* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Intestinal microbiota and metabolites—Implications for broiler
Survey
Document related concepts
Traveler's diarrhea wikipedia , lookup
Horizontal gene transfer wikipedia , lookup
Transmission (medicine) wikipedia , lookup
Trimeric autotransporter adhesin wikipedia , lookup
Microorganism wikipedia , lookup
Disinfectant wikipedia , lookup
Metagenomics wikipedia , lookup
Community fingerprinting wikipedia , lookup
Magnetotactic bacteria wikipedia , lookup
Bacterial cell structure wikipedia , lookup
Marine microorganism wikipedia , lookup
Probiotics in children wikipedia , lookup
Triclocarban wikipedia , lookup
Phospholipid-derived fatty acids wikipedia , lookup
Transcript
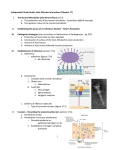
©2013 Poultry Science Association, Inc. Intestinal microbiota and metabolites—Implications for broiler chicken health and performance1 T. Rinttilä and J. Apajalahti2 Alimetrics Ltd., Koskelontie 19B, FIN-02920 Espoo, Finland Primary Audience: Researchers, Nutritionists, Veterinarians SUMMARY Microbiota in the intestine of an animal species has evolved together with the host. This coevolution has produced specific host-microbe combinations, called superorganisms, with the best possible fitness in a given environment. Intestinal microbiota has an enormous metabolic potential and it affects both the nutrition and health of the host. The importance of intestinal microbiota for the performance of broiler chickens has been studied for decades. The microbial community structure of the cecum is significantly more complex and less well characterized than that of the crop and small intestine. Culture-independent molecular techniques have recently been introduced to bring fresh insights into the intestinal system of the broiler chicken. As a consequence, there is growing evidence of connection between the apparent metabolizable energy of the diet and microbiota composition in the hindgut of the host. This connection can be due to direct conversion of some dietary components into high-energy metabolites by specialized bacteria. The utilization of such fermentation end products would provide more energy for the host, which could be measured as improved feed conversion efficiency. However, it is also possible that cecal microbial profile is a reflection of feed digestion and nutrient absorption efficiency in the proximal intestine. Disorders in these functions cause excess nutrient bypass to the lower intestine, providing new, easily available substrates for bacteria that could otherwise not compete in that habitat. The 2 causal links described are likely to coexist in real life situations, but their relative dominance warrants further well-designed studies. Key words: metabolite, intestinal microbiota, broiler chicken health, broiler chicken performance 2013 J. Appl. Poult. Res. 22:647–658 http://dx.doi.org/10.3382/japr.2013-00742 INTRODUCTION Microbiota in the animal intestine has evolved together with the host. As a consequence, the gastrointestinal tract (GIT) could be considered a metacommunity, comprising many local microbial communities in different ecological niches. Each individual gut compartment has its 1 unique physiochemical characteristics and each location is inhabited by a specialized microbial composition [1]. The microorganisms inhabiting the intestinal tract of humans and animals outnumber the cells of a host [2]. The vast majority of gut bacteria reside in the distal intestine, where densities approach 1011 to 1012 cells/g, the highest recorded for any microbial habitat [3]. Presented as a part of the Informal Nutrition Symposium “Metabolic Responses to Nutrition and Modifiers” at the Poultry Science Association’s annual meeting in Athens, Georgia, July 9, 2012. 2 Corresponding author: [email protected] 648 The gut microbiota therefore extends the host’s genome such that the host can be described as host-microbe superorganism [4]. One of the preconditions for a long-term coadaptation of the gut microbiota is the uninterrupted transfer of maternal intestinal bacteria to the next generation. If this lineage breaks, the superorganism ceases to exist and the adapted microbiota is replaced by a random selection of bacteria from the environment that are able to colonize the GI tract. It is worth considering whether such natural inoculation is achieved in today’s commercial hatchery systems and whether the characteristics evolved are desirable in animal production. For example, microbiota beneficial for the broiler breeders may not be ideal for the health and performance of shortlived broiler chickens. Independent studies carried out with germfree mice and human volunteers have elucidated that obesity is likely to be associated with the capability of the colon microbiota to harvest energy from the diet [5]. Knowledge on the intestinal microbiota of man is mainly motivated by health applications, one sideline of which is obesity. However, intestinal health is also highly important for production animals. When productivity is not threatened by disease, the most important driver of performance is the capability of the animal to convert feed into carcass as efficiently as possible. Therefore, the question arises as to whether the microbiota associated with obesity in humans could provide us with information of how to improve energy harvesting and FCR in production animals such as broiler chickens. Humans have a highly variable genetic background, a unique microbiota inherited from the mother, and variable dietary preferences. Thus, finding significant associations between the structure of intestinal microbiota and various phenotypic traits is likely to be more challenging than finding such associations in flocks of broiler chickens with a relatively uniform genetic background, diet, and bacterial pressure from the environment. In spite of this apparently easy experimental setting, there is no consensus regarding the type of microbiota related to an ideal animal performance. Until recently, intestinal microbiota was studied using conventional culturing techniques, which apply differential media to select for spe- JAPR: Symposium cific populations of bacteria based on their metabolic requirements. Although the culture-based techniques enable the recovery of microbial isolates for further biochemical and physiological investigations under both anaerobic and aerobic conditions, only a proportion (10–60%) of the bacterial species present in the intestine are estimated to be cultivable under standard laboratory conditions [6–9]. To overcome these drawbacks, culture-independent molecular analysis tecniques have been introduced to obtain a better understanding of the gut microbiota. However, in studies involving microbiota analysis, the complexity of interpretation increases with the use of applied methods, all of which have inherent biases of varying degrees. Hence, the results of independent studies with different sample preparation and analysis techniques need to be interpreted with caution. INTESTINAL MICROBIOTA Establishment of the Intestinal Microbiota Development of the broiler chicken intestinal microbiota starts at the moment of hatching. The newly hatched chicks are initially exposed to microbes that originate from the surface of the egg shells, which are populated by bacteria from the intestine of mother and the surrounding environment. Hence, the microbial inoculum obtained at the early stage of the posthatch period is critical for the establishment of the gut microbial community. The effect of the first inoculum may last over the entire lifetime of a broiler chicken by directing the development of the immune system and the intestinal microbiota [10]. Bacterial densities in the chicken GIT have been shown to increase rapidly after hatching. In a study by Apajalahti et al. [10], bacterial numbers in the proximal and distal intestine of the broiler chicks reached 108 and 1010 cells/g of digesta 1 d posthatching, respectively. The maximum density of 109 cells/g of ileal digesta and 1011 per gram of cecal digesta was reached less than 1 wk, after which the microbiota remained relatively stable for the following 30 d. The GIT is initially colonized by facultative aerobes such as Enterobacteriaceae, Lactobacillus, and Streptococcus, as the intestinal environment of chicks shows a positive oxidation or reduction potential at hatching [11–14]. However, Rinttilä and Apajalahti: INFORMAL NUTRITION SYMPOSIUM oxygen consumption by these bacteria alters the lower gut environment to more reducing conditions, which facilitates subsequent growth and colonization of the extremely oxygen-sensitive obligate anaerobes [11, 13, 15]. Microbial Community of the Upper GIT The small intestine, which consists of the duodenum, jejunum, and ileum, is the GIT compartment where most of the digestion and absorption of nutrients occurs [16]. Bacteria in the small intestine use the same readily fermentable nutrients that are being used by the host. Therefore, the small intestine is a gut compartment where commensal microbiota and the host compete for dietary nutrients [17]. The host can recover part of the energy lost to microbes by absorbing and metabolizing bacterial fermentation products, lactic and volatile fatty acids (VFA). Due to the low pH and rapid passage of intestinal contents, duodenal bacterial counts are low. Subsequently, digesta flows from the duodenum through the jejunum to the end of the ileum, where the digestive enzyme activities are reduced and bile acids are deconjugated [18]. As a consequence, the bacterial numbers increase up to 108 cells/mL of digesta in the distal ileum [10]. With methods based on sequencing of specific DNA fragments, there is relatively good consensus of the microbial community structure of the in the chicken small intestine [12, 19–24]. By far the most dominant bacteria are lactobacilli, which represent 80 to 90% of the total microbiota, the remainder mainly consisting of enterobacteria and enterococci. The most dominant cecal bacteria are also found in the distal ileum. It is possible that these bacteria enter the ileum during cecal emptying or reverse peristalsis and are not metabolically active in the small intestine. Microbial Community of the Lower Intestine Due to the low redox potential prevailing distal intestine, the numbers of obligate anaerobic microbes are several orders of magnitude higher than those of aerobes or facultative anaerobes [25]. Dietary compounds and intestinal secretions that are not absorbed in the small intestine reach the lower intestine, where they provide the 649 dense microbial population with nutrients and energy for growth [26]. Hence, intense bacterial fermentation in the hindgut does not lead to energy losses for the host, as these bacteria scavenge energy and nutrients from feed residues that are already beyond the endogenous digestion system of the host. Accordingly, any bacterial metabolites produced from such residues that have any value for the host will improve the yield of overall energy and nutrient capture from the diet. It is possible that with poorly digestible diets, a considerable proportion of total dietary energy comes from bacterial fermentation. In recent years, the application of 16S ribosomal (r) RNA gene sequencing approaches has provided new insights into characterization of the broiler chicken cecal microbial composition [21, 23, 27]. Several studies have revealed that cecal microbiota in healthy chickens is dominated by representatives of the order Clostridiales. For a long time, representatives of this order were grouped according to their 16S rDNA sequence homology to Clostridial clusters I to XIX. During the last few years, several bacterial phylotypes that previously could not be assigned to proper taxonomic clusters have now found a place in newly described families and genera. In the chicken cecum, the most common families within the order Clostridiales are Lachnospiraceae, Ruminococcaceae, and Veillonellaceae [27]. These appear to represent more than 50% of the cecal bacteria, the remainder belonging mainly to the orders Bacteroidales and Lactobacillales [20, 28]. The genus Clostridium is well known to people involved in the poultry industry, mainly because of one pathogenic species, Clostridium perfringens. Unfortunately, this species has affected the reputation of the entire order Clostridiales, even though the genus Clostridium is only 1 of 123 genera within the order and the species C. perfringens is only distantly related to the majority of the 129 species in the genus Clostridium. Therefore, the generic term clostridia should not be automatically associated with poor health and performance in broiler chickens. In fact, all the existing evidence suggests that the great majority of the bacteria belonging to the bacterial class Clostridia are nonpathogenic, and even beneficial for animals and for farm profits. 650 Functions of the Gut Microbiota The conventional view that all microorganisms cause disease has been replaced by the insight that most microbe-host interactions are commensal (with the microbes benefiting while the host is unharmed) or even mutualistic (in which both partners benefit) [1]. The benefits of commensal intestinal bacteria are well known: they help in digestion and synthesis of dietary compounds; are involved in the gastrointestinal development; regulate intestinal epithelial proliferation, inflammatory immune responses, and host energy metabolism; synthesize vitamins; and fill a microbiological niche that might otherwise be colonized by potentially harmful enteric microorganisms [29–33]. In return, the intestinal microorganisms are provided with secure growth conditions and a constant stream of nutrients [2]. The diet-derived complex carbohydrates not degraded in the small intestine, such as nonstarch polysaccharides (NSP) and resistant starch, are the main sources of carbon and energy for the commensal microbiota in the lower intestine. Dietary protein and protein from pancreatic enzymes and gastrointestinal secretions also contribute, to some extent, to the supply of substrates to intestinal bacteria [26]. In addition, the gut microbiota degrades and processes large quantities of substances derived from the host, such as mucins and sloughed epithelial cells. Hence, the intestinal microbiota has a substantial catalytic potential, which leads to the formation of microbial metabolites with beneficial or adverse health effects. The amounts and types of fermentation products formed by intestinal bacteria depend on the relative amounts of each substrate available and the fermentation strategy (biochemical characteristics and catabolite regulatory mechanisms) of bacteria involved in the fermentation process [34]. Importance of Short-Chain Fatty Acids The principal end products of intestinal microbial fermentation are short-chain fatty acids (SCFA), such as acetate, butyrate, propionate, succinate, and lactate [21, 35, 36]. Human studies have shown that up to 95% of the SCFA produced during carbohydrate fermentation are JAPR: Symposium used by the host, providing 5 to 15% of the total energy requirements; whereas pigs may obtain as much as 30% of their energy from SCFAproducing hindgut fermentation [37, 38]. Ruminants obtain almost all their energy from SCFA produced in the rumen. In addition to energyyielding activity, SCFA formation in chicken cecum reduces pH of the intestinal environment, which may inhibit acid-sensitive pathogenic bacteria, such as members of the family Enterobacteriaceae, by dissipating the proton motive force across the bacterial cell membrane [39]. Among the SCFA, butyrate has gained particular interest, as it is the preferred energy source for the enterocytes and is known to regulate cellular differentiation and proliferation within the intestinal mucosa, thereby increasing intestinal tissue weight [40, 41]. The epithelium provides a highly selective barrier that prevents the passage of toxic and proinflammatory molecules from the external milieu into the submucosa and systemic circulation [42]. Hence, the contribution of butyrate and other SCFA to epithelial development is essential in the maintenance of normal intestinal barrier functions. Although butyrate production is distributed across many Clostridial clusters of the phylum Firmicutes [43], it is mainly produced by members of Roseburia spp. and Eubacterium rectale, both members of the family Lachnospiraceae, as well as by Faecalibacterium prausnitzii-like organisms of family Ruminococcaceae [21, 44, 45]. These bacterial groups are susceptible to changes in dietary carbohydrate intake, which has an effect on lower intestine butyrate concentrations [46]. Lactic acid is the strongest of the common SCFA produced by GIT bacteria, and therefore it tends to reduce residual pH more than other SCFA. Lactate produced by lactic acid bacteria is rarely present in the hindgut in high quantities, as it is normally rapidly absorbed from the intestine or used as a substrate for lactate-utilizing bacteria, such as representatives of the genera Eubacterium, Anaerostipes, Veillonella, and Megasphaera [47, 48]. However, human studies have shown that, in certain conditions, abnormal accumulation of lactic acid occurs as a result of carbohydrate malabsorption in the upper intestine and subsequent overgrowth of lactic acidproducing bacteria due to increased delivery of readily metabolized carbohydrates to the lower Rinttilä and Apajalahti: INFORMAL NUTRITION SYMPOSIUM intestine [49]. Consequently, production of lactic acid exceeds the buffering and absorptive capacity of the colon and the pH of the habitat decreases, with concomitant inhibition of the activity of lactate-utilizing microorganisms. This cascade of events may eventually lead to impaired well-being of the host. There is only a little evidence of the lactic acid accumulation in chicken cecum. Taylor [50] discovered elevated d-lactic acid concentrations in the hindgut of the laying hens caused by sudden changes in diet type or processing. Protein and Amino Acid Fermentation in the Lower Intestine The lower intestine of warm-blooded animals has been described as a site of intense protein turnover [51]. The active proteolytic microbial community in the hindgut includes species belonging to the genera Bacteroides, Clostridium, Fusobacterium, Streptococcus, and Enterococcus [52]. Once carbohydrate sources are exhausted in the broiler chicken cecum, sources of protein material are fermented and metabolized to salvage energy. Although proteins and amino acids provide a less significant energy source in the lower intestine, their importance lies mainly in the formation of potential systemic toxins and carcinogens as a result of protein or amino acid fermentation by putrefactive bacteria. Common examples of undesired metabolic end products include phenols and indoles (as a result of anaerobic fermentation of the aromatic amino acids, tyrosine and tryptophan, by intestinal bacteria), ammonia (as a result of oxidative or reductive deamination of amino acids), and amines (as a result of amino acid decarboxylation in the gut). Ammonia is easily absorbed through the intestinal epithelium into the blood circulation of broiler chickens and has toxic effects on enterocytes, whereas amines have been associated in several physiological processes in living cells [53, 54]. In addition to being toxic to the host, these end products of protein fermentation tend to increase the pH of the intestinal contents. The general assumption is that low pH is beneficial for gut health, as the growth of the acid-sensitive pathogenic microorganisms is suppressed in such conditions [17]. As the bacteria gener- 651 ally favor fermentable sources of carbohydrates, protective measures against excessive putrefactive activity and the undesired metabolites in the hindgut can be achieved by adding dietary fiber in the diet or limiting the intake of poorly digested proteins. Role of Intestinal Microbiota Against Enteric Pathogens The mucosal surface of the broiler chicken GIT is the largest body surface in contact with the external environment. It is a complex ecosystem combining the gastrointestinal epithelium, immune cells, and resident microbiota [55]. The host is normally protected by the physical barrier created by colonized commensal microbes, which protects the epithelium from attack by invading harmful microorganisms or opportunistic species usually present in the gut in low numbers. However, the intestinal mucosa can be exposed to various enteric pathogens. In the first step of the infection process, pathogenic microorganisms adhere to the brush border of intestinal cells, enabling them to exploit the underlying signaling pathways [56, 57]. Furthermore, some enteric microbial pathogens have developed specialized systems to produce virulence factors. After normal host-cell processes have been weakened, these systems enable the pathogen to penetrate and cross the epithelial barrier [58]. The enteric microbial pathogens commonly target the host cell cytoskeleton during the cell penetration step. It is exploited for various purposes, such as gaining entry into cells, moving within and between cells, and forming vacuoles to create a specialized niche, which facilitates the pathogen’s chances of survival and ability to proliferate [59]. The role of the commensal GIT microbiota in suppressing pathogen colonization is likely to be multifactorial. In addition to the production of a wide variety SCFA, which are bacteriostatic for a subset of bacterial species either directly or by reducing pH of the intestinal environment, the commensal microbiota contribute to the colonization resistance against pathogenic microbes. Moreover, some members of the microbiota also generate bacteriocins, small peptide molecules with microbicidal or microbiostatic properties [60]. In addition, intestinal microbes apply a 652 delicate microbe-to-microbe signaling network against pathogenic invaders, as well as using it to optimize the composition and numbers of appropriate members of the microbial ecosystem. This coordinated bacterial communication, termed quorum sensing, can occur within a single bacterial species or between diverse species, and enables the regulation of different processes, especially when environmental conditions are appropriate [61]. A variety of different molecules can be used as signals. Common classes of soluble signaling molecules include oligopeptides in gram-positive bacteria, N-acyl homoserine lactones in gram-negative bacteria, and a family of autoinducers known as autoinducer-2 in both gram-negative and positive bacteria [62]. Cecal Microbial Community and Metabolizability of Dietary Energy The composition of the broiler chicken microbial community is likely to be affected by the source of the microbes that first meet the virgin intestine by the diet. If hatching takes place in an environment where the microbial load has been minimized, individual chicks may pick up a random inoculum from their surroundings, which may lead to differences in the intestinal physiology of individual birds in a flock. It is worth noting that intentional inoculation of the chicks with a competitive exclusion culture at hatch might render a flock more uniform, not only with regard to microbiota composition, but also to FCR. JAPR: Symposium In our studies on microbiota of the chicken intestine over the years, it has become apparent that on farms with good health and FCR history, the between-bird variability of the dominant microbial community is small, whereas on problem farms the flock is less uniform. Figure 1 clearly illustrates this. Ten commercial Finnish farms were randomly sampled for microbiota analysis 3 wk after arrival of the chicks. Three pools of 3 birds per farm were analyzed for cecal microbiota structure by a PCR-independent DNA-based method that fractionates total bacterial chromosomes according to their guanine + cytosine content (G+C). Three weeks later, the flock was taken to slaughter and carcass weights were recorded for each farm. Final FCR was calculated and compared against the G+C profiles. The results clearly demonstrated that, on farms with better overall FCR, the variability in the cecal microbiota composition 3 wk before slaughter was much lower than on farms with poorer FCR. It is intriguing to think that on any farm there may be individual birds that have highly different efficiency to convert diet into body mass. The flock may appear uniform in size, but some animals may consume much more feed than others. The farm- and flock-specific situation can be revealed by determining a variability index by G+C profiling. In a study carried out by McCracken at Queen’s University (Belfast, Northern Ireland), broiler chickens were individually caged, fed with the same wheat-based diet, and analyzed Figure 1. Chicken cecal microbial profiles arranged according to the FCR of the farm from which they originated. Bacterial profiles from 5 farms with the lowest FCR are on the left and those from 5 farms with the highest FCR on the right. G+C = guanine + cytosine. Rinttilä and Apajalahti: INFORMAL NUTRITION SYMPOSIUM for metabolizable energy from 7 to 28 d. Cecal bacterial profile was determined by G+C profiling and compared against the ability of the individual chicken to harvest energy from the diet. It was found that metabolizable or gross energy was significantly different between individual birds, ranging from less than 60 to more than 80%, and there was no good correlation between FCR and BW gain [63]. Figure 2 shows the microbial profiles of the 8 most efficient birds in terms of energy harvesting and the 8 least efficient birds. There was a highly significant correlation between cecal microbial community structure and the energy harvesting efficiency of the host. Cecal bacteria from the most efficient birds were dominated by bacteria with G+C content around 45%. In inefficient birds, this peak was replaced by 2 major peaks, representing bacteria with G+C content around 37 and above 60%, respectively. These 2 totally independent studies demonstrated that cecal bacterial composition was similarly related to FCR on commercial farms in Finland and to metabolizable energy at the experimental facility in Northern Ireland. In both cases bacteria with G+C ~45% represented birds with good performance, whereas an increased proportion of bacteria with G+C ~37% and ~65% represented broiler chickens with reduced efficiency to harvest energy from the diet and convert feed into carcass mass. Chromosomal DNA representing the diagnostics peaks of the G+C profiles was collected and analyzed in more detail by 16S rRNA gene sequencing to identify the actual bacterial phy- 653 lotypes behind the shifts in the G+C profiles. Eight taxonomic units were identified and a rapid quantitative real-time PCR assays were designed for their quantification. The abundance of these bacterial groups was used to design an algorithm that correlated with performance. The algorithm provided a value referred to as Performance-linked Microbial Index (PMI), which was first applied to all samples from the 2 studies. In Figure 3, the PMI values are plotted against FCR and ME/gross energy. As expected, FCR and PMI showed a highly significant negative correlation, whereas ME and PMI showed a positive correlation. The association of broiler chicken gut microbiota and bird performance as measured by FE was also investigated in a study by Torok et al. [27]. Three feeding trials were performed in various geographic locations with differences in diet composition, broiler breed, and bird age. The results of the gut microbial profiling with terminal restriction fragment length polymorphism technique revealed 8 common performance-linked operational taxonomic units within both the ilea and ceca across the trials. Targeted 16S rRNA gene cloning and sequencing of these operational taxonomic units revealed that they represented 26 bacterial species or phylotypes, which clustered phylogenetically into 7 groups related to Lactobacillus spp., Ruminococcaceae, Clostridiales, Gammaproteobacteria, Bacteroidales, Clostridiales, or Lachnospiraceae, and unclassified bacteria or clostridia. Some of the potential performance-related phylotypes showed high sequence identity with classified bacteria Figure 2. Chicken cecal microbial profiles of individually caged birds arranged according to diet metabolizability. The profiles on the left are from the 8 birds with the highest metabolizability and those on the right from the 8 birds with the lowest metabolizability. The black line in each diagram represents the mean of the 8 profiles. GE = gross energy; G+C = guanine + cytosine. 654 JAPR: Symposium Figure 3. Relationship between performance-linked microbial index and feed utilization efficiency expressed as FCR (upper panel) and ME (lower panel). GE = gross energy. such as Lactobacillus salivarius, Ruminococcus torques, and Faecalibacterium prausnitzii. It is also noteworthy that birds on diets exhibiting significantly lower performances in this study generally showed more pronounced betweenbird variation in gut microbial communities. As demonstrated by the trials described above, there is a correlation between cecal microbiota composition and the efficiency of the host to extract energy from the diet and to deposit that energy into improved FCR. However, the question of how the intestinal bacterial community composition relate to relevant metabolic changes and to broiler chicken performance is not completely understood. Turnbaugh et al. [64] discovered in a study with obese and lean twins that obese individuals had a larger proportion of genes for digesting fat, protein, and carbohydrates, which possibly facilitates their ability to extract and store energy from food. Furthermore, the same researchers demonstrated that the transfer of microbes from a conventional mouse to a germ-free individual led to different energy-harvesting efficiency depending on whether the inoculum was derived from an obese or a lean mouse, suggesting that specific microbes are able to provide the host with an energy-efficient phenotype [5]. Consequently, it is likely that certain microbial species or groups in the gut microbiome encode substantial specialized metabolic capacities, including the efficient degradation of otherwise indigestible components of the diet with subsequent production of SCFA. As mentioned previously, SCFA are a valuable energy source for the host, with butyrate being especially important as a preferred energy source for the intestinal epithelium, thus ensuring that dietary nutrients and high-energy SCFA are taken up by the host at the best possible efficiency. Accordingly, butyrate-producing Rinttilä and Apajalahti: INFORMAL NUTRITION SYMPOSIUM representatives of the bacterial order Clostridiales can be expected to be especially beneficial for the animal performance. An alternative explanation for the connection between FCR and the composition of cecal microbiota is that digestive disorders or compromised absorption of nutrients in the small intestine results in an abnormal nutrient bypass to the lower intestine. Hence, dietary components, which under normal circumstances would be digested and absorbed in the upper intestine, fail to do so for some reason. This could be a consequence of epithelial damage caused by coccidiosis, necrotic enteritis, or a high level of antinutritional lectins in the diet. It is also possible that an exceptionally high passage rate of digesta leads to poor nutrient absorption and high nutrient bypass [65]. Poor nutrient absorption reduces FCR and, simultaneously, the overflow of nutrients to the lower intestine favors bacteria that grow rapidly on easily available substrates. In such situations, a correlation between FCR and cecal microbiota is apparent but the favored bacteria do not directly affect FCR. Instead, the shift in microbiota composition is a reflection of reduced nutrient absorption. It is most likely that both of the abovementioned causal links exist in real life situations. Even though there are 2 clearly different mechanisms, they are not mutually exclusive and most likely coexist. In the ideal situation, the main dietary nutrients, starch and protein, are effectively digested and absorbed in the small intestine. The only dietary components that reach cecum are NSP, which are hydrolyzed and metabolized by cecal bacteria to high-energy SCFA, with a sufficient proportion of butyrate. In this situation the proportion of bacteria capable of utilizing NSP is high, and butyrate-producing bacteria of the order Clostridiales are well represented. POTENTIAL STRATEGIES TO OPTIMIZE CECAL MICROBIOTA COMPOSITION There is mounting evidence that intestinal microbiota of the broiler chickens is associated with good animal performance. However, if poor bacterial profile cannot be transformed into a good profile, with a concomitant improvement in animal performance, this knowledge has 655 no practical value. If optimization of microbial balance in the cecal environment leads to good performance, it would be of great importance to establish or develop strategies to provide a competitive advantage for representatives of taxa identified as being beneficial or taxa providing similar functions. There are many potential ways of achieving this. One option could be to develop starter cultures that directly provide the most important bacterial species. This is challenging because the desirable beneficial bacteria are strict anaerobes. Such products are commercially available for ruminants (e.g., Megasphaera elsdenii), but significant development work is required for poultry applications. An alternative to early inoculation is to provide dietary ingredients that favor the growth of the desirable bacteria. In principle, it is possible to affect microbial balance by nutritional strategies including an appropriate selection of the main feed raw materials. In addition, various special feed additives, such as prebiotics, feed enzymes, and other products that potentially provide substrates for the NSP-digesting and SCFA-producing cecal bacteria, could serve as efficient microbiota modulators and indirectly inhibit the growth of pathogenic bacteria. In some cases it is possible that cecal microbiota composition does not directly affect FCR, but only reflects feed digestibility, conditions in the proximal intestine, and consequent nutrient bypass to the cecum. In such cases, the potential use of knowledge obtained is different. One approach to use that information could be in diagnostic testing on the farm to determine the intestinal condition and uniformity of the flock. In wider farm surveys, common factors that lead to a lack of between-bird uniformity and ways to improve the situation could be identified. With early diagnosis, it could even be possible to take corrective actions before the birds are slaughtered. CONCLUSIONS AND APPLICATIONS 1. Growing evidence exists of a connection between good animal performance and microbiota composition in the lower GIT of broiler chickens. 2. More well-designed research is warranted to investigate the influence of dietary JAPR: Symposium 656 factors on physiology and metabolism of the microorganisms residing in the intestinal tract, as well as on absorption of fermentation end products. 3. Today, numerous potential in vitro methods are available for prescreening a wide range of dietary ingredients under laboratory conditions for the properties described in this paper. 4. It is essential that the microbial modulators with the most potential would be tested in in vivo broiler chicken feeding trials to elucidate the interactions of the dietary composition and microbial fermentation, especially with regard to animal health and performance. REFERENCES AND NOTES 1. Dethlefsen, L., M. McFall-Ngai, and D. A. Relman. 2007. An ecological and evolutionary perspective on human–microbe mutualism and disease. Nature 449:811–818. 2. Savage, D. C. 1977. Microbial ecology of the gastrointestinal tract. Annu. Rev. Microbiol. 31:107–133. 3. Whitman, W. B., D. C. Coleman, and W. J. Wiebe. 1998. Prokaryotes: The unseen majority. Proc. Natl. Acad. Sci. USA 95:6578–6583. 4. Kleerebezem, M., and E. E. Vaughan. 2009. Probiotic and gut lactobacilli and bifidobacteria: Molecular approaches to study diversity and activity. Annu. Rev. Microbiol. 63:269–290. 5. Turnbaugh, P. J., R. E. Ley, M. A. Mahowald, V. Magrini, E. R. Mardis, and J. I. Gordon. 2006. An obesityassociated gut microbiome with increased capacity for energy harvest. Nature 444:1027–1031. 6. Mead, G. C. 1989. Microbes of the avian cecum: Types present and substrates utilized. J. Exp. Zool. Suppl. 3:48–54. 7. Amann, R. I., W. Ludwig, and K. H. Schleifer. 1995. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 59:143–169. 8. Vaughan, E. E., F. Schut, H. G. Heilig, E. G. Zoetendal, W. M. de Vos, and A. D. Akkermans. 2000. A molecular view of the intestinal ecosystem. Curr. Issues Intest. Microbiol. 1:1–12. 9. Eckburg, P. B., E. M. Bik, C. N. Bernstein, E. Purdom, L. Dethlefsen, M. Sargent, S. R. Gill, K. E. Nelson, and D. A. Relman. 2005. Diversity of the human intestinal microbial flora. Science 308:1635–1638. 10.Apajalahti, J., A. Kettunen, and H. Graham. 2004. Characteristics of the gastrointestinal microbial communities, with special reference to the chicken. World’s Poult. Sci. J. 60:223–232. 11.Barnes, E. M., G. C. Mead, D. A. Barnum, and E. G. Harry. 1972. The intestinal flora of the chicken in the period 2–6 weeks of age, with particular reference to the anaerobic bacteria. Br. Poult. Sci. 13:311–326. 12.Lu, J., U. Idris, B. Harmon, C. Hofacre, J. J. Maurer, and M. D. Lee. 2003. Diversity and succession of the intestinal bacterial community of the maturing broiler chicken. Appl. Environ. Microbiol. 69:6816–6824. 13.Wise, M. G., and G. R. Siragusa. 2007. Quantitative analysis of the intestinal bacterial community in one- to three-week-old commercially reared broiler chickens fed conventional or antibiotic-free vegetable-based diets. J. Appl. Microbiol. 102:1138–1149. 14.Gong, J., H. Yu, T. Liu, J. J. Gill, J. R. Chambers, R. Wheatcroft, and P. M. Sabour. 2008. Effects of zinc bacitracin, bird age and access to range on bacterial microbiota in the ileum and caeca of broiler chickens. J. Appl. Microbiol. 104:1372–1382. 15.Gong, J., R. J. Forster, H. Yu, J. R. Chambers, P. M. Sabour, R. Wheatcroft, and S. Chen. 2002. Diversity and phylogenetic analysis of bacteria in the mucosa of chicken caeca and comparison with bacteria in the caecal lumen. FEMS Microbiol. Lett. 208:1–7. 16.Renner, R. 1965. Site of fat absorption in the chick. Poult. Sci. 44:861–864. 17.Apajalahti, J. 2005. Comparative gut microflora, metabolic challenges, and potential opportunities. J. Appl. Poult. Res. 14:444–453. 18.Rehman, H. U., W. Vahjen, W. A. Awad, and J. Zentek. 2007. Indigenous bacteria and bacterial metabolic products in the gastrointestinal tract of broiler chickens. Arch. Anim. Nutr. 61:319–335. 19.Knarreborg, A., R. M. Engberg, S. K. Jensen, and B. B. Jensen. 2002. Quantitative determination of bile salt hydrolase activity in bacteria isolated from the small intestine of chickens. Appl. Environ. Microbiol. 68:6425–6428. 20.Apajalahti, J., and A. Kettunen. 2006. Microbes of the chicken gastrointestinal tract. Pages 124–137 in Avian Gut Function In Health And Disease. CABI Publishing, Wallingford, UK. 21.Bjerrum, L., R. M. Engberg, T. D. Leser, B. B. Jensen, K. Finster, and K. Pedersen. 2006. Microbial community composition of the ileum and cecum of broiler chickens as revealed by molecular and cellular-based techniques. Poult. Sci. 85:1151–1164. 22.Dumonceaux, T. J., J. E. Hill, S. M. Hemmingsen, and A. G. Van Kessel. 2006. Characterization of intestinal microbiota and response to dietary virginiamycin supplementation in the broiler chicken. Appl. Environ. Microbiol. 72:2815–2823. 23.Gong, J., W. Si, R. J. Forster, R. Huang, H. Yu, Y. Yin, C. Yang, and Y. Han. 2007. 16S rRNA gene-based analysis of mucosa-associated bacterial community and phylogeny in the chicken gastrointestinal tracts: From crops to ceca. FEMS Microbiol. Ecol. 59:147–157. 24.Torok, V. A., K. Ophel-Keller, M. Loo, and R. J. Hughes. 2008. Application of methods for identifying broiler chicken gut bacterial species linked with increased energy metabolism. Appl. Environ. Microbiol. 74:783–791. 25.Marteau, P., P. Pochart, J. Dore, C. Bera-Maillet, A. Bernalier, and G. Corthier. 2001. Comparative study of bacterial groups within the human cecal and fecal microbiota. Appl. Environ. Microbiol. 67:4939–4942. 26.Cummings, J. H., and G. Macfarlane. 1991. The control and consequences of bacterial fermentation in the human colon. J. Appl. Bacteriol. 70:443–459. 27.Torok, V. A., R. J. Hughes, L. L. Mikkelsen, R. Perez-Maldonado, K. Balding, R. MacAlpine, N. J. Percy, and K. Ophel-Keller. 2011. Identification and characterization Rinttilä and Apajalahti: INFORMAL NUTRITION SYMPOSIUM of potential performance-related gut microbiotas in broiler chickens across various feeding trials. Appl. Environ. Microbiol. 77:5868–5878. 28.Zhu, X. Y., T. Zhong, Y. Pandya, and R. D. Joerger. 2002. 16S rRNA-based analysis of microbiota from the cecum of broiler chickens. Appl. Environ. Microbiol. 68:124– 137. 29.Forder, R. E. A., G. S. Howarth, D. R. Tivey, and R. J. Hughes. 2007. Bacterial modulation of small intestinal goblet cells and mucin composition during early posthatch development of poultry. Poult. Sci. 86:2396–2403. 30.Klasing, K. C. 2007. Nutrition and the immune system. Br. Poult. Sci. 48:525–537. 31.Guarner, F. 2006. Enteric flora in health and disease. Digestion 73(Suppl. 1):5–12. 32.Noverr, M. C., and G. B. Huffnagle. 2004. Does the microbiota regulate immune responses outside the gut? Trends Microbiol. 12:562–568. 33.Rakoff-Nahoum, S., J. Paglino, F. Eslami-Varzaneh, S. Edberg, and R. Medzhitov. 2004. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118:229–241. 34.Macfarlane, G. T., and S. Macfarlane. 1997. Human colonic microbiota: Ecology, physiology and metabolic potential of intestinal bacteria. Scand. J. Gastroenterol. Suppl. 222:3–9. 35.Topping, D. L., and P. M. Clifton. 2001. Short-chain fatty acids and human colonic function: Roles of resistant starch and nonstarch polysaccharides. Physiol. Rev. 81:1031–1064. 36.Hooper, L. V., T. Midtvedt, and J. I. Gordon. 2002. How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu. Rev. Nutr. 22:283– 307. 37.Bergman, E. N. 1990. Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol. Rev. 70:567–590. 38.Cummings, J. H., E. W. Pomare, W. J. Branch, C. P. Naylor, and G. T. Macfarlane. 1987. Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28:1221–1227. 39.van Der Wielen, P. W., S. Biesterveld, S. Notermans, H. Hofstra, B. A. P. Urlings, and F. van Knapen. 2000. Role of volatile fatty acids in development of the cecal microflora in broiler chickens during growth. Appl. Environ. Microbiol. 66:2536–2540. 40.Le Blay, G., H. M. Blottiere, L. Ferrier, E. C. Le Foli, J. P. Bonnet, C. Galmiche, and C. Cherbut. 2000. Shortchain fatty acids induce cytoskeletal and extracellular protein modifications associated with modulation of proliferation on primary culture of rat intestinal smooth muscle cells. Dig. Dis. Sci. 45:1623–1630. 41.Fukunaga, T., Y. Sasaki, T. Araki, T. Okamoto, T. Yasuoka, T. Tsujikawa, Y. Fujiyama, and T. Bamba. 2003. Effects of the soluble fibre pectin on intestinal cell proliferation, fecal short chain fatty acid production and microbial population. Digestion 67:42–49. 42.Niba, A. T., J. D. Beal, A. C. Kudi, and P. H. Brooks. 2009. Bacterial fermentation in the gastrointestinal tract of non-ruminants: Influence of fermented feeds and fermentable carbohydrates. Trop. Anim. Health Prod. 41:1393– 1407. 43.Pryde, S. E., S. H. Duncan, G. L. Hold, C. S. Stewart, and H. J. Flint. 2002. The microbiology of butyrate forma- 657 tion in the human colon. FEMS Microbiol. Lett. 217:133– 139. 44.Hold, G. L., A. Schwiertz, R. I. Aminov, M. Blaut, and H. J. Flint. 2003. Oligonucleotide probes that detect quantitatively significant groups of butyrate-producing bacteria in human feces. Appl. Environ. Microbiol. 69:4320– 4324. 45.Louis, P., P. Young, G. Holtrop, and H. J. Flint. 2010. Diversity of human colonic butyrate-producing bacteria revealed by analysis of the butyryl-CoA:acetate CoA-transferase gene. Environ. Microbiol. 12:304–314. 46.Duncan, S. H., P. Louis, and H. J. Flint. 2007. Cultivable bacterial diversity from the human colon. Lett. Appl. Microbiol. 44:343–350. 47.Belenguer, A., S. H. Duncan, G. Holtrop, S. E. Anderson, G. E. Lobley, and H. J. Flint. 2007. Impact of pH on lactate formation and utilization by human fecal microbial communities. Appl. Environ. Microbiol. 73:6526–6533. 48.Harmsen, H. J. M., G. C. Raangs, T. He, J. E. Degener, and G. W. Welling. 2002. Extensive set of 16S rRNAbased probes for detection of bacteria in human feces. Appl. Environ. Microbiol. 68:2982–2990. 49.Duncan, S. H., P. Louis, and H. J. Flint. 2004. Lactate-utilizing bacteria, isolated from human feces that produce butyrate as a major fermentation product. Appl. Environ. Microbiol. 70:5810–5817. 50.Taylor, R. 2002. Hindgut function in the laying hen: A report for the Rural Industries Research and Development Corporation. Publication No. 02/043, Newcastle, Australia. 51.Macfarlane, G. T., J. Cummings, and C. Allison. 1986. Protein degradation by human intestinal bacteria. J. Gen. Microbiol. 132:1647–1656. 52.Macfarlane, G. T., G. R. Gibson, E. Beatty, and J. H. Cummings. 1992. Estimation of short-chain fatty acids production from protein by human intestinal bacteria based on branched-chain fatty acids measurements. FEMS Microbiol. Ecol. 101:819–822. 53.Macfarlane, S., and G. T. Macfarlane. 1995. Proteolysis and amino acid fermentation. Pages 75–100 in Human Colonic Bacteria: Role in Nutrition, Physiology and Pathology. G. R. Gibson and G.T. Macfarlane, ed. CRC Press, Boca Raton, FL. 54.Capano, G., K. J. Bloch, E. J. Schiffrin, J. A. Dascoli, E. J. Israel, and P. R. Harmatz. 1994. Influence of the polyamine, spermidine, on intestinal maturation and dietary antigen uptake in the neonatal rat. J. Pediatr. Gastroenterol. Nutr. 19:34–42. 55.McCracken, V. J., and R. G. Lorenz. 2001. The gastrointestinal ecosystem: A precarious alliance among epithelium, immunity and microbiota. Cell. Microbiol. 3:1–11. 56.Boyle, E. C., and B. B. Finlay. 2003. Bacterial pathogenesis: Exploiting cellular adherence. Curr. Opin. Cell Biol. 15:633–639. 57.Torres, A. G., X. Zhou, and J. B. Kaper. 2005. Adherence of diarrheagenic Escherichia coli strains to epithelial cells. Infect. Immun. 73:18–29. 58.Cossart, P., and P. J. Sansonetti. 2004. Bacterial invasion: The paradigms of enteroinvasive pathogens. Science 304:242–348. 59.Gruenheid, S., and B. B. Finlay. 2003. Microbial pathogenesis and cytoskeletal function. Nature 422:775– 781. 60.Corr, S. C., Y. Li, C. U. Riedel, P. W. O’Toole, C. Hill, and C. G. Gahan. 2007. Bacteriocin production as a mecha- 658 nism for the anti-infective activity of Lactobacillus salivarius UCC118. Proc. Natl. Acad. Sci. USA 104:7617–7621. 61.Kaper, J. B., and V. Sperandio. 2005. Bacterial cellto-cell signaling in the gastrointestinal tract. Infect. Immun. 73:3197–3209. 62.Miller, M. B., and B. L. Bassler. 2001. Quorum sensing in bacteria. Annu. Rev. Microbiol. 55:165–199. 63.McCracken, K. J., T. C. Murphy, M. R. Bedford, and J. Apajalahti. 2006. Proc. XII WPSA Eur. Poult. Conf. September 10–14, Verona, Italy. World’s Poult. Sci. Assoc., Beekbergen, the Netherlands. JAPR: Symposium 64.Turnbaugh, P. J., M. Hamady, T. Yatsunenko, B. L. Cantarel, A. Duncan, R. E. Ley, M. L. Sogin, W. J. Jones, B. A. Roe, J. P. Affourtit, M. Egholm, B. Henrissat, A. C. Heath, R. Knight, and J. I. Gordon. 2009. A core gut microbiome in obese and lean twins. Nature 457:480–484. 65.Hughes, R. J. 2007. The rate of passage of digesta influences energy metabolism in broiler chickens. Proc. Aust. Poult. Sci. 16:63–66.