51 Sequence Analysis The genome projects are - Rose
... much more difficult than sequencing the DNA that codes for the protein, we have sequence information available for a great many proteins which have not had their three dimensional structure determined. In the early 1960s, C.B. Anfinsin and F. White unfolded a protein called ribonuclease. Removing th ...
... much more difficult than sequencing the DNA that codes for the protein, we have sequence information available for a great many proteins which have not had their three dimensional structure determined. In the early 1960s, C.B. Anfinsin and F. White unfolded a protein called ribonuclease. Removing th ...
Expect Values
... •Should they get a 0 (non-identical) or a 1 (identical) or something in between? ...
... •Should they get a 0 (non-identical) or a 1 (identical) or something in between? ...
Biology and computers
... similarity but have two different functions. For example, human gamma-crystallin is a lens protein that has no known enzymatic activity. It shares a high percentage of identity with E. coli quinone oxidoreductase. These proteins likely had a common ancestor but their functions diverged. ...
... similarity but have two different functions. For example, human gamma-crystallin is a lens protein that has no known enzymatic activity. It shares a high percentage of identity with E. coli quinone oxidoreductase. These proteins likely had a common ancestor but their functions diverged. ...
L2_Principle of protein folding in the cellular environment
... • Proteins that help the folding of other proteins, usually through cycles of binding and release, without forming part of their final native structure. • Increase in the efficiency, not the specificity, of protein folding • Change in emphasis from post-translational modification to co-translational ...
... • Proteins that help the folding of other proteins, usually through cycles of binding and release, without forming part of their final native structure. • Increase in the efficiency, not the specificity, of protein folding • Change in emphasis from post-translational modification to co-translational ...
The goal of protein structure prediction by threading is to find a best
... The pairwise interaction between residues depends on both the geometric features of positions close in the 3D structure, and on the specific amino acids that are aligned to those positions. There are two methods that attempt to capture these complicated dependencies. In the Filtered Neighbors Thread ...
... The pairwise interaction between residues depends on both the geometric features of positions close in the 3D structure, and on the specific amino acids that are aligned to those positions. There are two methods that attempt to capture these complicated dependencies. In the Filtered Neighbors Thread ...
Interactive Software for the study of membrane biology: lipid
... biochemistry courses. Visualization of structures generally facilitates the understanding of many related topics of membrane composition, structures, and protein interactions but they lack in many events that occurs in membranes. Also, at the present time, animations exploring solubilization and rec ...
... biochemistry courses. Visualization of structures generally facilitates the understanding of many related topics of membrane composition, structures, and protein interactions but they lack in many events that occurs in membranes. Also, at the present time, animations exploring solubilization and rec ...
Document
... De Novo Structure Prediction DEIVKMSPIIRFYSSGNAGLRTYIGDHKSCVMCTYWQNLLTYESGILLPQRSRTSR ...
... De Novo Structure Prediction DEIVKMSPIIRFYSSGNAGLRTYIGDHKSCVMCTYWQNLLTYESGILLPQRSRTSR ...
Proteins & Nucleic Acids - St. Mary Catholic Secondary School
... within the chain and R-group interactions with the environment. Tertiary structure is also aided by prosthetic groups that are inorganic compounds that act as a central point for bonding within the protein. Quaternary structure occurs when a few tertiary structures fit together to act as one functio ...
... within the chain and R-group interactions with the environment. Tertiary structure is also aided by prosthetic groups that are inorganic compounds that act as a central point for bonding within the protein. Quaternary structure occurs when a few tertiary structures fit together to act as one functio ...
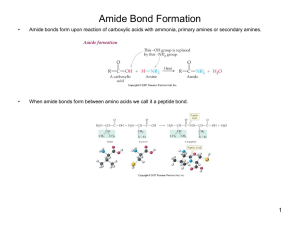
Amide Bond Formation
... Roles of proteins: – Gene expression is due to proteins – Almost all enzymes are proteins – Many hormones are proteins or peptides – Proteins form structural tissue – Storage and transportation of many molecules is possible due to proteins (think back to passive transport of hydrophilic molecules in ...
... Roles of proteins: – Gene expression is due to proteins – Almost all enzymes are proteins – Many hormones are proteins or peptides – Proteins form structural tissue – Storage and transportation of many molecules is possible due to proteins (think back to passive transport of hydrophilic molecules in ...
ans - Gogarten Lab
... 20. Which of the following is NOT part of the explanation for how complex functional molecules were assembled, despite the vastness of protein space? A. Gaia directs protein evolution, through negative feedback loops, to the correct region of protein space. B. There are multiple unrelated soluti ...
... 20. Which of the following is NOT part of the explanation for how complex functional molecules were assembled, despite the vastness of protein space? A. Gaia directs protein evolution, through negative feedback loops, to the correct region of protein space. B. There are multiple unrelated soluti ...
PowerPoint 0.8MB - The Biomolecular Modeling & Computational
... Is fold recognition useful? • In how many ways do protein fold? – 104 protein structures determined ...
... Is fold recognition useful? • In how many ways do protein fold? – 104 protein structures determined ...
PPT
... • Rosetta Stone has done fairly well at CASP competitions– David Baker at U. of Washington • Accuracy of predictions still in question ...
... • Rosetta Stone has done fairly well at CASP competitions– David Baker at U. of Washington • Accuracy of predictions still in question ...
Most Proteins Don`t Exist!
... It is just a little over fifty years since Max Perutz and John Kendrew were awarded the Nobel Prize for determining the three dimensional structure of proteins, known as the tertiary structure. Four years before Perutz and Kendrew received their Nobel Prize, Fred Sanger had received his first Nobel ...
... It is just a little over fifty years since Max Perutz and John Kendrew were awarded the Nobel Prize for determining the three dimensional structure of proteins, known as the tertiary structure. Four years before Perutz and Kendrew received their Nobel Prize, Fred Sanger had received his first Nobel ...
Document
... Sequence identity and homology: poor coverage the two proteins have the same fold,both bind heme and oxygen in same place: good independent structural/functional evidence for homology... Yet alignments of their sequences reveal only 24% identity. There are also many examples of related globins and ...
... Sequence identity and homology: poor coverage the two proteins have the same fold,both bind heme and oxygen in same place: good independent structural/functional evidence for homology... Yet alignments of their sequences reveal only 24% identity. There are also many examples of related globins and ...
Lecture 5
... Subset of turn-lovers. Glycine is special because it is so flexible, so it can easily make the sharp turns and bends needed in a b-turn. Proline is special because it is so rigid; you could say that it is pre-bend for the b-turn. Aspartic acid, asparagine, and serine have in common that they have sh ...
... Subset of turn-lovers. Glycine is special because it is so flexible, so it can easily make the sharp turns and bends needed in a b-turn. Proline is special because it is so rigid; you could say that it is pre-bend for the b-turn. Aspartic acid, asparagine, and serine have in common that they have sh ...
Modelling proteomes
... expanded irregular unique shape precisely ordered stable/functional globular/compact helices and sheets ...
... expanded irregular unique shape precisely ordered stable/functional globular/compact helices and sheets ...
C h e m g u id e –... PROTEINS: STRUCTURE
... f) What is represented by the string-like sections of the protein? g) Proteins are described in terms of their primary, secondary and tertiary structures (and quaternary, but we aren’t concerned with that at this level). What level of structure are we talking about in the diagrams above? 3. The over ...
... f) What is represented by the string-like sections of the protein? g) Proteins are described in terms of their primary, secondary and tertiary structures (and quaternary, but we aren’t concerned with that at this level). What level of structure are we talking about in the diagrams above? 3. The over ...
Amino acid sequence alignment of a `small` citrate synthase from
... considered for many years to contain a single molecular form of CS. There is also evidence that CS isoenzymes may have different metabolic roles [6]. Citrate synthase isoenzymes have recently been purified to homogeneity, from Pseudomonas aeruginosa PAC514 [7]. We have determined the N-terminal 23 a ...
... considered for many years to contain a single molecular form of CS. There is also evidence that CS isoenzymes may have different metabolic roles [6]. Citrate synthase isoenzymes have recently been purified to homogeneity, from Pseudomonas aeruginosa PAC514 [7]. We have determined the N-terminal 23 a ...
Three Dimensional Protein Structures
... Conformation: Spatial arrangement of atoms that depend on bonds and bond rotations. Proteins can change conformation, however, most proteins have a stable “native” conformation. The native protein is folded through weak interactions: a) Hydrophobic interactions b) Hydrogen-bonds c) Ionic interaction ...
... Conformation: Spatial arrangement of atoms that depend on bonds and bond rotations. Proteins can change conformation, however, most proteins have a stable “native” conformation. The native protein is folded through weak interactions: a) Hydrophobic interactions b) Hydrogen-bonds c) Ionic interaction ...
Übung: Monte Carlo, Molecular Dynamics
... I try to make a continuous form of the functions by fitting the data to a polynomial function like scorerij k1 k2 x k3 x2 k4 x3 ... Why would a sane person do this ? Where may it break down ? ...
... I try to make a continuous form of the functions by fitting the data to a polynomial function like scorerij k1 k2 x k3 x2 k4 x3 ... Why would a sane person do this ? Where may it break down ? ...
Full Text
... INTRODUCTION Discrete protein sequence motifs are widely used to describe homology between proteins and establish relationships between well-known and new protein sequences. More specifically, discrete motifs identify amino acids sharing important properties conserved in evolution. Further, they are ...
... INTRODUCTION Discrete protein sequence motifs are widely used to describe homology between proteins and establish relationships between well-known and new protein sequences. More specifically, discrete motifs identify amino acids sharing important properties conserved in evolution. Further, they are ...
Structural alignment

Structural alignment attempts to establish homology between two or more polymer structures based on their shape and three-dimensional conformation. This process is usually applied to protein tertiary structures but can also be used for large RNA molecules. In contrast to simple structural superposition, where at least some equivalent residues of the two structures are known, structural alignment requires no a priori knowledge of equivalent positions. Structural alignment is a valuable tool for the comparison of proteins with low sequence similarity, where evolutionary relationships between proteins cannot be easily detected by standard sequence alignment techniques. Structural alignment can therefore be used to imply evolutionary relationships between proteins that share very little common sequence. However, caution should be used in using the results as evidence for shared evolutionary ancestry because of the possible confounding effects of convergent evolution by which multiple unrelated amino acid sequences converge on a common tertiary structure.Structural alignments can compare two sequences or multiple sequences. Because these alignments rely on information about all the query sequences' three-dimensional conformations, the method can only be used on sequences where these structures are known. These are usually found by X-ray crystallography or NMR spectroscopy. It is possible to perform a structural alignment on structures produced by structure prediction methods. Indeed, evaluating such predictions often requires a structural alignment between the model and the true known structure to assess the model's quality. Structural alignments are especially useful in analyzing data from structural genomics and proteomics efforts, and they can be used as comparison points to evaluate alignments produced by purely sequence-based bioinformatics methods.The outputs of a structural alignment are a superposition of the atomic coordinate sets and a minimal root mean square deviation (RMSD) between the structures. The RMSD of two aligned structures indicates their divergence from one another. Structural alignment can be complicated by the existence of multiple protein domains within one or more of the input structures, because changes in relative orientation of the domains between two structures to be aligned can artificially inflate the RMSD.