* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download aliphatic amino acid structures
Paracrine signalling wikipedia , lookup
Signal transduction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Silencer (genetics) wikipedia , lookup
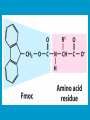
Peptide synthesis wikipedia , lookup
Gene expression wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Magnesium transporter wikipedia , lookup
Expression vector wikipedia , lookup
Biosynthesis wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Genetic code wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Point mutation wikipedia , lookup
Interactome wikipedia , lookup
Metalloprotein wikipedia , lookup
Homology modeling wikipedia , lookup
Biochemistry wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein purification wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Western blot wikipedia , lookup
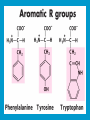
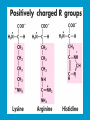
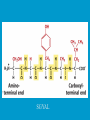
Dr. Rita P.-Y. Chen (陳佩燁) Assistant Research Fellow Institute of Biological Chemistry Academia Sinica 1 Amino acid •Chiral center •Optically active: rotate planepolarized light •Amino acid in protein: L stereoisomer Absolute configuration: D,L system Not all L-amino acids are levorotatory (rotating polarized light to the left) Specifying onfiguration: RS system L-amino acid has S configuration 4 7 8 aliphatic amino acid structures CH Isoleucine (I) (Ile) CH3 CH2 CH3 9 Side Chains with Alcohol Groups • Serine (Ser, S) and Threonine (Thr, T) have uncharged polar side chains Catalytic role, phosphorylation, o-linked glycosylation, hydrogen bond 10 Structures of aspartate, glutamate, asparagine and glutamine 11 Structures of histidine, lysine and arginine imidazole group d-Guanido group 12 Methionine and cysteine First a.a. Catalytic role, disulfide bond 13 Fig 3.4 Formation of cystine Disulfide bond 14 Aromatic amino acid structures Indole group phosphorylation 15 Proline has a nitrogen in the aliphatic ring system • Proline (Pro, P) - has a three carbon side chain bonded to the a-amino nitrogen • The heterocyclic pyrrolidine ring restricts the geometry of polypeptides 16 p t 23 Uncommon a.a. Prothrombin Ca2+ binding protein Plant cell wall Collagen Collagen elastin myosin 21th a.a. Added during protein synthesis UGA codon glutathione peroxidases Other amino acids (not constituents of proteins) : metabolite Key intermediate in biosynthesis of Arg and in urea cycle Urea p843 25 補充教材 26 Amino acid can act as acids and bases 27 較易解離 較易解離 The characteristic pH at which the net electric charge is zero is called the isoelectric point or isoelectric pH, designated pI. For glycine, which has no ionizable group in its side chain, the isoelectric point is simply the arithmetic mean of the two pKa values: 0 pI = (2.19+4.25)/2 = 3.22 0 pI = (6+9.17)/2 = 7.59 Peptides and Proteins SGYAL 9.6 4.25 +2 +1 0 -1 pI = (4.25+9.6)/2 =6.93 10.53 2.34 • • • • Artificial Sweetener, ex. in Diet Coke 200 times sweeter than sugar D-form a.a. substitution is bitter 苯酮尿症(Phenylketonuria) 患者不可使用, use Alatame instead 35 催產素 血管收縮素 36 Fig. 3-13, p.75 Glutathione • an important water-phase antioxidant and essential cofactor for antioxidant enzymes 37 Protein size is varied Polypeptides have characteristic amino acid compositions Some proteins have chemical groups other than a.a. • Non a.a. part – prosthetic group Protein purification • Fractionation: protein solubility depends on temperature, pH, salt • Dialysis • Ultrafiltration: N2 purge, centrifugation • Column chromatography • Electrophoresis 42 Column chromatography • Cation exchange chromatography: • Protein carries postive charge (cation) • Buffer pH must be lower than protein pI • No sample volume limit • Size exclusion chromatography • Big protein runs faster • Sample volume is limited • Column is usually long • Affinity chromatography • Separate proteins by their binding specificities • No sample volume limit Electrophoresis: (1) SDS-PAGE • Proteins migrate according to their size and shape • One SDS bind for every two residues • Protein is denatured, subunits will be separated cathode Anode Coomassive blue staining 4 subunits Electrophoresis: (2) Isoelectric focusing • Determine protein pI • Use ampholytes to create get with pH gradient • Proteins stop migration when pH = pI Two-dimensional electrophoresis Protein sequencing • Protein function depends on its sequence • 20 -30 % proteins are polymorphic • Most proteins contain crucial regions that are essential to their function and whose sequence is therefore conserved. 56 1953, Frederick Sanger worked out the sequence of insulin 1958 Nobel Prize in Chemistry 1980 Nobel Prize in Chemistry: DNA sequencing Only identify the first a.a. Pehr Edman: Edman degradation Large proteins must be sequenced in smaller segment • • • • • Breaking disulfide bond Cleaving the polypeptide chain Sequencing of peptides Ordering peptide fragments Locating disulfide bond 59 Breaking disulfide bond Cleaving the polypeptide chain Locating disulfide bond • Do the same thing except breaking disulfide bond • See which peptide fragments are missing or which peptide fragment (longer) appears 63 Other methods to solve protein sequence • Translate from DNA:Genome, proteome • Mass spectrometry 64 The Nobel Prize in Chemistry 1984 --for his development of methodology for chemical synthesis on a solid matrix Robert Bruce Merrifield Rockefeller University 1921-2006 Homologous proteins • Paralog: homologous genes within a single species that diverged by gene duplication. • Ortholog: genes in different species that derive from a common ancestor. Orthologous genes may or may not have the same function. 73 Molecular evolution • 1960s Zuckerkandle and Pauling use nucleotide and protein sequence to explore evolution • 1970s Carl Woese used ribosomal RNA sequence (archaebacteria is different from other bacteria) • Not every protein is a good target (choose protein with essential function ex. cellular metabolism EF-1a) • Lateral gene transfer ex. Antibiotic-resistent gene • At some position, only particular amino substitutions can be tolerated • Electronic search, multiple sequence alignment • Gap, penalty 74 Determining how closely related the proteins are – Blosum (blocks substitution matrix) Blosum 62 table (62% identity) •Based on short conserved blocks •Unique chemical properties – higher score •Each substitution has a score, based on its frequency Signature sequence – useful sequence segment in taxonomy EF-1a/EF-Tu family • Sequence in signature sequence might be quite distinct Evolutionary tree • Length of line is proportional to the number of a.a. substitution • From the sum of length, we can know how close two species are • From different proteins, we can obtain different evolutionary trees 78 79