* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Li, 2004
Survey
Document related concepts
Transcript
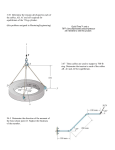
A map of the interactome network of the metazoan C. elegans. Li S, Armstrong CM, Bertin N, Ge H, Milstein S, Boxem M, Vidalain PO, Han JD, Chesneau A, Hao T, Goldberg DS, Li N, Martinez M, Rual JF, Lamesch P, Xu L, Tewari M, Wong SL, Zhang LV, Berriz GF, Jacotot L, Vaglio P, Reboul J, Hirozane-Kishikawa T, Li Q, Gabel HW, Elewa A, Baumgartner B, Rose DJ, Yu H, Bosak S, Sequerra R, Fraser A, Mango SE, Saxton WM, Strome S, Van Den Heuvel S, Piano F, Vandenhaute J, Sardet C, Gerstein M, Doucette-Stamm L, Gunsalus KC, Harper JW, Cusick ME, Roth FP, Hill DE, Vidal M. Science. 2004 Jan 23;303(5657):540-3 MEDGEN 505 Jan Stöpel Goal How do protein-protein interactions relate to multicellularity? Approach 1. Select candidate genes 2. Use high-throughput Y2H to screen for interactions 3. Verify coverage 4. Verify accuracy of interaction 5. Build network 6. …future use Candidate genes by BLAST ancient S. cerevisae C. elegans A. thaliana D.melanogaster H. sapiens worm multicellular (3024) Making Y2H constructs Used Invitrogen Gateway® system utilizes phage recombination QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. • fast • automated • allows large scale tagging and plasmid shuffling Yeast 2 hybrid If proteins interact, don’t interact, reporter reporter gene isgene switched remains on inactive and strain can grow not grow on selective on selective media media QuickTime™ QuickTime™and andaa TIFF TIFF(Uncompressed) (Uncompressed)decompressor decompressor are areneeded neededtotosee seethis thispicture. picture. lost 105 candidates due to autoactivation or toxicity Screen 1873 Gal4-BD constructs Vs. AD-wormcDNA contains ~11000 ORFs AD-yfg multiple junctions AD-ORFeome contains ~11000 ORFs AD-yfg single junction Screen 858 1299 1892 Results # interactions ∑ DB-X/AD-Y Core-1/ Core2 858/1299 2135 502/1039 NonCore 1892 1892 531/1395 ADworm/ AD-ORF 2783/ 1505 4027 239 (overlap) Coverage coverage efficiency: ~ 10% Literature search for known and verified interactions of any ORF used as bait. 108 interactions AD-wormcDNA+ AD-ORFeome DB-X (literature) (8+2)/108 Verification using GST-bait and Myc-prey fusion in Myc immunoprecipitation accuracy 82% 59% 35% Building a network sources: HT-Y2H literature search interlogs scaffolds (4027) (108) (949) (?) Interlogs: in silico approach to identify conserved interactions Y2H data set from S.cerevisiae BLASTed against C.elegans proteome scaffolds: Y2H data sets for biological processes of C.elegans protein degradation, vulval development, DNA damage response, germline formation Evolution is using old and new 748 1314 836 Transcriptome vs Interactome