* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Metabolic Interactions of aPurple Sulfur Bacterium and a Facultative
Metabolic network modelling wikipedia , lookup
Bioluminescence wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Butyric acid wikipedia , lookup
Biochemistry wikipedia , lookup
Community fingerprinting wikipedia , lookup
Magnetotactic bacteria wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
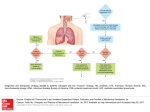
Metabolic Interactions of a Purple Sulfur Bacterium and a Facultative Anaerobe Laura K. 1 Baumgartner and Adam C. 2 Martiny Microbial Diversity 2001 Marine Biological Laboratory Woods Hole, MA . 1 U niversity of Connecticut Department of Marine Sciences, 1084 Shennecossett Rd, Groton CT 06340, LKB00002Uconn.edu 2 University of Denmark, BioCentrum-DTU, Buil ding 301, 2800 Lyngby, Denmark, [email protected] . Introduction Purple Sulfur Bacteria: Overview and An Unusual Co-Culture Purple sulfur bacteria (PSB) are generally anaerobic photolithoautotrophs that conduct anoxygenic photosynthesis with sulf ide as the electron donor (Pfenning and Truper 2001). However, some strains hav e been shown to grow heterotrophically as well as lithotrophically (Bauld et al. 1987), and some strains can utilize acetate, mal ate, pyruvate, and succinate (Krassilnikova and Kondratyeva, 1984). PSB are commo n in marine environments such as microbial mat s (van Gemerden 1993). We examined an unusual colony formatio n that arose in PSB enrichments •- 1) from Sippewisset Salt Marsh, Cr”? Cod , Massachusetts. Under the . Figure 1: Satellite colonies in the prim ary enrichment. The white bacteri um is surrounded by a halo of purple bacterium displaying enhanced growth. dissecting microscope, the initial colonies consisted of a white disc-shaped colony surrounded by several different color morphs of red to purple colonies that are presumed to be PSB. These satellite colonies developed in all but the highes t dilution of the shake tubes, and their developmen t appeared to occur in stages: first the (darker) PSB appeared as small colonies throughout the tube, then the white colo (strain ACMOI) appeared, and finally nies the PSB colonies nearest the ACMOI gre w relative to the more distant PSB colonies and more colonies appeared near AC MOI. This phenomenon was interesting for two reasons. First, because the media was highly limiting (see methods and Append ix A), the metabolic strategy of AC MOI was unclear. Second, the enhanced growth and spatial distribution of the PSB imp lied some interaction between the two strains. 2 • We were interested in several questions. First, what is the iden tity of ACMOI? Second, is ACMOI growing on some product of the PSB, or is it growing from a component of the media (including agar)? Third, what is the inter action between ACMOI and the PSB that resulted in the satellite colony form ation? These questions were examined with both traditional microbiological and mole cular analyses. Methods Sample source and Enrichment The bacteria were enriched and isolated from a sediment samp le from Sippewisset Salt Marsh, Cape Cod, MA. The sediment was dark pink and is known to be enriched for PSB. A small inoculum (—1 g) was initially enriched in CR medium (see Appendix A). After nine days of growth, 0.5 mL of the enric hment was diluted in shake tubes with CR medium in 1% agar (deep agar dilution series, see Appendix A). Further liquid cultures of the PSB also utilized CR medium as a base , and all enrichments grown in the light were given broad-spectrum light from stand ard incandescent bulbs. Specific Shake Enrichments Further shake enrichments were carried out to examine the metabolism of PSBI and ACMOI. Mixed cultures (predominantly ACMOI) were grown in the dark to assess the phototrophic requirements of ACMOI and PSBI. For PSB I, enrichments included the addition of galactose, glucose, or 0.2 im filter-sterilize d supernatant from an ACMOI culture. ACMOI was also grown in shake tubes with 0.1% bromothymol blue to assess pH change (pH-indicator, blue at 7.6 and yellow at 6.0). Liquid Cultures Both strains were maintained in CR medium. Mixed cultu res (predominantly ACMO1) were grown in liquid CR medium in the dark to asses s the phototrophic requirements of ACMOI and PSB1. Metabolic preferenc es of ACMOI and PSBI were assessed by enrichment on various substrates. Concentr ated (by centrifugation) ACMOI or PSBI monoculture were added to 4 ml vials cont aining CR medium and one of the following substrates (2mM): acetate, butyrate, form ate, fumarate, galactose, glucose, lactate, malate, propionate, succinate, as well as 5% supernatant, ‘10% supernatant, 25% supernatant, 50% supernatant, and CR medium (control) for the PSBI (the supernatant percentages represent the perc entage of filter-sterilized supernatant from a galactose-enriched ACMOI culture in the total media volume). After three days of growth, the optical density of each enrichme nt was measured spectrometrically as absorbance at either 550nm (ACM OI) or 660 nm (PSBI). Plating ACMOI was plated aerobically and anaerobically on seaw ater and freshwater 1.5% agar plates, with and without malate, (all combina tions, see Appendix A for media). PSBI was plated on freshwater malate and seawater POB plates aerobically and anaerobically in continuous light and dark condition s. However, no growth was achieved on any PSBI plates, and the results will there fore not be discussed. DNA Extraction and Amplification Satellite colonies were extracted from the 102 dilutio n of the original shake series by removing the agar from the tube and extracting individ ual colony formations with suction. DNA was extracted from these colonies with Mo Bio Ultra Clean Soil DNA kits and amplified with 8F and 1495 reverse primers at 55 °C annealing temperature. 3 • • . Cloning, Restriction Fragment Length Pol ymorphism, and Sequencing The amplified 168 rDNA was cloned with the Invit rogen TOPA TA Cloning kit. Initial clones were grown on ampicillin-X-gal plate s for 48 hours at 37°C. Fifty clones were picked at random and transferred to another ampicillin-X-Gal plate and allowed to grow for 48 hours at 37 oC. PCR from 35 of thes e clones was then amplified using MI3F and MI3R primers. This PCR product was then digested with Rsal and Hpall for 2 hours at 37°C and separated on a 2% agarose gel for comparison of digested bands. Products that appeared to have distinct band patte rns were sent to Accugenix for sequencing. 16S rDNA from ACMO1 was extracted from pure culture with Promega Wizard Genomic DNA kit and amplified via similar meth ods, and the full 16S rRNA was sequenced in 6 runs by Accugenix. Phylogeny for all sequences was conducted with ARB using Ribosomal Database Project databas es. Results and Discussion Purple Sulfur Bacterium (PSBI) Microscopic, Spectral Observations At higher magnification (400-l000x), the purple sulfur bacteria isolates are short rods or long cocci with internal sulfur granules, many of which are dividing and all of which are motile (Figure 2), possibly Chromatium (Jane Gibson, personal communication). Cells have been observed to reverse direction of rotation, and under transmission electron microscopy, the cells have a single polar flagellum. Spectral analysis of PSBI revealed peaks at 590, 800, and 850 nm, all of which are indicative of bacteriochlorophyll a. PSBI rapidly utilized sulfide, and sulfide depl eted cultures (no internal granules) regained internal granules within 10 hours with the addition of 2mM S 2 (as Na S, 2 pH7). Also, PSBI displayed chemotaxis with sulfide gradients, moving away from high concentrations of added sulfide on a microsco pe slide to an intermediate concentration, and then moving back towards the point of addi tion as sulfide was consumed. Figure 2: PSBI at 400x and I000x. The left image is from a sulfide-starved cultu re, and the bacteria contain no internal granules. The right image is from a sulfide-enrich ed culture, and internal granules are evid ent as highly refractive spheres. 4 Metabolic Observations PSB and ACMOI were grown in mixed culture in shake tubes in the dark with no apparent PSB colonies forming. When the enrichments were place d in constant light, PSB colonies appeared, indicating that PSB may be an oblig ate phototroph. However, PSBI grown axenically did not form the large colonies that were observed in mixed culture. The addition of 0.2 jim filter-sterilized supernatant from ACMO1 cultures enhanced colony growth while media alone had no affect (micro scopically, the supernatant did not contain any ACMOI). This experiment will be discussed in greater detail below, but the result hinted that some metabolic product of ACMOI was stimulating PSBI growth. Therefore PSB were enriched with vario us carbon sources (sugars, fatty acids, and different concentrations of supernata nt from ACMO1) and the optical density of the enrichments was measured after two days (Figure 3) Enrichments with fatty acid additions showed enhanced growth over the control, as did enrichments with sugars and supernatant. However, the greater degree of enhancement displayed by the fatty acid enrichments appe ars to indicate that PSBI utilized these substances more easily or gained more from them. Shake tubes of PSBI with various carbon additions were also carried out, but these shakes had no apparent growth within the time of this study. . Optical Density after 2 Days E 4.ICD .cD 0.6 0.5 0.4 0.3 —o 0.2 .11.0 0.1 0. 0 Substrate Figure 3: Optical density of PSBI cultures at 660 nm after 2 days. Sugars and supematant from ACMOI slightly enhanced growth over the control, while enrichments with fatty acids showed strongly enhanced growth. Isolation and characterization of strain ACMOI ACMO1 was isolated from the white central colony in the shake-tube. This was done by removing (sucking) a piece of the colony and trans ferring it to liquid media and shake tubes and incubate it in the dark. Light microscopy showed a non-motile, black 5 ( rod 2-10 m long (depending on growth conditions), which was consistent with earlier observations of the mixed culture. The pure culture was incubated under anaerobic cond itions with various carbon/energy sources and was shown to grow on agar and agarose as the sole energy source as well as hexoses (see Table 1). Table 1. Conditions that initially promoted cirowth of ACMOI Media (incu. dark) Media (incu. light) Glucose Galactose Agar Agarose Glucuronic acid + + + + ACMOI was streaked aerobically on seawaterand freshwater-base plates with and without malate. The strain grew only on seaw ater-based plates both in the presence and absence of malate. The strain clear ly degraded agar and produced a small halo around the colony (see Figure 4). . Figure 4: An ACMOI colony on a agar plate. Note the darkened halo of degraded agar. The combined data suggested that the strain was able to utilize agar as a sole carbon/energy source. Gas was produced durin g degradation of agar and gas bubbles were formed in the shake tubes (see Figure 5a). These bubbles could be CO 2 or H . 2 6 . Figure 5a (left) and b (right): a) Gas bubbles in a shake of ACMOI. b) Shake dilutions of ACMOI with bromothymol blue. Note the yellow color indicat ing increased acidity in the lower dilutions. Bromothymol blue was added to measure a pH-change during degradation of agar (see Figure 5b). The results show a decrease in pH at high densities of ACMOI, which is contributed to the production of fatty acids. Attempted identification of this acid using GC and HPLC was unsuccessful. Since the strain did not grow on freshwater-based plates, it poin ted toward a requirement of a certain salt concentration. This was tested by incubating ACMOI in galactose containing media with various salt concentrations (see Figure 6). Figure 6: ACMOI enrichments with varied salt concen trations. A minimum salt concentration of 0.5-1% is required for growth. 7 ACMO1 was grown on various substrates to further investiga te and characterize its metabolic properties under anaerobic conditions (see Figu re 7). The results show increased growth on galactose compared to various fatty acids . Optical Density 4-’ (U 0.21 U) 0 C (U .0 0.18 I 0.15 0 (I) 0.12 .11(0 0.09 C U) 0.06 to (U 0 4-’ 0. 0.03 0.00 0 ? -C’ . C, Substrate Figure 7: Optical density of ACMOI at 560 nm indicating different cell densities on different substrates. Biolog plates were also used for a more complete investigation of the strains metabolic properties under aerobic and anaerobic condition s. Unfortunately, no color development was observed after 4 days of incubation indicat ing either no growth or no production of reducing equivalents. Therefore, it is not clear whether the strain can perform aerobic respiration or is simply aerotolerant. Molecular Analysis . The restriction fragment length polymorphism yielded 17 distinct band patterns that were sent for sequencing (Figure 8). A phylogen etic analysis of the sequences using the neighbor-joining algorithm from these RFLP-gr oups fell into 6 groups: cytophagas, chromatiaceae, rhodobacters, vibrios, spiro chaetes, and psuedoalteromonads (Figures 9-11, products noted as PS#, with numbers corresponding to Figure 8). The sequence from ACMOI also fell into the cytophaga, near Cytophaga fermentans. The Chromatium sequence s may be representative of PSBI. Vibrios and rhodobacters are common in mat envi ronments, as are spirochaetes, which are also known to utilize agar degradati on products. While Figure 9 shows all of the products together, each sequence was careful ly examined in smaller trees to ensure accurate positioning. 8 . Figure 8: RFLP pattern from clone library DNA. Numbered lanes were judged to have distinct band patterns and PCR product corresponding to these lanes was sequenced. Numbers correspon d to the numbers in the phylogenetic trees. “L” denotes a ladder. Phylogenetic tree Cytophage Purple Non-sulfur Becteria Chromatie, Purple Sulfur acteria Psuedoalteromonads Vibrios Spirochaetes 0.1 Figure 9: Phylogenetic tree from clone library (“PS”) and ACMOI PCR products. Note that sequences similar to ACMOI appeared in the clone library, as did purple sulfur- arid purple non sulfur bacteria, a vibrio, a spirochaete, and a psue doalteromonad. 9 4 Phyogenetic tree BzetIine3 Cytophaga fermentans 1 P 512 JLP524 P519 PS29 I .hRCI1e11198313489 I SUCCIflIC8flS odoratum odoratus . • 8zeti i rier Anaeroflexus maritimus Figure 10: Phylogenetic positioning of the Cytophaga-related sequences, including the full sequence from ACMOI. 10 Chromatium purpuratum Unknown 1144, phototrophic 90Th I nus — PS2 PSi 5 L_PS31 Phototrophic Bacteria Ppc8acte — 1 P pcBaet5 I PpcBack3 99LRosea PpcBact2 PpcUact4 578Purpu Unkno4 Gamma —Gamma. 1 • SuuEndos Rbmtlar In — — Unknoli43 S78Roseu 35TV 101 a tepidum Chromatium vinosum cBact6 nromatium okenii Thiocystis qelatinosa . 0. 1 Figure 11: Phylogenetic positioning of the Chromati um-related sequences. PSBI is believed to be a Chromatium. Interaction Both the strain from the central white colony and the purple sulfur bacterium were isolated. The two strains did therefore not have an oblig ate syntrophic relationship. It was more likely that one strain provided a substrate that promoted growth of the other. As isolated culture ACMOI formed large colonies in shake tubes and grew up overnight in liquid media containing galactose. In contrast, PSB I formed tiny colonies compared to the mixed culture. This suggested that ACMO1 provided a substrate from degradation of agar that promoted growth of PSB . To test this hypothesis, supernatant from galactose- and glucose-containing media inoc ulated overnight with ACMOI was sterile-filtrated and added to a 7 day old shake-tub e culture of PSB. As a control, normal media was added to a higher density shak e. . 11 . Figure 12: Shake tubes of PSB with supematant or media added superficially. The supernatant of galactose and glucose degraded by ACMOI enhanced the growth of PSB. The top-liquid was inspected microscopically to ensure that no ACMOI bacteri a was transferred and proliferated. This experiment was repeated in liquid culture (see Figure 3). Here one can see that an increase in the amount of supernatant enhances the growth of PSB, and thereby confirming the result. Additionally, Figure 3 shows that the growth of PSB is generally improved by addition of short-chain fatty acids. Combined with the fact that acid is produced during degradation of agar, these results could point toward an excretion of a short-chain fatty acid by ACMOI, which in turn is then utilized, possibly assimilated, by PSB during growth. Another explanation could be that the gas formed during degr adation of agar by ACMOI is hydrogen, which provides PSB with reducing equivalen ts that could be used for reduction of CO . Hydrogen has not been directly measured but one might 2 expect that produced CO 2 would dissolve and not produce bubbles. On the othe r hand , if larger amounts CO 2 is generated it could form bubbles. Only a direct meas urem ent of the gas could discriminate between the two possibilities. To test if hydrogen could be a possible metabolic linkage betw een ACMOI and PSB, a liquid PSB culture was incubated with either a 2 /C0 or 2 N /C0 headspace. H Additionally, acetate was added (Figure 13). 12 Figure 13: Optical density of PSBI grown with nitrogen or hydrog en and with or without acetate. Both acetate and hydrogen can be used growth by PSB and either one or both of the substrates are possibly transferred between ACMO1 and PSB. Since the supernatant significantly enhanced growth where one would expect a low concentration of H , it is reasonable to say that at least a soluble compound is involv 2 ed, presumably a short-chain fatty acid. Finally, a constant number of PSBI was added to shake tubes with a dilution of ACMOI in an attempt to regenerate the satellite formation. PSBI only formed satellite colonies at low densities of ACMOI. This indicates that a chemic al gradient of a compound or relative cell densities may influence this phenomenon. Conclusions Phylogenetic analysis based on a full-length 168 rDNA sequence placed ACMOI in the genus Cytophaga. Cytophagas are known to be small rods, that possess gliding motility, are able to cleave 3-glycosidic bond, grow aerobic as well anaerobic, and some of which vary in length depending on growth conditions (pleom orphic) (Reichenbach 2001). This is consistent with the observations of ACMOI, even though gliding motility was not tested. Additionally, members from this group have appe ared in clone libraries from a sulfide rich salt marsh in California. Agar degradati ons is commonly observed in this group and it has been reported that hydrogen as well short-c hain fatty acids are possible fermentations products. Based on molecular and phenot ypic data, there is strong evidence that ACMOI belongs to the Cytophaga. Phylogenetic data generated from clone library and 16S sequ encing showed an organism belonging to the group Chromatium. Morphological examination of PSBI revealed a highly motile rod with internal sulfur granules and bacteriochlorophyll-a that is strictly phototrophic, all characteristic of Chromatium (Pfennig and Truper 2001). Combined, these results indicate that PSBI belong to the Chrom atiaceae. There is strong evidence for transfer of a metabolic product generated by ACMO1 from degradation of agar to PSBI. The presence of the fermen tation product enhances 13 growth of PSBI. It was not possible to directly identify the compound but eviden ce points towards either hydrogen or a short-chain fatty acid. Direct measurement s with either GC or HPLC are needed for a conclusive answer. The formation of PSB colonies surrounding a white colony of ACMOI is therefore generated by the gradien t of one (or both) of these compounds and is dependent of the density of both strains. Future Work Continued examination of this phenomenon would focus mainly on what compounds are created by ACMOI and exchanged with PSBI. The compo unds could potentially be identified with HPLC and GC as well as some wet chemistry analysis. Enrichments of ACMOI for gas production/identification analysis would identif y whether 2 or CO H 2 is being produced, and labeled galactose could be followed through both cultures to examine the products and their utilization. Also, growth rate experiments for both cultures on various substrates would further clarify their metabolism, and molecular analysis of PSBI would definitively identify it as a purple sulfur- or purple non-su lfur bacterium. Acknowledgements We would like to acknowledge all of the faculty and students of Microbial Diversity 2001 for their help and support, particularly Jane Gibson, Alfred Spoorman, Carrie Harwood, Bianca Brahamsha, John Waterbury, Dan Buckley, Jochen Mueller, and Louie Kerr. . 14 Literature Cited Bauld J, Favinger JL, Madigan MT, Gest H. 1987. Obligately haloph ilic Chromatium vinosum from Hamelin Pool, Shark Bay, Australia. Curr. Microbiol. 14(6): 335339. Krassilnikova EN, Kondratyeva EN. 1984. Growth of different purple bacterial species belonging to the genus Ectothiorhodospira in the dark. Mikrobiolog iya 53(3): 526-528. Pfennig N and Truper HG. 2001. The Family Chromatiaceae. in: The Prokaryotes Online. Springer-Verlag. http:link-springer-ny.com:6635 Reichenbach H. 2001. The Order Cytophagales. In: The Prokaryotes Online. Springer-Verlag. http:Iink-springer-ny.com:6635 van Gemerden H. 1993. Microbial mats: a joint venture. Mar. Geol. 113: 3-25. 0 15 Appendix A: PS Medium, shake instructions, fw and sw malate plates . 16 __ - Enrichment and isolation of anoxygenic phototrop liic bacteria Anoxygenic phototrophic bacteria are physiologically and phylogenetically diverse. In the following series of enrichments, different incubation cond itions will be used in order to obtain a wide variety of purple sulfur, purple nonsulfur, and gree n phototrophic bacteria. Parameters changed include the type of electron donor adde d (organic/inorganic), the pH, and light quality. After isolation in pure culture, strains will be characterized based on their j,igment composition, the formation and localization of elemental sulfu r globules, cell morphology and (partial) I 6S rRNA gene sequence. 1. Green and purple sulfur bacteria Green and purple sulfur bacteria grow preferentially phot olithoitotrophically by oxidizing sulfide, elemental sulfur, polysulfides or thiosulfate as elect ron-donating substrate in the light. This growth mode will thus be used to selectively enrich these bacteria from a variety of environmental samples. . Green sulfur bacteria grow at pH values between 6.5 and 7.0, at high sulfide concentrations of up to 5 mM, absorb light of wavelengths below 760 nm, and are able to exploit low light intensities. In contrast, purple sulfur bacteria grow only at lowe r sulfide concentrations, at pH values above 7.1, absorb light in the infrared wavelength rang e (up to 1030 nm), and require higher light intensities. Consequently, different light sources (fluorescent light, ER light), as well as different pH values and sulfide concentrationsof the growth medium are employed to selectively enrich either for green sulfur bacteria or purple sulfur bacteria. Subsamples from different sources (laminated mats from sand flats, black mats, pink “berries”, fluffy red/pink cell material from puddles, pink sand from sand bars in tidal rivers) are inoculated into mineral media CR or CL: I Media (per liter) CR NaC1 - S CL 20g — S 3 Ca 2 2 O H CI PO 2 KH 4 KC1 NHCI 0.15 g 0.20 g 0.50g 0.25 g After autoclaving and cooling under an 2 /C0 (90/10) atmosphere, add sterile solutions: N NaHCO 3 17 mM (final) N S O 2 9H a 1 mM (final) 2.5 mM (final) 12 vitamin sol. I ml 1 ml Trace element sol. SLI2 SLIO Final pH 7.2-7.3 6.7-6.8 Light source >800nm fluorescent light >960nm (selects for BChlb) -L— Y”/L.’ -- Composition of solutions used in construction of media during MBL 2000 lOOx FW-Base for Freshwater media (S-. C-. and N-free) Water NaCI MgC 6 O 2 H l CaC • 0 2 2H l PO 2 KH 4 KCI Keep on shelf in clean bottle I liter bOg 40 g 10 g 20 g 50g lx SW-Base for Marine media (S-. C-, and N-free) Water 20 liter NaCl 400 g MgC 6 O 2 60 g H l CaC • 0 2 2H 3g l P0 2 KH 4 4g KCI log Keep in Clean Na/gene plastic bottle lOOx Ammonium Chloride solution for freshwater or marine media Water CI 4 NH Store in clean bottle I liter 25 grams I 000x EDTA-Chelated Trace Elements Stock Solution Water EDTA Adjust pH to 6.0 with NaOH Add the following: 7H 4 FeSO O 2 B0 H 3 MnC • 0 2 4H I CoC 6 O 2 H l NIC • 2 6H20 I CuC • 0 2 2H I ZnS • 4 0 2 7H O N M • 4 0 2 2H oO a Sodium Vanadate N S 5 3 O 2 eO H a W Na • 4 0 2 2H O Filter sterilize 987 ml 5200 mg 2100 mg 30 mg 100 mg 190 mg 24 mg 2 mg 144 mg 36 mg 25 mg 6 mg 8 mg 12-Vitamin Solution Q Phosph 100 ml l000x ate buffer, 10mM, pH7.2 Riboflavin 10 mg Thiamine•HCI 100 mg L-Ascorbic acid 100 mg D-Ca-pantOtheflate 100 mg Folic acid 100 mg Niacinamide 100 mg Nicotinic acid 100 mg 4-Aminobenzoic acid 100 mg Pyridoxine•HCI 100 mg Lipoic acid 100 mg NAD 100mg Thiamine pyrophosphate 100 mg Titrate with NaOH until dissolved; Filter sterilize and freeze in 10 ml aliquots I 000x Vitamin B Solution Water 100 ml Cyanocobalamin 100 mg Titrate with HCI until dissolved; filter sterilize and freeze in 10 ml aliquots Sundry Solutions: I M MOPS Buffer, pH 7.2, filtered I M MES Buffer, pH 6.8, filtered I M MES Buffer, pH 6.5, filtered I M Sodium Sulfate, store in clean bottle on slelf I M Sodium Bicarbonate, autoclaved under 100% 2 CO in serum bottles I M Methanol, filtered 1 M Sodium Lactate, filtered 1 M Sodium Acetate, autoclaved 0.5 M 4-Hydroxybenzoic acid, titrated with NaOH until crystals dissolved, filtered 0.5 M Trimethoxybenzoic acid, titrated with NaOH until crystals dissolved, filtered 1 M Sodium Sulfide, stored under N 2 at 40 C (throw away when no longer clear) I % (wlv) Resazurin Dye, stored on shelf in clean bottle 0.4 M Fe(lll) oxyhydroxide, prepared by neutralizing Fe(lll)Cl with NaOH, washed 3x with H 0 2