The aconitase of Escherichia cok purification of the
... with fragments of lac DNA, and the other was traced to the recC and ptrIII genes at 61 min. These genes each contain a small sequence that is similar to part of the citB probe : ATCTGGCTTCACTGCTAAtGCA in recC and AACCCTtATATTCCTGaTGATtTC in ptrIII (Finch et al., 1986a, b). Because the citB probe fai ...
... with fragments of lac DNA, and the other was traced to the recC and ptrIII genes at 61 min. These genes each contain a small sequence that is similar to part of the citB probe : ATCTGGCTTCACTGCTAAtGCA in recC and AACCCTtATATTCCTGaTGATtTC in ptrIII (Finch et al., 1986a, b). Because the citB probe fai ...
SNP Analysis of the PTC Gene Using PCR
... into the air. A nearby colleague exclaimed how bitter the powder tasted, but Fox (who was closer to the chemical) tasted nothing. Interested, both men took turns tasting the chemical. Fox continued to find the chemical tasteless while his college found it bitter. Next, Fox tested a large number of pe ...
... into the air. A nearby colleague exclaimed how bitter the powder tasted, but Fox (who was closer to the chemical) tasted nothing. Interested, both men took turns tasting the chemical. Fox continued to find the chemical tasteless while his college found it bitter. Next, Fox tested a large number of pe ...
Skeletal muscle actin mRNA. Characterization of the 3
... Chick skeletal muscle A—A. Hybridizations were performed under conditions of excess RNA (as described previously (27). Hybridization reactions were incubated at 68°C for 1-5 hours. Each reaction contained in a total volume of 20 pi about 2000 cpm of 32 P-labelled DNA. ...
... Chick skeletal muscle A—A. Hybridizations were performed under conditions of excess RNA (as described previously (27). Hybridization reactions were incubated at 68°C for 1-5 hours. Each reaction contained in a total volume of 20 pi about 2000 cpm of 32 P-labelled DNA. ...
Rolling circle transcription on smallest size double stranded DNA
... Transcription kinetics of the RNAP T7 on small circular templates .................................................. 30 Summary and prospects .................................................................................................................... 35 Materials and methods ................ ...
... Transcription kinetics of the RNAP T7 on small circular templates .................................................. 30 Summary and prospects .................................................................................................................... 35 Materials and methods ................ ...
Query Results
... Layout and flowchart of the SAGExplore server. The core of the server is a MySQL database of virtual genomic SAGE tags with confidence assignments The module I is linked to NCBI BLAST server and to the Saccharomyces Genome Database (SGD) and does not require any input from the user. The modules II ...
... Layout and flowchart of the SAGExplore server. The core of the server is a MySQL database of virtual genomic SAGE tags with confidence assignments The module I is linked to NCBI BLAST server and to the Saccharomyces Genome Database (SGD) and does not require any input from the user. The modules II ...
Genetic Control of X Chromosome Inactivation in Mice: Definition of
... the choice of which chromosome is inactivated is essentially random, but can be biased by alleles at the X-linked X controlling element (Xce). Although this locus was first described nearly four decades ago, the identity and precise genomic localization of Xce remains elusive. Within the X inactivat ...
... the choice of which chromosome is inactivated is essentially random, but can be biased by alleles at the X-linked X controlling element (Xce). Although this locus was first described nearly four decades ago, the identity and precise genomic localization of Xce remains elusive. Within the X inactivat ...
document
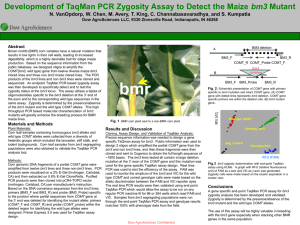
... sequenced. An endpoint TaqMan PCR based zygosity assay was then developed to specifically detect and to test the zygosity status at the bm3 locus. The assay utilizes a biplex of oligonucleotides specific to the bm3 deletion at the 3’ end of the exon and to the corresponding wild type sequences in th ...
... sequenced. An endpoint TaqMan PCR based zygosity assay was then developed to specifically detect and to test the zygosity status at the bm3 locus. The assay utilizes a biplex of oligonucleotides specific to the bm3 deletion at the 3’ end of the exon and to the corresponding wild type sequences in th ...
Protocol for Real-Time RT-PCR - MGH-PGA
... Some genes are expressed transiently or only in certain tissues. In our experience, this is the most likely cause for negative PCR results. Please read literature for the gene expression patterns. One caveat is that microarrays are not always reliable at measuring gene expressions. After switching t ...
... Some genes are expressed transiently or only in certain tissues. In our experience, this is the most likely cause for negative PCR results. Please read literature for the gene expression patterns. One caveat is that microarrays are not always reliable at measuring gene expressions. After switching t ...
HapTree-X: An integrative Bayesian framework for haplotype
... compromising accuracy. We prove theoretical bounds on the precise improvement of accuracy as a function of coverage which can be achieved from differential expression-based methods alone. Thus, the advantage of our integrative approach substantially grows as the amount of RNA-seq data increases. ...
... compromising accuracy. We prove theoretical bounds on the precise improvement of accuracy as a function of coverage which can be achieved from differential expression-based methods alone. Thus, the advantage of our integrative approach substantially grows as the amount of RNA-seq data increases. ...
Subset-Based Ant Colony Optimisation for the Discovery of Gene
... Many examples of GWAS data analysis exist in the literature that demonstrate the association between a single SNP and disease. There are known single associations for Type II diabetes and traits such as height for instance. However, there is a considerable amount of missing heritability, for example ...
... Many examples of GWAS data analysis exist in the literature that demonstrate the association between a single SNP and disease. There are known single associations for Type II diabetes and traits such as height for instance. However, there is a considerable amount of missing heritability, for example ...
Linkage analysis
... If not specific enough than you may analyze different disorders that can map to different genomic loci ...
... If not specific enough than you may analyze different disorders that can map to different genomic loci ...
USB® Thermo Sequenase Cycle Sequencing Kit
... Electrophoresis equipment—While a standard, non-gradient sequencing gel apparatus is sufficient for much sequencing work, the use of field-gradient (‘wedge’) or salt-gradient gels will allow much greater reading capacity on the gel(13, 14). A power supply offering constant voltage operation at 2000V ...
... Electrophoresis equipment—While a standard, non-gradient sequencing gel apparatus is sufficient for much sequencing work, the use of field-gradient (‘wedge’) or salt-gradient gels will allow much greater reading capacity on the gel(13, 14). A power supply offering constant voltage operation at 2000V ...
Carotene genes from cassava-pchavarriaga.pdf
... clones, using psy and pdes PCR products as probes, or by using their cDNA sequences to produce primers for RACE (Rapid Amplification of cDNA Ends) technology, to isolate psy and pdes full leght cDNAs. Screening of genomic cassava libraries from CM 523-7 and Per 297 with psy and pdes orthologous fr ...
... clones, using psy and pdes PCR products as probes, or by using their cDNA sequences to produce primers for RACE (Rapid Amplification of cDNA Ends) technology, to isolate psy and pdes full leght cDNAs. Screening of genomic cassava libraries from CM 523-7 and Per 297 with psy and pdes orthologous fr ...
Fifteen years of genomewide scans for selection: trends, lessons
... comparing variation in 2773 protein-coding genes between normal and dwarf forms of the whitefish Coregonus clupeaformis found very few divergence outliers that were protein-coding mutations, suggesting an abundance of regulatory mutations under selection (Hebert et al. 2013). Yet, it is important to ...
... comparing variation in 2773 protein-coding genes between normal and dwarf forms of the whitefish Coregonus clupeaformis found very few divergence outliers that were protein-coding mutations, suggesting an abundance of regulatory mutations under selection (Hebert et al. 2013). Yet, it is important to ...
Electrophoretic karyotypes of clinically isolated yeasts
... to that of FC18, whose chromosome sizes have been calibrated as 2.86, 2-66, 2.20, 1.90, 1.70, 1.36, 1-18 and 1.08 Mb. These chromosomal bands each represent different chromosomal DNA molecules except the largest band (2.86 Mb), which is a mixed band containing both homologues of one chromosome as we ...
... to that of FC18, whose chromosome sizes have been calibrated as 2.86, 2-66, 2.20, 1.90, 1.70, 1.36, 1-18 and 1.08 Mb. These chromosomal bands each represent different chromosomal DNA molecules except the largest band (2.86 Mb), which is a mixed band containing both homologues of one chromosome as we ...
Molecular Imaging - Engineering Computing Facility
... circumstances. This is mainly due to the fact that most imaging modalities only provide information at the anatomical level, which is insufficient for diseases at the genetic or molecular level. For instance, in cancer diagnosis and followup, it is inappropriate to merely use the physical shape and ...
... circumstances. This is mainly due to the fact that most imaging modalities only provide information at the anatomical level, which is insufficient for diseases at the genetic or molecular level. For instance, in cancer diagnosis and followup, it is inappropriate to merely use the physical shape and ...
Application Note: Targeted sequencing and chromosomal haplotype
... complete loci on the basis of crosslinking physically proximal sequences. Unlike other targeted sequencing methods, TLA works without prior detailed locus information, as one primer pair is sufficient to amplify tens to hundreds of kilobases of DNA surrounding that locus. In a separate application o ...
... complete loci on the basis of crosslinking physically proximal sequences. Unlike other targeted sequencing methods, TLA works without prior detailed locus information, as one primer pair is sufficient to amplify tens to hundreds of kilobases of DNA surrounding that locus. In a separate application o ...
PPT - Glasnost
... # The 'slices' program - slicing arrays. @sequences = ( 'TTATTATGTT', 'GCTCAGTTCT', 'GACCTCTTAA', 'CTATGCGGTA', 'ATCTGACCTC' ); print "@sequences\n"; @seq_slice = @sequences[ 1 .. 3 ]; print "@seq_slice\n"; print "@sequences\n"; @removed = splice @sequences, 1, 3; print "@sequences\n"; print "@remov ...
... # The 'slices' program - slicing arrays. @sequences = ( 'TTATTATGTT', 'GCTCAGTTCT', 'GACCTCTTAA', 'CTATGCGGTA', 'ATCTGACCTC' ); print "@sequences\n"; @seq_slice = @sequences[ 1 .. 3 ]; print "@seq_slice\n"; print "@sequences\n"; @removed = splice @sequences, 1, 3; print "@sequences\n"; print "@remov ...
Navigating the HapMap - Oxford Academic
... a difference in frequency between cases and controls, and hence an association will be seen with the trait in question. How near these polymorphisms need to be to the disease allele on average is still somewhat open to debate [4**], but is generally dependent on the population history of the sample, ...
... a difference in frequency between cases and controls, and hence an association will be seen with the trait in question. How near these polymorphisms need to be to the disease allele on average is still somewhat open to debate [4**], but is generally dependent on the population history of the sample, ...
Supplements - Haiyuan Yu
... protein. Loci are converted by separating the coding sequence of the transcript into codons, and aligning amino acids 1–n in the protein directly to codons 1–n in the transcript. All possible single-nucleotide variants in the transcript codon (3 possible alternate nucleotides × 3 possible positions ...
... protein. Loci are converted by separating the coding sequence of the transcript into codons, and aligning amino acids 1–n in the protein directly to codons 1–n in the transcript. All possible single-nucleotide variants in the transcript codon (3 possible alternate nucleotides × 3 possible positions ...
Pairing of homologous regions in the mouse genome is associated
... chromatin, position of the gene in nuclear space, and an intricate network of chromosome associations in trans determine the expression state of a particular gene. Often, co-regulated genes are found in the same transcription factory, bringing together various regions from different chromosomes [1]. ...
... chromatin, position of the gene in nuclear space, and an intricate network of chromosome associations in trans determine the expression state of a particular gene. Often, co-regulated genes are found in the same transcription factory, bringing together various regions from different chromosomes [1]. ...
Contribution of IKBKE and IFIH1 gene variants to SLE susceptibility
... located in the other gene regions persisted. Conditioning on the other SNVs, significant associations from these two SNVs at the gene 50 -region were retained in most cases; whereas they lost their significant P-values when conditioned on each other. These observations indicate that there are two inde ...
... located in the other gene regions persisted. Conditioning on the other SNVs, significant associations from these two SNVs at the gene 50 -region were retained in most cases; whereas they lost their significant P-values when conditioned on each other. These observations indicate that there are two inde ...
Laboratory 7: PCR and Ligation Reaction
... 4. Hold at 4C The run should take about 2-2.5 hours. After setting up their reactions, students should make their agarose gels. A typical minigel with 10 lanes will be sufficient for two students. The agarose concentration should be 0.8-1.0%. Part III. Analysis of PCR products Amplified genomic or ...
... 4. Hold at 4C The run should take about 2-2.5 hours. After setting up their reactions, students should make their agarose gels. A typical minigel with 10 lanes will be sufficient for two students. The agarose concentration should be 0.8-1.0%. Part III. Analysis of PCR products Amplified genomic or ...
High-resolution haplotype structure in the human genome
... allele fell exclusively or nearly exclusively on one of the major haplotype patterns and simply created a subtype of that pattern. This underscores that, when we refer to limited haplotype diversity, we are not implying complete sequence identity among chromosomes with the same haplotype, but rather ...
... allele fell exclusively or nearly exclusively on one of the major haplotype patterns and simply created a subtype of that pattern. This underscores that, when we refer to limited haplotype diversity, we are not implying complete sequence identity among chromosomes with the same haplotype, but rather ...
Abbott RealTime HIV-1
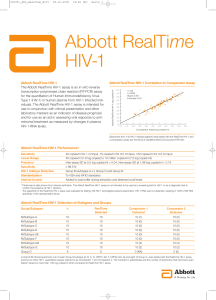
... A total of 88 clinical specimens, ten of each Group M subtype (A, B, C, D, CRF01-AE, F, CRF02-AG, G) and eight of Group O, were tested with the RealTime HIV-1 assay and by two other HIV-1 quantitative assays referred to as Comparator 1 and Comparator 2. The numbers in parentheses are the number of s ...
... A total of 88 clinical specimens, ten of each Group M subtype (A, B, C, D, CRF01-AE, F, CRF02-AG, G) and eight of Group O, were tested with the RealTime HIV-1 assay and by two other HIV-1 quantitative assays referred to as Comparator 1 and Comparator 2. The numbers in parentheses are the number of s ...
Molecular Inversion Probe

Molecular Inversion Probe (MIP) belongs to the class of Capture by Circularization molecular techniques for performing genomic partitioning, a process through which one captures and enriches specific regions of the genome. Probes used in this technique are single stranded DNA molecules and, similar to other genomic partitioning techniques, contain sequences that are complementary to the target in the genome; these probes hybridize to and capture the genomic target. MIP stands unique from other genomic partitioning strategies in that MIP probes share the common design of two genomic target complementary segments separated by a linker region. With this design, when the probe hybridizes to the target, it undergoes an inversion in configuration (as suggested by the name of the technique) and circularizes. Specifically, the two target complementary regions at the 5’ and 3’ ends of the probe become adjacent to one another while the internal linker region forms a free hanging loop. The technology has been used extensively in the HapMap project for large-scale SNP genotyping as well as for studying gene copy alterationsand characteristics of specific genomic loci to identify biomarkers for different diseases such as cancer. Key strengths of the MIP technology include its high specificity to the target and its scalability for high-throughput, multiplexed analyses where tens of thousands of genomic loci are assayed simultaneously.