AP Biology 12
... MP3 Tutor: Control of Gene Expression chapter 18 website D8: Describe some mechanisms by which gene expression is regulated in prokaryotes (and ...
... MP3 Tutor: Control of Gene Expression chapter 18 website D8: Describe some mechanisms by which gene expression is regulated in prokaryotes (and ...
PG1005 Lecture 17 Gene Transcription
... -ribosmal RNA forms complex with multimolecular protein machinery to form the ribosome. Central to some of the processing steps involved in the production of mature mRNA ...
... -ribosmal RNA forms complex with multimolecular protein machinery to form the ribosome. Central to some of the processing steps involved in the production of mature mRNA ...
File
... variety of mechanisms that range from those that prevent transcription to those that prevent expression after the protein has been produced. The various mechanisms can be placed into one of ...
... variety of mechanisms that range from those that prevent transcription to those that prevent expression after the protein has been produced. The various mechanisms can be placed into one of ...
CHAPTER 31
... subunits) has an additional ligand-(hormone) binding domain. The DNAbinding domains of nuclear hormone receptor proteins possess globular structural domains in which four cysteines are tetrahedrally coordinated with a divalent zinc ion. Two of these zinc clusters are present on each subunit and they ...
... subunits) has an additional ligand-(hormone) binding domain. The DNAbinding domains of nuclear hormone receptor proteins possess globular structural domains in which four cysteines are tetrahedrally coordinated with a divalent zinc ion. Two of these zinc clusters are present on each subunit and they ...
Figure 10-14: Cooperative binding of activators.
... bromodomains that specifically bind to the acetyl groups. Therefore, a gene bearing acetylated nucleosomes at its promoter have a higher affinity for the transcriptional machinery than the one with unacetylated nucleosomes. ...
... bromodomains that specifically bind to the acetyl groups. Therefore, a gene bearing acetylated nucleosomes at its promoter have a higher affinity for the transcriptional machinery than the one with unacetylated nucleosomes. ...
Capacity Matrix Name: Date Started: Date Completed: Class/Course
... way. (e.g. Explain or go beyond) ...
... way. (e.g. Explain or go beyond) ...
DNA cr.eu updated plg latest
... beads represent nucleosomes. • Nucleosomes consist of eight proteins known as histone with approximately 147 base pairs of DNA wound around them; in euchromatin, this wrapping is loose so that the raw DNA may be accessed. • Each core histone possesses a `tail' structure, which can vary in several wa ...
... beads represent nucleosomes. • Nucleosomes consist of eight proteins known as histone with approximately 147 base pairs of DNA wound around them; in euchromatin, this wrapping is loose so that the raw DNA may be accessed. • Each core histone possesses a `tail' structure, which can vary in several wa ...
7.2.7 Describe the promoter as an example of non
... gene’s location. It is the binding site of RNA polymerase--the enzyme that constructs mRNA from the DNA template during Transcription. ...
... gene’s location. It is the binding site of RNA polymerase--the enzyme that constructs mRNA from the DNA template during Transcription. ...
Document
... • An operon includes a promoter, an operator, and one or more structural genes that code for all the proteins needed to do a job. – Operons are most common in prokaryotes. – The lac operon was one of the first examples of gene regulation to be discovered. – The lac operon has three genes that code f ...
... • An operon includes a promoter, an operator, and one or more structural genes that code for all the proteins needed to do a job. – Operons are most common in prokaryotes. – The lac operon was one of the first examples of gene regulation to be discovered. – The lac operon has three genes that code f ...
Hardcastle, A., et. al. Pharmacodynamic markers of response to
... PD markers for inhibitors of HSP90 The molecular chaperone HSP90 maintains the conformation, stability and function of oncogenic client proteins (eg. ERBB2, AKT and CDK4) HSP90 inhibitors cause degradation of client proteins, disruption of signalling pathways and antitumour activity Several a ...
... PD markers for inhibitors of HSP90 The molecular chaperone HSP90 maintains the conformation, stability and function of oncogenic client proteins (eg. ERBB2, AKT and CDK4) HSP90 inhibitors cause degradation of client proteins, disruption of signalling pathways and antitumour activity Several a ...
The aim of the thesis was to characterize chosen expression vectors
... Different properties of these vectors (level of expression of the cloned gene, leaky expression without inducer, dependence of expression level on inducer concentration and cell population homogeneity) were found by determination of expression level of the model gfpuv gene by fluorescence intensity ...
... Different properties of these vectors (level of expression of the cloned gene, leaky expression without inducer, dependence of expression level on inducer concentration and cell population homogeneity) were found by determination of expression level of the model gfpuv gene by fluorescence intensity ...
Ellie Degen
... As a control, we verified the function of various lysine mutations on histone methylation. Samples were run on 15% SDS-PAGE gels and transferred onto a nitrocellulose membrane. A ponceau stain was used to check for even loading of samples. After application of primary and secondary antibodies, the b ...
... As a control, we verified the function of various lysine mutations on histone methylation. Samples were run on 15% SDS-PAGE gels and transferred onto a nitrocellulose membrane. A ponceau stain was used to check for even loading of samples. After application of primary and secondary antibodies, the b ...
Unit 6B Learning Targets
... a. Transcription factors bind to specific DNA sequences and/or other regulatory proteins. b. Some of these transcription factors are activators (increase expression), while others are repressors (decrease expression). c. The combination of transcription factors binding to the regulatory regions at a ...
... a. Transcription factors bind to specific DNA sequences and/or other regulatory proteins. b. Some of these transcription factors are activators (increase expression), while others are repressors (decrease expression). c. The combination of transcription factors binding to the regulatory regions at a ...
genes, which corresponds to a greater than 1000
... Differential regulation of gene expression , in a precise temporal and spatial pattern during development, is thought to be partly mediated by site specific DNA binding proteins which promote a selective activation of qene transcription (1). From studies on XenopusTFl IA, a factor selectively requir ...
... Differential regulation of gene expression , in a precise temporal and spatial pattern during development, is thought to be partly mediated by site specific DNA binding proteins which promote a selective activation of qene transcription (1). From studies on XenopusTFl IA, a factor selectively requir ...
Slide 1
... (2) How to change the rate of a specific cellular activity? (3) Rapid vs slower change (4) Varying amount vs specific activity of a protein (5) Coordinating simultaneous changes in related proteins (6) How to achieve fine/differential regulation ...
... (2) How to change the rate of a specific cellular activity? (3) Rapid vs slower change (4) Varying amount vs specific activity of a protein (5) Coordinating simultaneous changes in related proteins (6) How to achieve fine/differential regulation ...
No Slide Title
... Effects of nuclear organization on gene expression Chromosomes in Mitosis SMC proteins:Condensins and Cohesins Centromeres and Kinetochores Paper: Loss of the Suv39h Histone Methyltransferases Impairs Mammalian Heterochromatin and Genome Stability ...
... Effects of nuclear organization on gene expression Chromosomes in Mitosis SMC proteins:Condensins and Cohesins Centromeres and Kinetochores Paper: Loss of the Suv39h Histone Methyltransferases Impairs Mammalian Heterochromatin and Genome Stability ...
Control of gene expression in prokaryotes and eukaryotes
... Gene expression is transcription of DNA to make RNA and then using the RNA to make proteins. This process can’t be left on indefinitely. The turning on and off of genes is critical to the development of an organism and the organism functioning properly throughout its life. Eukaryotic control Pretran ...
... Gene expression is transcription of DNA to make RNA and then using the RNA to make proteins. This process can’t be left on indefinitely. The turning on and off of genes is critical to the development of an organism and the organism functioning properly throughout its life. Eukaryotic control Pretran ...
Biology CELL VIABILITY AND DNA DAMAGE IN MRC5 AND HeLa
... compared with control (p<0.05). Interestingly, after knockout of HIST1H1B gene the percentage of DNA fragmentation in both cell cultures were approximately equal to 22.3% in MRC5 and 24.4% in HeLa. Levels of cells viability and DNA damage in MRC5 and HeLa cells before and after ...
... compared with control (p<0.05). Interestingly, after knockout of HIST1H1B gene the percentage of DNA fragmentation in both cell cultures were approximately equal to 22.3% in MRC5 and 24.4% in HeLa. Levels of cells viability and DNA damage in MRC5 and HeLa cells before and after ...
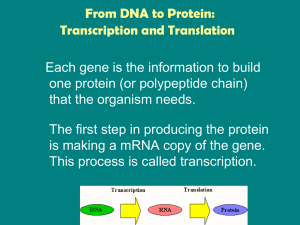
From DNA to Protein: Transcription and Translation
... The Process of Transcription The process of transcription is similar to DNA replication in that the DNA is unwound and complementary nucleotides are added. Differences: • Only a gene is copied, not the whole chromosome. • RNA nucleotides are added instead of DNA nucleotides. – Uracil is paired wit ...
... The Process of Transcription The process of transcription is similar to DNA replication in that the DNA is unwound and complementary nucleotides are added. Differences: • Only a gene is copied, not the whole chromosome. • RNA nucleotides are added instead of DNA nucleotides. – Uracil is paired wit ...
Epigenetics of Cancer
... Epigenetics/epigenomics- a definition • Any process that alters gene activity without changing the DNA sequence and leads to modifications that can be transmitted to daughter cells. • Epigenomics: global study of epigenetic changes across the entire genome ...
... Epigenetics/epigenomics- a definition • Any process that alters gene activity without changing the DNA sequence and leads to modifications that can be transmitted to daughter cells. • Epigenomics: global study of epigenetic changes across the entire genome ...
Gene Section JARID1A (jumonji, AT rich interactive domain 1A (RBBP2-like))
... 31 exons over 105 kb. ...
... 31 exons over 105 kb. ...
Supplementary Material
... with CBP and represses transcription in cell culture (Steffan et al., 2000). Furthermore, it was shown that Htt peptide, through its polyglutamine stretch, interacts primarily with the histone acetyltransferase (HAT) domain of CBP as well as other HAT co-activators (p300 and P/CAF), and inhibits the ...
... with CBP and represses transcription in cell culture (Steffan et al., 2000). Furthermore, it was shown that Htt peptide, through its polyglutamine stretch, interacts primarily with the histone acetyltransferase (HAT) domain of CBP as well as other HAT co-activators (p300 and P/CAF), and inhibits the ...
AP Biology PowerPoint Ch 19
... able to discuss mechanisms for regulating DNA and protein synthesis (know several ways) How control of DNA can lead to cancer. ...
... able to discuss mechanisms for regulating DNA and protein synthesis (know several ways) How control of DNA can lead to cancer. ...
Histone acetylation and deacetylation

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation. Histone acetylation and deacetylation are essential parts of gene regulation. These reactions are typically catalysed by enzymes with ""histone acetyltransferase"" (HAT) or ""histone deacetylase"" (HDAC) activity. Acetylation is the process where an acetyl functional group is transferred from one molecule (in this case, Acetyl-Coenzyme A) to another. Deacetylation is simply the reverse reaction where an acetyl group is removed from a molecule.Acetylated histones, octameric proteins that organize chromatin into nucleosomes and ultimately higher order structures, represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin. Condensation can be brought about by processes including deacetylation and methylation; the action of methylation is indirect and has no effect upon charge.