* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Slide 1
Survey
Document related concepts
Magnesium transporter wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Genome evolution wikipedia , lookup
Gene expression wikipedia , lookup
Protein adsorption wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Western blot wikipedia , lookup
Molecular evolution wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Homology modeling wikipedia , lookup
Transcript
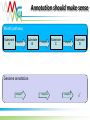
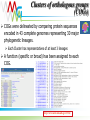
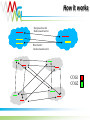
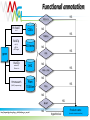
Functional annotation Datasources Konstantinos Mavrommatis [email protected] MGM workshop. 19 Oct 2010 Let’s get started… Information from databases is used to predict the function of a protein (functional annotation). Product name Enzyme catalog number Domain architecture … MGM workshop. 19 Oct 2010 But what is function? cobalamin biosynthetic enzyme, cobalt-precorrin-4 methyltransferase (CbiF) molecular/enzymatic (methyltransferase) Reaction (methylation) Substrate (cobalt-precorrin-4) Ligand (S-adenosyl-L-methionine) metabolic (cobalamin biosynthesis) physiological (maintenance of healthy nerve and red blood cells, through B12). MGM workshop. 19 Oct 2010 Functional annotation Predict the biochemistry and physiology of an organism based on its genome sequence Explain known biochemical and physiological properties MGM workshop. 19 Oct 2010 Homologs/Orthologs/Paralogs MGM workshop. 19 Oct 2010 Function prediction Function transfer by homology Homology implies a common evolutionary origin. not retention of similarity in any of their properties. Homology ≠ similarity of function. Punta & Ofran. PLOS Comp Biol. 2008 MGM workshop. 19 Oct 2010 Trust transfer of annotation ? Punta & Ofran. PLOS Comp Biol. 2008 MGM workshop. 19 Oct 2010 Dos and Don’ts Type Don’t Do Homology Same function Probability for same function Orthology Same function Probability for same function Paralogy Same function Probability for same function Sequence similarity Same function Probability for same function High sequence similarity Same function Probability for same function Same sequence Same function Probability for same function MGM workshop. 19 Oct 2010 What if nothing is similar ? Subcellular localization Gene context Special features Prediction of binding residues (DISIS, bindN) S~S S~S Periplasm Cytoplasm MGM workshop. 19 Oct 2010 Annotation should make sense Model pathway Substrate A Enzyme 1 Substrate B Enzyme 2 Substrate C Enzyme 3 Substrate D Genome annotation ? Enzyme 1 Enzyme 2 ? Enzyme 3 ✓ MGM workshop. 19 Oct 2010 Annotation should make sense MGM workshop. 19 Oct 2010 Databases Databases used for the analysis of biological molecules. Databases contain information organized in a way that allows users/researchers to retrieve and exploit it. Why bother? Store information. Organize data. Predict features (genes, functions ...). Understand relationships (metabolic reconstruction). MGM workshop. 19 Oct 2010 Primary nucleotide databases EMBL/GenBank/DDBJ (http://www.ncbi.nlm.nih.gov/,http://www.ebi.ac.uk/embl) Archive containing all sequences from: genome projects sequencing centers individual scientists patent offices The sequences are exchanged between the three centers on a daily basis. Database is doubling every 10 months. Sequences from >140,000 different species. 1400 new species added every month. Year 2004 2005 2006 2007 2008 Base pairs 44,575,745,176 56,037,734,462 69,019,290,705 83,874,179,730 99,116,431,942 Sequences 40,604,319 52,016,762 64,893,747 80,388,382 98,868,465 MGM workshop. 19 Oct 2010 Primary protein sequence databases Contain coding sequences derived from the translation of nucleotide sequences GenBank Valid translations (CDS) from nt GenBank entries. UniProtKB/TrEMBL (1996) Automatic CDS translations from EMBL. TrEMBL Release 40.3 (26-May-2009) contains 7,916,844 entries. MGM workshop. 19 Oct 2010 Errors in databases There are many errors in the primary sequence databases: In the sequences themselves: Sequencing errors. Cloning vectors sequences. In the annotations: Inaccuracies, omissions, and even mistakes. Inconsistencies between some fields. MGM workshop. 19 Oct 2010 Redundancy Redundancy is a major problem. Entries are partially or entirely duplicated: e.g. 20% of vertebrate sequences in GenBank. { { { Partial and complete sequence duplications MGM workshop. 19 Oct 2010 NCBI Derivative Sequence Data Curators RefSeq TATAGCCG AGCTCCGATA CCGATGACAA Labs Genome Assembly TATAGCCG TATAGCCG TATAGCCG TATAGCCG GenBank UniGene Algorithms MGM workshop. 19 Oct 2010 RefSeq Curated transcripts and proteins. reviewed by NCBI staff. Model transcripts and proteins. generated by computer algorithms. Assembled Genomic Regions (contigs). Chromosome records. MGM workshop. 19 Oct 2010 Secondary protein databases Uniprot/SWISS-PROT (1986) (http://ca.expasy.org/spro) a curated protein sequence database high level of annotation (such as the description of the function of a protein, its domains structure, post-translational modifications, variants, etc.) a minimal level of redundancy high level of integration with other databases MGM workshop. 19 Oct 2010 Classification databases Groups (families/clusters) of proteins based on… Overall sequence similarity. Local sequence similarity. Presence / absence of specific features (active site, signal peptides… ). Structural similarity. ... These groups contain proteins with similar properties. Specific function, enzymatic activity. General function. Evolutionary relationship. … MGM workshop. 19 Oct 2010 Overall sequence similarity MGM workshop. 19 Oct 2010 Clusters of orthologous groups (COGs) COGs were delineated by comparing protein sequences encoded in 43 complete genomes representing 30 major phylogenetic lineages. Each Cluster has representatives of at least 3 lineages A function (specific or broad) has been assigned to each COG. http://www.ncbi.nlm.nih.gov/COG/ MGM workshop. 19 Oct 2010 How it works Reciprocal best hit Bidirectional best hit Blast best hit Unidirectional best hit COG1 COG2 MGM workshop. 19 Oct 2010 Profiles & Pfam A method for classifying proteins into groups exploits region similarities, which contain valuable information (domains/profiles). These domains/profiles can be used to detect distant relationships, where only few residues are conserved. MGM workshop. 19 Oct 2010 Regions similarity MGM workshop. 19 Oct 2010 Pfam http://pfam.sanger.ac.uk HMMs of protein alignments (local) for domains, or global (cover whole protein) MGM workshop. 19 Oct 2010 TIGRfam Full length alignments. Domain alignments. Equivalogs: families of proteins with specific function. Superfamilies: families of homologous genes. HMMs http://www.tigr.org/TIGRFAMs/ MGM workshop. 19 Oct 2010 KEGG orthology MGM workshop. 19 Oct 2010 Composite pattern databases To simplify sequence analysis, the family databases are being integrated to create a unified annotation resource – InterPro Release 30.0 (Dec10) contains 21178 entries Central annotation resource, with pointers to its satellite dbs http://www.ebi.ac.uk/interpro/ MGM workshop. 19 Oct 2010 * It is up to the user to decide if the annotation is correct * MGM workshop. 19 Oct 2010 ENZYME MGM workshop. 19 Oct 2010 ENZYME http://ca.expasy.org/enzyme/ MGM workshop. 19 Oct 2010 KEGG Contains information about biochemical pathways, and protein interactions. http://www.kegg.com MGM workshop. 19 Oct 2010 Functional annotation YES IMG term PSI BLAST 1e-2 COGs NO YES TIGRfam BLASTp <1e-10, >45% id, >70% length KO terms NO gene YES COG BLASTp evalue<10, 20 best hits Hmmsearch (BLAST preprocessing) IMG NO YES COG + pfam Pfam TIGRfam NO YES Pfam NO YES BLAST http://imgweb.jgi-psf.org/img_er_v260/doc/img_er_ann.pdf NO Product name hypothetical (based on translation tables) MGM workshop. 19 Oct 2010 Sequencing projects & Metadata http://www.genomesonline.org MGM workshop. 19 Oct 2010 Literature search PubMed http://www.ncbi.nlm.nih.gov/Pubmed MGM workshop. 19 Oct 2010 Specialized databases There is a large number of databases devoted to specific organisms. For some model organisms there are often concurrent systems. These databases are typically associated to sequencing or mapping projects. MGM workshop. 19 Oct 2010 Other specialized databases Signal transduction, regulation, protein-protein interactions Gene 3D structures expression TRANSFAC (Transcription Factor database) GXD PDB(Mouse (Protein Gene Data Expression Bank) Database) BRITE (Biomolecular Relations in Information Transmission and The MMDB Stanford (Molecular Microarray Modelling Database Data Expression database) Base) Mapping DIP (Database of NRL_3D Interacting Proteins) (Non-Redundant Library of GDB (Genome Data Base) 3DInteraction Structures) BIND (Biomolecular Network database) EMG (Encyclopedia of Mouse Genome) SCOP (Structural Classification of BioCarta MGD (Mouse Genome Database) Proteins) Biochemical pathways INE (Integrated Rice Genome Explorer) Polymorphism KLOTHO (Biochemical Compounds Declarative database) Protein quantification (Allele Frequency Database) BRENDA (enzymeALFRED information system) SWISS-2DPAGE Molecular LIGAND (similar to Enzymeinteractions but with more information for substrates) PDD (Protein Disease Database) DIP (Database of Interacting proteins) Gene order and co-occurrence Sub2D (B. subtilis 2D Protein Index) BIND (Biomolecular Interaction STRING Network Database) MGM workshop. 19 Oct 2010 List of databases http://www.oxfordjournals.org/nar/database/c MGM workshop. 19 Oct 2010 Databanks interconnection Blocks MIMMAP REBASE PDBFINDER ALI PROSITEDOC OMIM ProDom PROSITE SWISSNEW ENZYME DSSP SWISSDOM HSSP FSSP GenBank PDB MOLPROBE SWISS-PROT NRL_3D ECDC EPD YPDREF PMD EMBL YPD EMNEW TFSITE TrEMBLNEW ProtFam FlyGene TrEMBL PIR TFACTOR Not all databases are updated regularly. Changes of annotation in one database are not reflected in others. MGM workshop. 19 Oct 2010 Summary Gene annotation should make sense in the context of the organism We have main archives (Genbank), and currated databases (Refseq, SwissProt), and protein classification database (COG, Pfam), and many, many more… They help predict the function, or the network of functions. Systems that integrate the information from several databases, visualize and allow handling of data in an intuitive way are required QUESTIONS? MGM workshop. 19 Oct 2010