* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Features of Hybrids
Point mutation wikipedia , lookup
Non-coding DNA wikipedia , lookup
RNA interference wikipedia , lookup
Ridge (biology) wikipedia , lookup
Human genome wikipedia , lookup
Genetic engineering wikipedia , lookup
Non-coding RNA wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Transposable element wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genomic imprinting wikipedia , lookup
Oncogenomics wikipedia , lookup
Long non-coding RNA wikipedia , lookup
RNA silencing wikipedia , lookup
Pathogenomics wikipedia , lookup
Minimal genome wikipedia , lookup
Public health genomics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Koinophilia wikipedia , lookup
Helitron (biology) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression programming wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene expression profiling wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Human–animal hybrid wikipedia , lookup
Genome evolution wikipedia , lookup
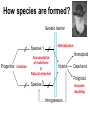
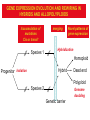
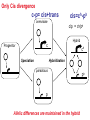
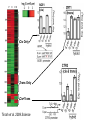
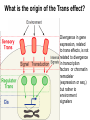
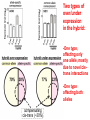
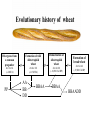
Weizmann Institute of Science Transcriptome rewiring in hybrids and polyploids Or What hybrids can teach us on biodiversity How species are formed? Genetic barrier Hybridization Species 1 Progenitor Isolation Accumulation of mutations & Natural selection Homoploid Hybrid Dead end Polyploid Species *2 Genome doubling Introgression Features of Hybrids: The benefits Heterotic features Broader range of adaptation Novel traits The troubles Embryonic lethality Adult inferiority Hybrid necrosis Sterility (chromosome pairing or nuclear-cytoplasmic interactions) Hybrid (homoploid) species Allopolyploid species AA X BB (2x) (2x) AB AABB (4x) Studying interspecific hybrids and allopolyploids in Two model systems Merging of wheat genomes Merging of yeast genomes (sensu stricto, n=16) P1 P2 + S. Paradoxux S. cerevisiae F1 Ae. sharonensis (genome SlSl) Amphiploid (SlSlAmAm) T. monococcum (genome AmAm) Collaboration with Naama Barkai’s lab Today’s question: What goes on at the molecular level during speciation/hybridization/ polyploidization ? What drives divergence in gene expression and how is gene expression rewired in hybrids/polyploids The yeast system Sensu Stricto (5-20 Mya) P1 P2 + S. Paradoxux S. cerevisiae F1 Growth vigor in the cerevisiae x paradoxus hybrid Max slope 0.06 0.05 0.04 S. paradoxus hybrid S. cerevisiae 0.03 0.02 0.01 How is gene expression rewired ? Are new patterns of expression related to heterosis? GENE EXPRESSION EVOLUTION AND REWIRING IN HYBRIDS AND ALLOPOLYPLOIDS Accumulation of mutations Cis or trans? Species 1 merging Novel patterns of gene expression Hybridization Homoploid Hybrid Progenitor Isolation Dead end Polyploid Species 2 Genome doubling Genetic barrier Only Cis divergence c-p= cis+trans cerevisiae cis=ch-ph c/p = ch/ph Hybrid c Progenitor Speciation ch Hybridization paradoxus p Allelic differences are maintained in the hybrid ph Only trans divergence c>p cerevisiae ch = ph Hybrid c Progenitor Speciation ch Hybridization paradoxus p Allelic differences are abolished in the hybrid ph Two-species array S. cerevisiae S. paradoxus TGCTCGTCGGGTAGCTAATAGCTAGTGAGATAGGTCCAGCACCATGTCACG TGCCCGTCGTGTAGCTACTGCGTAGAGATGTACGTGCAGTACCATCTCACG *** ***** ******* * *** ** ** ** *** ***** ***** Select 12 homoeoallele-specific 60mers oligos for 4400 genes Tirosh et al. 2009 Science Analyze allele expression in parental versus hybrid background P H P-H Cis Only Trans Only Cis+Trans Tirosh et al. 2009 Science Genome-wide analysis of cis-trans effects Alterations in cis factors are the prominent cause of divergence in gene expression Responsiveness to the environment is determined essentially by alterations in trans factors What is the origin of the Trans effect? Divergence in gene expression, related to trans effects, is not related to divergence in transcription factors or chromatin remodeler (expression or seq.) but rather to environment signalers Genes with both Cis and trans effects-Impact on over/under expression Same-direction mutations Enhancing Opposite direction mutations Compensating Prominence of genes with both cis and trans effects among overexpressed and underexpressed genes Two types of over/under expression in the hybrid: -One type, affecting only one allele, mostly due to novel cistrans interactions -One type affecting both alleles Novel expression patterns can be achieved “overnight” in hybrids through novel cis/trans, or trans interactions Compensatory mutations and Hybridspecific trans effect explain overexpression in hybrids In the hybrid (under rich medium) respiration genes are preferentially overexpressed, maybe causing heterosis. Thanks to: Naama Barkai Naama Barkai Itay Tirosh Itay Tirosh Michal Keinan- Dena Leshkowitz Sharon Reikhav Cathy MelamedBessudo Naomi AviviRagolsky Khalil Kashkush Moshe Feldman Evolutionary history of wheat Divergence from a common progenitor Formation of wild allotetraploid wheat 2n=2x=14 (~4 MYA) 2n=4x=28 (~0.5 MYA) PP AA BB DD BBAA Domestication of allotetraploid wheat 2n=4x=28 (~10,500 Cal BP) BBAA Formation of bread wheat 2n=6x=42 (~9,500 Cal BP) BBAADD Aegilops tauschii DD F1 BAD S1 Triticum durum BBAADD BBAA Heterosis and Hybrid necrosis TQ113 F1 S1 TTR16 Novel genetic and epigenetic variation released upon hybridization and polyploidization Gene silencing/activation Changes in DNA methylation Transposons activation DNA elimination The small RNAs hypothesis-A mechanism related to all of the above Deep sequencing of small RNA w/Illumina/Solexa system: -- Isolation of small RNAs --Amplification --Quality control (trimming “primers” from all tags, NNs, junk..) --Bioinformatic analysis (alignment, annotation, normalization..) #reads 327063 130015 102323 68524 62126 42354 25798 Signatures TCGCTTGGTGCAGATCGGGACTCGTATGCCGTC TGACAGAAGAGAGTGAGCACTCGTATGCCGTCT GGCGGATGTAGCCAAGTGGATCGTATGCCGTCT GGGCCTGTAGCTCAGAGGATCGTATGCCGTCTT TTGACAGAAGAGAGTGAGCACTCGTATGCCGTC TGAGAAGGTAGATCATAATAGCTCGTATGCCGT GCGTCTGTAGTCCAACGGTTCGTATGCCGTCTT MiR168 Analysis of small RNA data Total reads Reads after QC # signatures >30 reads Parent 1 Denome DD 3,064,285 1,202,651 4292 Parent 2 Genome BBA A 5,633,938 1,576,108 7967 F1 hybrid S1 polyploid total 6,421,516 1,689,159 6756 3,458,759 1,561,663 2772 18,578,498 6,029,581 21,787 Annotation: Hits in Micro-RNAs siRNAs corresponding to repeats, transposons Genes, tRNAs rRNA.. Transposons make up about 80% of the wheat genome and therefore present a major hazard to genome stability Adapted from Kronmiller and Wise, Plant Physiol. Vol. 146(1): 45-59, 2008 85% of small RNAs corresponding to transposons are down-regulated in the polyploid n=1113 100% 90% percent(cumulative) (cumulative) percent 80% 70% 60% hybrid polyploid 50% 40% 30% 20% 10% -16 -14 -12 -10 -8 -6 -4 0% -2 -1 0 1 2 4 6 Log2(#log2(reads/MPV) reads / # reads in MPV) <MPV hybrid 39% polyploid 85% ~MPV 42% 12% >MPV 19% 3% 8 10 12 Wis2-1A retrotransposon activation correlates with down-regulation of siRNAs Transcriptional activation of Wis2-1 in S1 Kashkush et al. Nat Genet 2003 Down-regulation of Wis2-1 siRNAs in S1 Keinan-Eichler et al. Unpublished Total small RNA count for Wis2-1A: TTR16 TQ113 MPV F1 S1 756 528 642 475 0 Northern (Wis probe) •Small RNAs are highly responsive to Polyploidization •They can be induced -- miRNAs or Repressed --siRNAs •The suppression of siRNAs correlates with transposons activation Extensive genetic and epigenetic rewiring occurs in hybrid and polyploids through novel cis-trans interactions that alter the activity of genes, repeats and small RNAs. Phenotypic consequences range from heterotic effects to genetic incompatibilities. Release of transposon silencing might lead to hybrid dysgenesis Thanks to: Naama Barkai Naama Barkai Itay Tirosh Itay Tirosh Michal Keinan- Dena Leshkowitz Sharon Reikhav Cathy MelamedBessudo Naomi AviviRagolsky Khalil Kashkush Moshe Feldman