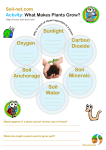
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download a multi-omics approach to alleviating
Soil respiration wikipedia , lookup
Soil salinity control wikipedia , lookup
Canadian system of soil classification wikipedia , lookup
Crop rotation wikipedia , lookup
Terra preta wikipedia , lookup
Soil compaction (agriculture) wikipedia , lookup
Plant nutrition wikipedia , lookup
No-till farming wikipedia , lookup
Soil food web wikipedia , lookup
Project proposal form – 2017 entry Project title: Bacterial biofertilisation of phosphorus in the rhizosphere of agricultural crops: a multi-omics approach to alleviating food security Host institution: University of Warwick Theme: Organisms, omics and biogeochemistry Key words: metagenomics, metaproteomics, Pi-scavenging, exoenzymes, rhizosphere Supervisory team: Professor Elizabeth Wellington, School of Life Sciences, University of Warwick, [email protected], Dr Alex Jones, School of Life Sciences, University of Warwick, [email protected] ; Dr Rob Finn, European Institute of Bioinformatics, [email protected] Project Highlights: Cutting-edge multi-omics approach Microbial physiology linked to crop production Scaling from the laboratory to the field Overview: This project focuses on bacterial extracellular (exo) enzymes involved in the remineralisation and solubilsation of complex organic phosphates and insoluble inorganic phosphates. These processes are thought to be involved in soil fertility and thus provide agricultral crops with inorganic phosphate (Pi) required for growth. Since the solubility of Pi salts is poor, and phopshorus (P) present in organic forms (Po) is not directly available for uptake by the roots, the supply of Pi in many soils is insufficient to maintain plant growth. Whilst bacteria have long been exploited for their enzymes there is a critical need to consider all the pathways for Pi mobilisation in soil in order to manipulate the plant microbiome to reduce our reliance on rock phopshate (Pi), which is a finite commodity. The alkaline and acid phosphatases, phytases, nucleases and phosphonatases are regarded as key microbial enzymes involved in microbial phosphate mineralisation. Recent work in our lab using Warwicks cutting-edge proteomics facility revealed a number of exoenzymes secreted from the cell in response to Pi-depletion in various rhizobacterial strains (RBS) from the genus Pseudomonas and Flavobacteria (Lidbury et al. 2016, Fig. 1). We now aim to take a novel approach to study in situ the enzymes involved in Pi mobilization by using metaproteomics and applying our newly developed method of metaexoproteomics (Johnson- Rollings et al., 2014). Our aim is to focus on the extracellular proteins in soil, as enzymes involved in breakdown of insoluble polymers and organic complexes must be in the extracellular milieu in order to act on these substrates. Wilmes and Bond [3] pioneered protein extraction from environmental samples. The soil metaexoproteome (SMEP) alongside the soil metaproteome (SMP) will indicate enzymes involved in Pi mobilisation and combined with metagenomics will provide insights into the microbial phosphatome in soil. 50% 45% 40% 35% 30% 25% 20% 15% 10% Low Pi 5% High Pi 0% DSM4166 - replete DSM4166 - deplete Figure 1: Analysis of the exoproteome of Pseudomonas stutzeri DSM4166. Proteins in the exoprtoeome were semi-quantified Left panel from mass-spectroscopy data generated from the protein gel right panel. Methodology: We will use various RBS as inoculants in soil experiments to determine the validity of our protein extraction process. and also to asceretain whether or not the same exoproteins are secreted in response to low Pi in situ as in vitro. We will first use laboratory control pot experiments before moving into fields plots. Warwick is currently running several field trials where wheat and oil seed rape (OSR) have been grown in differing rotations. The metagenome (MG) and metatranscriptome (MT) of the soil +/- Pi will also be obtained from deep sequence analysis of total community DNA/RNA and this data will be used to inform analysis of the soil SMP and SMEP. Objectives 1. Determine the phosphatome of various rhizobacterial strains in situ 2. Use soil microcosms to determine the key proteins responsible for Pi mobilisation in the rhizopshere of OSR and wheat in different soils. 3. Determine the key proteins responsible Pi mobilisation in field plots. Training and skills: CENTA students will attend 45 days training throughout their PhD including a 10-day placement. In the first year, students will be trained as a single cohort on environmental science, research methods and core skills. Throughout the PhD, training will progress from core skills sets to master classes specific to the student's projects and themes. Extensive training in experimental techniques related to molecular analysis of environmental samples will be provided in our lab by a senior research technician and post-doctoral research assistant working on a project similar to this. This includes developing skills in the field of bioinformatics, which has become a staple tool in microbial ecology. This project will provide a unique opportunity to work with skilled bioninformaticians at the European Bionformatics Institute (EBI), where a training course is available. The student will join a metagenomics network (ComMet) and gain access to training workshops and meetings in the UK. Partners and collaboration (including CASE): The experimental expertise in Wellington lab will be complimented by detailed expert knowledge provided by Jones, the director of Warwick Proteomics Centre where protein extracts will be analysed. Current work with Jones aims to improve resolution of peptides using a gel-independent Thermo Scientific™ Orbitrap Fusion Tribrid mass spectrometer. Bioinformatics expertise offered by Rob Finn will assist in resolving the protein identities from peptide hits using metagenome data bases. Data bases of enzymes predicted to be in the phosphatome are currently being developed by Finn and Wellington using in vitro data derived from published studies and genomic data bases. Possible timeline: Year 1: Use various RBS to generate data on the in situ expression of Pi responsive exoenzymes. Year 2: Work with plant-soil systems to analyse rhizosphere MG in response to Pi using HiSeq and use EBI portal to resolve gene diversity. Use this database to assist in resolving SMP and SEMP developed from protein extracts of rhizosphere. Year 3: Field trial experiments on wheat and OSR using the same methods developed in years 1 and 2. Further reading: 1) Lidbury IDEA, Murphy ARJ, Scanlan DJ, Bending GD, Jones AME, Moore JD et al (2016). Comparative genomic, proteomic and exoproteomic analyses of three Pseudomonas strains reveals novel insights into the phosphorus scavenging capabilities of soil bacteria. Environ Microbiol doi:10.1111/14622920.13390. 2) Roca A, Pizarro-Tobías P, Udaondo Z, Fernández M, Matilla MA, Molina-Henares MA, Molina L, Segura A, Duque E, Ramos JL. (2013). Analysis of the plant growth-promoting properties encoded by the genome of the rhizobacterium Pseudomonas putida BIRD-1. Environ Microbiol. 15, 780-94. 3) Johnson-Rollings, A. S., Wright, H., Masciandaro, G., Macci, C., Doni, S., Calvo-Bado, L. A., Slade, S. E., Vallin Plou, C., Wellington, E. M. H. (2014). Exploring the functional soil-microbe interface and exoenzymes through soil metaexoproteomics. ISME J. 8, 2148-50. 4) Wilmes, P.; Bond, P. L., (2006). Metaproteomics: studying functional gene expression in microbial ecosystems. Trends Microbiol, 14, 92-97. Further details: Professor E M H Wellington School of Life Sciences The University of Warwick Coventry CV4 7AL United Kingdom Tel: 00442476 523184 Fax: 00442476 523701 Email: [email protected] http://www2.warwick.ac.uk/fac/sci/lifesci/people/ew ellington/