* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Poster - Department of Computer Science
Interactome wikipedia , lookup
Magnesium transporter wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Expression vector wikipedia , lookup
Metalloprotein wikipedia , lookup
Western blot wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Point mutation wikipedia , lookup
Protein purification wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Proteolysis wikipedia , lookup
De novo protein synthesis theory of memory formation wikipedia , lookup
Next Generation Evolutionary Sampling and Energy Function Guided Ab Initio Protein
Structure Prediction
Example of 3DIGARS-PSP modeling results on known Hard E. Coli and Protease Inhibitor proteins
Avdesh Mishra, Md Tamjidul Hoque
email: {amishra2, thoque}@uno.edu
Additional beta sheet region
Additional beta sheet region
One amino acid is not
assigned to beta sheet
One amino acid is
added to the beta sheet
One amino acid is not
assigned to beta sheet
Department of Computer Science
University of New Orleans, LA, USA
One amino acid is
added to the beta sheet
Introduction
The confirmation of a protein is vital to understand the function it
performs within the cell. Towards this goal, we developed a
computer program that applies a memory assisted evolutionary
algorithm to sample the energy hyper-surface of the protein
folding process, searching for the global minimum or the native
fold of the protein. Sampling of the energy hyper-surface of the
protein is achieved by novel mutation and crossover operations
based on angular rotation and translation capabilities.
Furthermore, the crossover operations in current generation are
enhanced by the use of the best parents selected from previous
generations. In addition, we employ a knowledge-based novel
energy function, 3DIGARS3.0, which can differentiate the native
structure that corresponds to the most thermodynamically stable
state, compare to the possible decoy structures most effectively.
The 3DIGARS3.0 energy function is an optimized combination of
crucial properties such as hydrophobic versus hydrophilic,
sequence-specific predicted accessibility and ubiquitous phi-psi
characterization.
Missing Helixes
Figure 1 | Cysteine Protease Inhibitor (PDB ID: 1nyc); towards left – superposition of 3DIGARSPSP model on native (initial seeds from Rosetta); towards right – superposition of top Rosetta model
(based on TMScore) on native.
Figure 2 | E. Coli protein (PDB ID: 1pohA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from Rosetta); towards right – superposition of top Rosetta model (based on
TMScore) on native.
Beta sheet predicted correctly
Figure 3 | E. Coli protein (PDB ID: 1pohA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from I-Tasser); towards right – superposition of top I-Tasser model (based on
TMScore) on native.
Missing beta and helix regions
Missing helixes
Helixes are gained
Additional beta region
Additional beta sheet, potential
area of improvement
Additional beta sheet, potential
area of improvement
Missing beta sheet region
Figure 4 | E. Coli protein (PDB ID: 2z9hA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from Rosetta); towards right – superposition of top Rosetta model (based on
TMScore) on native.
Figure 5 | E. Coli protein (PDB ID: 2z9hA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from I-Tasser); towards right – superposition of top I-Tasser model (based on
TMScore) on native.
Figure 6 | E. Coli protein (PDB ID: 2p7vA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from Rosetta); towards right – superposition of top Rosetta model (based on
TMScore) on native.
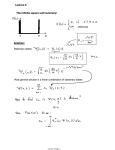
Methods
Additional helixes, potential
area of improvement
Missing Helixes
Backbone Models
Initialize Population for GA
using Single Point Angular
Mutation
Dataset of 4332
Protein Structures
Obtain Secondary
Structure (SS) and Φ, Ψ
Angles using DSSP
Save Best Model in Memory
Select 5% Elite Models
Missing beta sheet
Generate Frequency
Distribution of Φ, Ψ
Angles and SS Types
Perform Memory Assisted
Crossover @ 70 %
Missing helixes
Additional beta sheet
Additional beta sheets, potential
area of improvement
Additional beta sheets
Figure 7 | E. Coli protein (PDB ID: 2p7vA); towards left – superposition of 3DIGARS-PSP model on
native (initial seeds from I-Tasser); towards right – superposition of top I-Tasser model (based on
TMScore) on native.
Figure 8 | E. Coli protein (PDB ID: 1k4nA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from Rosetta); towards right – superposition of top Rosetta model (based on
TMScore) on native.
Note: Natives are shown in cyan and pink and Models are shown in red and yellow
Results
Fill Rest Randomly
Perform Angular Mutation
@ 60%
Calculate Fitness using
3DIGARS3.0
Save Models
Generation
< 2000
End
Best Models
Effective use of Ramachandran Plot
Effective initialization and use of associated memory
Development of new operator to implement move sets
Ongoing Research
Acknowledgements
Authors gratefully acknowledge the Louisiana Board of Regents through the
Board of Regents Support Fund, LEQSF (2013-16)-RD-A-19.
Figure 9 | E. Coli protein (PDB ID: 1k4nA); towards left – superposition of 3DIGARS-PSP model
on native (initial seeds from I-Tasser); towards right – superposition of top I-Tasser model (based on
TMScore) on native.
Discussions and Conclusions
In past we have shown that our energy function,
3DIGARS3.0 outperforms the state-of-arts method
significantly.
Also, in our prior work we have shown that our associate
memory based sampling algorithm provides superior
performance.
In this work, we are working on to find the right
combination of our energy function and the sampling
algorithm to have better prediction of 3D structure of protein
in comparison to the state-of-art approaches.
To this end, we have been able to successfully apply
dihedral angles mutation by rotation and crossover by
protein segment translation rules to enhance the mutation
and crossover operations of the sampling algorithms.
We are working on case by case basis to obtain an accurate
prediction of the useful secondary structures in a protein.
Towards this, we have utilized the Ramachandran Plot
information within our sampling algorithm.
We have found that the use of Ramachandran Plot yields in
significant improvement.
We are exploring on the topics such as effective use of
Ramachandran Plot, move sets and associated memory to
find more efficient and effective rules to apply within the
sampling algorithm.
We plan to further improve the PSP problem by combining
3DIGARS and sDFIRE energy function in near future to
make it further robust.