* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download DI4R-NBIS-Nanjiang
Survey
Document related concepts
Transcript
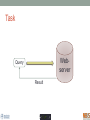
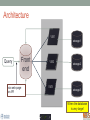
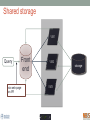
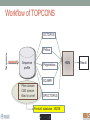
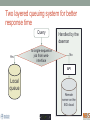
Scalable bioinformatics web-servers powered by cloud computing Nanjiang Shu National Bioinformatics Infrastructure Sweden at Science for Life Laboratory Department of Biochemistry and Biophysics at Stockholm University 2016-9-28 NBIS (National Bioinformatics Infrastructure Sweden) • A distributed national research infrastructure • Providing bioinformatics support to life science researchers in Sweden • NBIS is also the Swedish contact point to the European infrastructure for biological information ELIXIR • 79 staff at NBIS and the number is still increasing Content of support by NBIS • Genomics • Genome annotation and assembly • Mass spectrometry proteomics • Protein bioinformatics • Systems biology • Biostatistics and metabolomics • Data management and data publication • System development • Training (both internal and external) Demands for persistent data storage and web-tools hosting • Published data are likely to become non-available over time if not permanently archived (Heidorn PB. 2008) • Bioinformatics web-servers usually become dead or semi-dead after a few years if not centrally maintained • Development of a stable, robust and user-friendly web-server platform for hosting bioinformatics tools is needed Facts of the usage of web-servers Average running time per sequence vs. Methods • Running time various greatly Methods SCAMPI TOPCONS2 PconsC2 OCTOPUS Pcons-net 0 20000 40000 60000 80000 100000 120000 Average running time per sequence • Visiting frequency various greatly Demands and solutions • Flexible • Scalable • Robust Cloud computing Task Webserver Query Result Architecture VM1 storage1 Query Front end via web-page or API VM2 VM3 storage2 storage3 When the database is very large! Shared storage VM1 Query Front end via web-page or API VM2 storage VM3 Requirements for the data sharing method • Fast reading • Efficient caching system for repeated reading • Robust • Supporting multiple operating systems • Relatively easy to install Methods for data sharing NFS • Pros • Mature (since 1984) • Available on all major operating systems • Cons • Security problem • Limited by the network OneData • Pros • Silently merge small spaces to a large space • Flexible and secure authentication using token • Both web-interface and command-line interface • Cons • Not well tested (under development) Benchmark of NFS, OneData and Block storage • Two of the most widely used bioinformatics tools HMMSCAN PSIBLAST • The benchmark was carried out on one virtual machine (VM) with 4 CPUs and 8GB RAM. • OneData (oneprovider) server and the NFS server were installed on the same VM and data were stored on the same block storage volume. • All servers were running on the same EGI site (RECAS-BARI) • Randomly select 100 SWISSPORT sequences as test set. Benchmark for pure reading (wc -l) HMMSCAN PSIBLAST Most of the PSIBLAST process using database on OneData volumes failed First time reading vs repeated reading of (wc -l) First run Second run Our choice • NFS • Security problem can be minimized by sharing data within the same network under firewall. • OneData could still be a good candidate for publishing of large data • Secure and flexible authentication • Merging of multiple small spaces to a large space transparently for the user • Both web-browser and command line access Web-servers we have developed Pcons.net TOPCONS SCAMPI2 ProQ3 PconsC2/PconsC3 BOCTOPUS2 Powered by EGI cloud OCTOPUS SPOCTOPUS BOCTOPUS MPRAP TMBMODEL SubCons DGpred SCAMPI-single TOPCONS-single KalignP TOPCONS • A consensus predictor for the topology of membrane proteins and signal peptide Topology of membrane proteins Bacteriorhodopsin (PDB code 2brd) Image from G.v.H, 2006, Nature reviews N-terminal Out C-terminal I O I O I O In Workflow of TOPCONS OCTOPUS Sequence Philius Sequence profile Polyphobius SCAMPI • Pfam domain • CDD domain • Blast in uniref SPOCTOPUS Pre-built database, 300GB HMM Result TOPCONS One protein sequence (multiple sequences up to 100MB) Location of transmembrane helices and signal peptide NAR 2015 Most jobs are single-sequence jobs Two layered queuing system for better response time Query Yes Is single-sequence job from webinterface Handled by the daemon No API Local queue Remote server on the EGI cloud Single sequence jobs get fast response Performance of TOPCONS Predicting both the topology and signal peptide while still among the most accurate topology predictors NAR 2015 Usage of the web-server TOPCONS • Since 2015 • Number of finished sequences: 4517239 • Number of finished jobs: 54924 • Number of unique users (by IP address): 6997 • Number of countries the jobs have been submitted from: 73 • It has been cited 42 times during the past year Conclusions • The scalable web-server can be realized together with the cloud computing • NFS is suitable for sharing of large database among multiple virtual machines • The response time of the web-server can be improved by the two-layered queuing system • The life science community will benefit more from the bioinformatics tools with the user-friendly and responsive web-server implementation Acknowledgments • Swedish research council • System development platform at NBIS • Prof. Arne Elofsson at Stockholm University • Associate Prof. Björn Wallner at Linköping University