Slide 1
... generally modest6, 7. Here we used isobaric tag-based quantitative mass spectrometry to determine relative protein levels of 5,953 genes in lymphoblastoid cell lines from 95 diverse individuals genotyped in the HapMap Project8, 9. We found that protein levels are heritable molecular phenotypes that ...
... generally modest6, 7. Here we used isobaric tag-based quantitative mass spectrometry to determine relative protein levels of 5,953 genes in lymphoblastoid cell lines from 95 diverse individuals genotyped in the HapMap Project8, 9. We found that protein levels are heritable molecular phenotypes that ...
01 - Homework Now
... genetic ______________________. 2. A group of related genes that lie close together and that work together as a unit is called a(n) ______________________. 3. To break down lactose, Escherichia coli need three different ______________________, each of which is coded for by a different gene. 4. The t ...
... genetic ______________________. 2. A group of related genes that lie close together and that work together as a unit is called a(n) ______________________. 3. To break down lactose, Escherichia coli need three different ______________________, each of which is coded for by a different gene. 4. The t ...
Document
... complex as ssRNAs and initiate destruction of all cellular RNAs that share homology to the dsRNA. RNAi has been incredibly useful to researchers because it can be used to reduce the expression of genes that are tough to mutate. TFIID is a complex of proteins within the basal/general transcriptional ...
... complex as ssRNAs and initiate destruction of all cellular RNAs that share homology to the dsRNA. RNAi has been incredibly useful to researchers because it can be used to reduce the expression of genes that are tough to mutate. TFIID is a complex of proteins within the basal/general transcriptional ...
Gene Expression - the Biology Department
... • cis-acting elements; – DNA sequences that serve as attachments sites for the DNAbinding proteins that regulate the initiation of transcription. ...
... • cis-acting elements; – DNA sequences that serve as attachments sites for the DNAbinding proteins that regulate the initiation of transcription. ...
Biology Professor, Robert Osuna, Receives National Science
... Bacteria rely on numerous global gene regulators to rapidly control the activity of many of its genes in their attempt to protect themselves or benefit from a sudden change in their immediate environment. DksA, a fairly recently discovered bacterial gene regulator, plays an essential role in the reg ...
... Bacteria rely on numerous global gene regulators to rapidly control the activity of many of its genes in their attempt to protect themselves or benefit from a sudden change in their immediate environment. DksA, a fairly recently discovered bacterial gene regulator, plays an essential role in the reg ...
Chapter 18 - Regulation of Gene Expression - Bio-Guru
... ribosomes from binding • Any change in mRNA shape will prevent ribosome binding • Decreased length of poly-A tail will prevent translation ...
... ribosomes from binding • Any change in mRNA shape will prevent ribosome binding • Decreased length of poly-A tail will prevent translation ...
Simple tandem repeats in mammalian genomes
... repeats of the sequence CTC - are called microsatellites. For some microsatellites, therefore called "polymorphic", the number of repeats varies in different individuals "Genes" are defined as those parts of DNA-molecules that specify (encode) RNA or proteins. Only around 3% of the human genome enco ...
... repeats of the sequence CTC - are called microsatellites. For some microsatellites, therefore called "polymorphic", the number of repeats varies in different individuals "Genes" are defined as those parts of DNA-molecules that specify (encode) RNA or proteins. Only around 3% of the human genome enco ...
presentation source
... • Eukaryotes make use of transcription factors, complex multi-protein molecules that cause DNA to loop. • Therefore, blocking of regulatory proteins at some distance down a DNA sequence may effect a gene’s expression - may involve ‘enhancers’ • Binding of transcription factor begins at, but is not l ...
... • Eukaryotes make use of transcription factors, complex multi-protein molecules that cause DNA to loop. • Therefore, blocking of regulatory proteins at some distance down a DNA sequence may effect a gene’s expression - may involve ‘enhancers’ • Binding of transcription factor begins at, but is not l ...
12.5 Gene Regulation
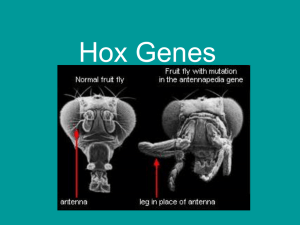
... • Show where differentiation should occur (cells and tissues) • Master control gene • Mutations in these genes can cause major developmental problems – Example: Drosophila melanogaster: replace the fly’s antennae with it’s legs – so legs were growing on the fly’s head ...
... • Show where differentiation should occur (cells and tissues) • Master control gene • Mutations in these genes can cause major developmental problems – Example: Drosophila melanogaster: replace the fly’s antennae with it’s legs – so legs were growing on the fly’s head ...
Presentation
... histones and DNA of chromatin influence both chromatin structure and gene expression Acetylation prevents histones from packing tightly, which allows genes to be expressed. Methylation causes histones to pack tightly so that genes are not expressed. ...
... histones and DNA of chromatin influence both chromatin structure and gene expression Acetylation prevents histones from packing tightly, which allows genes to be expressed. Methylation causes histones to pack tightly so that genes are not expressed. ...
Elucidating Principles of Gene Regulation from Stochastic Models
... The complexity of multicellular organisms arises largely from reusing many of the same genes in numerous combinations, rather than by the introduction of novel genes for each new celltype. Put another way, what makes you human is not so much which genes you have but how you use them. The instruction ...
... The complexity of multicellular organisms arises largely from reusing many of the same genes in numerous combinations, rather than by the introduction of novel genes for each new celltype. Put another way, what makes you human is not so much which genes you have but how you use them. The instruction ...
Gene Section LCP1 (lymphocyte cytosolic protein1) Atlas of Genetics and Cytogenetics
... Lai JL, Michaux L, Dastugue N, Vasseur F, Daudignon A, Facon T, Bauters F, Zandecki M. Cytogenetics in multiple myeloma: a multicenter study of 24 patients with t(11;14)(q13;q32) or its variant. Cancer Genet Cytogenet 1998 ...
... Lai JL, Michaux L, Dastugue N, Vasseur F, Daudignon A, Facon T, Bauters F, Zandecki M. Cytogenetics in multiple myeloma: a multicenter study of 24 patients with t(11;14)(q13;q32) or its variant. Cancer Genet Cytogenet 1998 ...
State of BER
... OptSSeq is a general tool for synthetic biology to tune pathway enzyme levels whose function can be linked to cell growth or survival. Ghosh, I. and Landick, R. OptSSeq: High-throughput sequencing readout of growth enrichment defines optimal gene expression elements for ...
... OptSSeq is a general tool for synthetic biology to tune pathway enzyme levels whose function can be linked to cell growth or survival. Ghosh, I. and Landick, R. OptSSeq: High-throughput sequencing readout of growth enrichment defines optimal gene expression elements for ...
Interferon-lambda and therapy for chronic hepatitis C virus infection
... elements (IBEs) that provide binding sites for phosphorylated IRF3 and/or IRF7. Similar binding sites are also present in the promoters of the IFN- λ genes . Therefore, it appears that the same set of transcription factors that regulate IFNB transcription also control expression of the IFN- genes. F ...
... elements (IBEs) that provide binding sites for phosphorylated IRF3 and/or IRF7. Similar binding sites are also present in the promoters of the IFN- λ genes . Therefore, it appears that the same set of transcription factors that regulate IFNB transcription also control expression of the IFN- genes. F ...
ALE #7
... 1. Please define the following important players in eukaryotic gene regulation: a. transcription factors – regulatory proteins that help RNA polymerase bind to the promoter. Thus they promote transcription. b. Activators - regulatory proteins that bind to enhancer sequences, interacting with transcr ...
... 1. Please define the following important players in eukaryotic gene regulation: a. transcription factors – regulatory proteins that help RNA polymerase bind to the promoter. Thus they promote transcription. b. Activators - regulatory proteins that bind to enhancer sequences, interacting with transcr ...