises: 1. Rnase A. RasMol can automatically identify turns in a coordinate file of a protein structure. Focus on the turns in Rnase A. To classify turns, one will need to include the backbone atoms N, Ca, C’ and O as type I and II turns have a characteristic hydrogen bond between C=O of residue i and HN of residue i+3. The following is a suggestion for such a display. RasMol> structure // have RasMol calculate SS structure RasMol> wireframe off RasMol> restrict protein RasMol> strands // trace backbone RasMol> select backbone and turns RasMol> wireframe 40 RasMol> select turns and *.ca // labels are nice RasMol> label %r%n RasMol> color labels green 2. Thioredoxin. To see some “classical” examples of reverse turns, we will use the small globular protein thioredoxin from E. coli (PDBid code 2TRX). Thioredoxin is one of a handful of proteins that shuttle reducing from NADH to ribonucleotides in the synthesis of deoxyribonucleotides. Visit the RCSB and download the structure. If the above (minus the first statement) is placed into a RasMol script, it can simply be applied using the “script” command. -8- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 Appendix I Plotting data using MicroSoft Excel ASCII text files containing columns of data can be opened for analysis in MS Excel. To open a text file and create a plot, follow the following steps. Importing ASCII files into Excel Step 1. Start Excel Step 2. Select File-Open to get a file window and select the text file containing your data. You will be prompted by the text Import wizard. Step 3. choose “delimited” and “finish” Step 4. remove the header from the file (click on the row number of the first row to delete and while holding down <SHIFT>, select the row number of the last row to delete. Select Edit/Delete so the data looks like the following with the residue number, φ,ψ in columns B, C and D, respectively. GLU THR ALA ALA ALA 2 3 4 5 6 -53.1 -72 -61.1 -65.4 -68.8 126.8 158.8 -39.6 -45.7 -36.9 Simple plot in Excel Step 1 select the letter of the column to be on the x-axis (C) then while pressing <SHIFT> select the column to be on the y-axis (D). Step 2 Click on the Chart Wizzard. Select a type like XY (Scatter) and then select <finish>. And a plot is produced. Step 3 You can make the plot look a bit nicer by double clicking the axis and setting the limits for both x and y to –180 to 180, and adding the labels for each axis. -9- Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 Appendix II Allowed regions in φ,ψ space The allowed regions were determined from 378 different crystal structures with a resolution of 2.5 A or better. The regions where the density of points (φ,ψ) from this statistic is highest are marked with color. The black regions contain 80% of all points, the medium grey, 95% and the light grey 98% and yellow regions together 95%, all colored regions together 98%. There is a different plot for all non-Gly/Pro residues, one for Val/Ile/Thr and one for Gly. Gly ”normal” Thr, Val, Ile - 10 - Södertörns högskola & Karolinska Institutet Structural biochemistry Computer lab 2 HT05 Answer sheet Use this form to fill in answers to the questions appearing in the text as you go through it. What are the values of the χ3 torsion in Rnase A? How many structural domains does the protein have? How does RasMol know where the secondary structures are? Is the beginning and end of the secondary structure unambiguous? Where might the active site be? Between which atoms do we find typical αhelical hydrogen bonds? How long are these typically (distance between donor-acceptor, or distance between hydrogenacceptor may be used, but specify which it is)? Do the values of φ and ψ fall in “allowed” regions? How do they compare to the average values for these secondary structures (see compendium 2.7.1-2.7.3)? What are the relative orientations of the strands in the sheet? Helix residue 6 7 8 9 10 11 φ -68.8° ψ -36.9° ω 178.1° Strand residue 81 82 83 84 φ -133.8° ψ 154.7° ω 178.0° - 11 -