* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Key Concepts -- Lecture 17 (BIOSYSTEMATICS 2) Spring 2009 IB
Population genetics wikipedia , lookup
Genome evolution wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Gene expression programming wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome (book) wikipedia , lookup
Y chromosome wikipedia , lookup
X-inactivation wikipedia , lookup
Human–animal hybrid wikipedia , lookup
Neocentromere wikipedia , lookup
Koinophilia wikipedia , lookup
Microevolution wikipedia , lookup
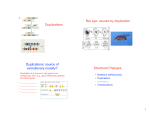
Key Concepts -- Lecture 17 (BIOSYSTEMATICS 2) Spring 2009 IB 168 Cytogenetics (chromosomal genetics). Differences in chromosome number or arrangements of chromosomal arms or segments are common in plants. (DNA was not known to be the genetic material until 1953, whereas chromosomes were associated with inheritance in 1903, ± simultaneous with rediscovery of Mendel's genetic work) Crosses are widely used in cytogenetic studies, although are not always necessary. Major phenomena involving changes in chromosome numbers: - Polyploidy: Addition of one or more entire set(s) of chromosomes. Common phenomenon in flowering plants, ferns, and lycophytes (not known in cycads; rare in conifers; occurs in gnetophytes), both in past and today. - Allopolyploidy believed to be especially common in plants: Chromosome doubling following hybridization (as opposed to chromosome doubling not involving hybridization = autopolyploidy, which also occurs in plants). Results in instantaneous evolution of a reproductively isolated lineage and a good example of reticulate evolution. High incidence of polyploidy may be in part due to selection for fixed hybrid genotypes (allopolyploids will breed true for the hybrid phenotype because of lack of pairing between chromosomes inherited from the different parental species). hybridization Species 1 haploid gamete Species 2 haploid gamete meiosis I fails Hybrid diploid sporophyte Hybrid diploid gametophyte selfing Hybrid diploid gametes True-breeding tetraploid sporophyte Hybrid diploid gametes Tetraploid sporophyte Diploid sporophyte Species 1 ANALYSIS OF GENOMIC SIMILARITY BETWEEN TETRAPLOID AND DIPLOID PLANT EQUAL NUMBER OF PAIRED AND UNPAIRED CHROMOSOMES IN TETRAPLOID X DIPLOID HYBRID IS CONSISTENT WITH THE TWO SPECIES SHARING ONE GENOME (AND THE OTHER GENOME CONTRIBUTED FROM ANOTHER TAXON) meiosis hybridization Diploid gametophyte Haploid gametophyte 1 chr. pair (bivalent) and 1 unpaired (univalent) chr. Dysploidy: change in chromosome number by less than a whole set of chromosomes (gains or losses), often by chromosomal rearrangements (types of macro-mutation) and loss or gain of a chromosome without significant loss or gain of gene dosage (= dysploidy; see example on next page). Chromosome rearrangements involve two simultaneous structural mutations (breakages and reunions) and therefore are highly unlikely to arise independently in different lineages; that is, are unlikely to exhibit homoplasy (as phylogenetic studies of such mutations has confirmed). Consequently, chromosome rearrangements (such as inversions, translocations, etc.) are excellent phylogenetic characters and have been used to infer relationships at deep and shallow levels of divergence. Also, chromosome rearrangements result in loss of interfertility between plants bearing the ancestral versus descendant arrangements (~50% loss of fertility with each reciprocal translocation, for example). loss of centric fragment Unequal reciprocal translocation of chromosome arms (macromutation) centric fragment New arrangement and reduction in chr. number by 1 without loss of genetic material Plant with 2 pairs of chr. (gametophyte) hybridization Taxon with derived arrangement (gametophyte) Taxon with ancestral arrangement (gametophyte) At meiosis, a chain of 3 chromosomes: diagnostic for homology between main arms of 2 small chr. and each of arm of large chr. Breeding relationships: Early biosystematists believed that presence or absence of crossability (= ability to generate a viable, but not necessarily fertile, hybrid) and hybrid vigor, and levels of interfertility (= fertility of hybrids) were good indications of relationship. For example, two species that cannot hybridize were considered more distantly related to one another than two species that could hybridize; also, two species that could form a highly fertile hybrid were considered more closely related than two species that formed sterile hybrid. These beliefs were based on the assumption that evolution is gradualistic, with progressive divergence between lineages manifested by gradually increasing reproductive isolation. These progressive stages, determined by crossability/interfertility experiments, formed the basis of classification. See Clausen (1951), figure 76 in handout for example of how crossing data influenced perception of relationships in Madia and Layia (the figure is essentially a couple of crossing diagrams laid on their sides and with crossing connections pulled down as evolutionary branches). Problems: (1) Exceptions numerous and of all possible types, even in the groups that formed the basis for these ideas. For example, internal reproductive isolation (loss of interfertility or crossability) arises well after ecological isolation and evolutionary divergence in many groups of woody plants (oaks, Ceanothus -- see Nobs 1963 figure on handout, Ribes, Pinus, etc.). Reproductive isolation by multiple chromosomal rearrangements may arise before morphological or ecological divergence in some annual plants (for example, in some members of Holocarpha -- see Clausen 1951 fig. 42 on handout). In short, internal barriers to gene flow are not necessary prior to evolutionary divergence in plants and the presence of internal barriers to gene flow between two groups will not necessarily result in their rapid divergence in morphological and/or ecological features (stabilizing selection or evolutionary constraints may cause them to remain similar, as noted in the speciation lecture by Brent Mishler). (2) Crossability and interfertility are plesiomorphic (ancestral) features and therefore cannot reliably diagnose clades or lineages; barriers to gene flow can arise rapidly and at irregular rates in different groups, so non-monophyletic groups are likely to be recognized using those criteria as the basis for classification. (3) Circularity in reasoning if you base classification on crossability/interfertility and then make conclusions about how speciation/diversification has occurred based on breeding data. Note: In general, perennials (including woody plants) and annuals appear to differ in the timing of origin of reproductive barriers, with different perennial lineages often retaining interfertility long after divergence and different annual lineages often becoming intersterile or losing crossability before much morphological or ecological change has occurred. This difference between annuals and perennials may reflect higher overall rates of molecular evolution in annuals (recently demonstrated). In part, more rapid evolution of reproductive isolation in annuals (at least with regard to pre-zygotic barriers) could be attributable to different levels of natural selection acting against gametic wastage. That is, annuals have one reproductive opportunity before dying; if they hybridize, they may leave behind only unfit progeny. Perennials, on the other hand, have multiple reproductive opportunities over a long timeframe and may not experience such strong selection against the ability to hybridize; if seed set is not the limiting factor on reproductive success, then producing occasional hybrids may not be deleterious. Bias: The focus on making hybrids in biosystematic methods may account for the widespread assumption among biosystematists that if hybrids could be made, then gene flow or reticulate evolution between those crossable taxa would eventually occur in nature. That prediction formed the basis for defining genera as groups of species among which reticulate evolution was possible. Reticulation may be possible, but we cannot predict the future and, again, crossability may not diagnose an actual lineage. Chemosystematics - Systematic study of variation in secondary metabolites (often defensive compounds sequestered in plants -- highly variable, even among close relatives). First became popular as a means of distinguishing hybrids (look for additive profile of compounds that combine the two parental profiles, by paper chromatography, HPLC, etc.). Broad array of aromatic compounds have been studied, including terpenes, flavonoids, and alkaloids. These methods were not in force until the early 1960s and have gradually waned as studies revealed extensive homoplasy in secondary compounds (not good phylogenetic characters at any level of divergence). Protein (enzyme) electrophoresis -- A set of methods involving electrophoresis (separating molecules of unlike charge, or of similar charge but different size, in an electric field, in some type of matrix) to examine the different allelic forms of enzymes, or duplicated versus unduplicated enzymes. Used in part by systematists to diagnose hybrids, as with chemosystematic data; used more widely by population geneticists/biologists to examine variation within and between populations.