* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download D. melanogaster
Mitochondrial DNA wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Gene expression programming wikipedia , lookup
Oncogenomics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Copy-number variation wikipedia , lookup
DNA barcoding wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Designer baby wikipedia , lookup
Genomic imprinting wikipedia , lookup
Y chromosome wikipedia , lookup
Transposable element wikipedia , lookup
Neocentromere wikipedia , lookup
X-inactivation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Koinophilia wikipedia , lookup
Non-coding DNA wikipedia , lookup
Microevolution wikipedia , lookup
Genome (book) wikipedia , lookup
Human genome wikipedia , lookup
Metagenomics wikipedia , lookup
Minimal genome wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Genome editing wikipedia , lookup
Helitron (biology) wikipedia , lookup
Public health genomics wikipedia , lookup
Genomic library wikipedia , lookup
Human Genome Project wikipedia , lookup
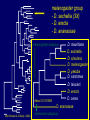
Genomics and Shallow Genomics in Drosophilidae: A Comparative Approach. Patrick M. O’Grady University of Vermont Department of Biology Talk Outline • Drosophilidae Genomics • Genomic Studies of Other Insects • Genome Enabled Studies Genomics in Drosophilidae I & II • Whole genome sequence of D. melanogaster (Rubin et al., 2000) and D. pseudoobscura (http://www.hgsc.bcm.tmc.edu/projects/drosophila/) have been determined. • Two additional taxa, D. yakuba and D. simulans, have been approved (and earmarked as “high priority”) by NIH for whole genome sequencing. (Begun & Langley; January 2003). D. pseudoobscura Sophophora The melanogaster and obscura species groups are sister taxa placed in the subgenus Sophophora. melanogaster ~150 species obscura ~60 species Drosophila (in part) D. mauritiana D. sechellia D. simulans D. melanogaster D. yakuba D. santomea D. teissieri D. erecta D. orena after Remsen & O’Grady (2002) melanogaster subgroup. • Full drosophilid genome is ~1/30th the size of the “typical” mammal. • 8X coverage, with basic annotation, can be done in 2-3 months. Full annotation will be over a longer time period. • Cost is about $1,000,000 per genome. How Genomes Get Funded… (the oversimplified version) • Write up a white paper – Feedback from larger community – Strong justification for the work • Paper goes to NIH/NHGRI panel(s) • If approved for sequencing, large genome centers bid to do the work Genomics in Drosophilidae III • Committee of drosophilid biologists met in March 2003 to discuss the possibility of proposing additional taxa for sequencing. – From a variety of backgrounds (ecology, evolutionary biology, systematics, population genetics, developmental biology, etc.). Concerns • Assembly and annotation of existing genomes. – Phylogenetic shadowing – Gene finding, role of regulatory sequences, genome component of transposable elements, adaptive evolution, functional validation of gene sequences. • Expand the taxonomic coverage to include a greater diversity of drosophilids. • Submitted a white paper proposing full genome sequencing for 8 species (Clark et al., June 2003) melanogaster group - D. sechellia (3X) - D. erecta - D. ananassae melanogaster subgroup D. mauritiana D. sechellia D. simulans D. melanogaster D. yakuba D. santomea D. teissieri D. erecta D. orena About 12-15 MYA D. ananassae after Remsen & O’Grady (2002) ananassae subgroup 2 6 1 melanogaster group Well-sampled range of 1-15 MY. obscura group -D. persimilis (3X) The obscura-melanogaster common ancestor was about 25 MYA. willistoni group - D. willistoni D. willistoni diverged from the common ancestor of the melanogaster-obscura groups about 35-40 MYA. 2 6 1 1 1 1 subgenus Drosophila - D. virilis - D. repleta - D. grimshawi The Sophophora-Drosophila divergence was 40-50 MYA. About the same as the divergence between each subgenus Drosophila group. All 8 are now ranked as “high priority.” Will have 12 full genomes over the next 2-3 years. “I’m completely confident that 10 years from now we’ll have the sequences of 50 Drosophila.” - Gerry Rubin, HHMI Genomics in Other Insects • Diptera: Aedes aegypti, Anopheles gambiae – http://klab.agsci.colostate.edu/ • Lepidoptera: Bombyx mori – http://www.ab.a.u-tokyo.ac.jp/lep-genome/new_lepgenome.htm • Hymenoptera: Apis mellifera – http://www.genome.gov/10002154 • Coleoptera: Tribolium castaneum – http://www.genome.gov/10002154 Diptera Strepsiptera Siphonaptera Mecoptera Trichoptera Lepidoptera Hymenoptera Neuroptera Megaloptera Raphidioptera Coleoptera Hemiptera Psocodea Plecoptera Dictyoptera Odonata after Whiting et al., (1997) • What is the best way to sample Diptera and other insect groups? – Phylogenetic approach? Target model systems? • What questions can be addressed using genomic data? – Generalist vs. specialist, widespread vs. rangerestricted, virulence, life history characters, etc... “In many ways we are like children in an enchanted forest, wandering almost aimlessly from discovery to discovery. For the moment, at least, that should be sufficient. At some point we will inevitably emerge into a clearing where principles and patterns in the organization and evolution of the genome are evident. Until then, let us be thankful that the pleasures of the forest are so numerous and diverse.” R. J. MacIntyre (1985) • Genome-enabled research – Systematics Mitogenomics: nearly Nearly complete complete – Genome Evolution – Comparative Genomics mt genomes Shallow nuclear genomics: multiple nuclear loci. • Nuclear loci – Goal is one gene per chromosome division – 68 are “in production” – Another 810 have been designed – 2R and 3L have not been heavily targeted to date • Examined about 200 species to date (~250 are planned. X Chromosome 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Chromosome 2L 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 Chromosome 2R 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 Chromosome 3L 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 Chromosome 3R 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 Chromosome 4 101 102 103 104 D.funebris103952 D.putrida103964 D.pallidipennis104815 D.cardini103963 D.dunni103969 D.guarani103966 D.ornatipennis103965 D.griseolineata103986 D.guttifera103968 D.immigrans103956 D.hypocausta103972 D.nasuta103957 D.lineosa103961 Z.tuberculatus105498 Z.badyi105640 Z.sepsoides105642 Z.ghesquierei105641 S.caliginosa105680 S.palmae106323 D.canalinea103953 D.pavani103955 D.hydei105429 D.navojoa105433 D.mojavensis106302 D.mercatorum106304 D.camargoi103971 D.gibberosa103960 D.nannoptera105440 D.acanthoptera105622 D.aracatas103962 D.montana103959 D.virilis105500 D.sordidula103954 D.robusta103967 D.melanica105499 D.polychaeta103958 D.crinumlily104814 D.melanogasterNM 079818 D.simulans105634 D.malerkotliana105504 D.pseudoobscura105505 H.duncani105637 S.latifasciaformis105638 D.percnosoma105685 D.waddingtoni105687 D.waddingtoni105707 D.conformis105686 D.melanoloma105708 D.austrosaltans106314 D.willistoni106322 C.procnemis106326 S.pattersoni105497 S.lebanonensis105639 Topology is similar when sampling 5,000 characters compared to 40,000. Support levels (bootstrap, jackknife) increase slowly. This amount of data is particularly useful for resolving rapid radiations (Hawaiian Drosophila). • Genome-enabled research – Systematics – Genome Evolution – Comparative Genomics Chromosome Evolution in Drosophila. Muller’s Elements species D. melanogaste r D. affinis D.willistoni D. virilis D. robusta D. grimshawi # A B C D E 4 X 2L 2R 2L 3R 5 XL 4 3 XR 2 3 XL 2R 2L XR 3 6 X 4 5 3 2 4 XL 3 2R XR 2L 6 X 3 2 5 4 F 4 5 6 4 6 – Transposition events from element to element are thought to be rare: gene content of major chromosome elements is conserved. – Extensive internal reshuffling via paracentric inversions. • First comparative genomic studies compared polytene chromosome banding patterns between different Drosophila species (Muller, 1940; Sturtevant and Novitski, 1941). Used to physically map genes In D. melanogaster we know the position of all genes Information concerning other species is lacking, but can easily be inferred by combining polytene and molecular data. • Comparisons of synteny and colinearity – Syntenic genes are on the same linkage group in different species. – Colinear genes are in the same order in different species. – What is the rate of genic transposition in the genome? – How often are so called “coadapted” gene complexes formed and lost. J. Ranz, et al., (1999) – – – – Margaret Kidwell (UA) Bill Heed (UA) Rob DeSalle (AMNH) Kenneth Kaneshiro (UH Manoa) • Technical Support: – Amy Turmelle, Jake Wintermute, Jim Bonacum