* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Lecture 7 notes - UC Davis Plant Sciences
Survey
Document related concepts
Genomic imprinting wikipedia , lookup
Gene expression wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Gene regulatory network wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene desert wikipedia , lookup
Community fingerprinting wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Genomic library wikipedia , lookup
Non-coding DNA wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Molecular evolution wikipedia , lookup
Transcript
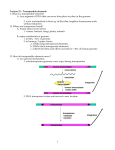
The problem: find the genes, regulatory elements, and repetitive elements in the middle of millions of ATCG Arial After a genome is completely sequenced, it can then be annotated. This operation is performed initially using computer programs that are designed to find genes (exons and introns, regulatory sequences such as TATA boxes and polyA addition sites), and repetitive elements and determine their structure. Once genes are identified, the gene order and structure can be compared with other species: comparative genomics. For prokaryotes annotation use http://compbio.ornl.gov/prodigal/server.html Arial Comparison of genomic sequences with cDNA, using the program GeneSeqer is a good strategy t t to t annotate t t genes (extrinsic ( t i i information) i f ti ) Arial Arial The absolute frequency of the pair A(k)A with k (0 to 50) nucleotides between the two As in th first the fi t 200 bp b in i a sett off 1761 h human genes and d 1753 iintrons. t A clear l ttriplet i l t pattern tt appears in the coding region but not in the introns. This pattern is a reflection of the characteristic codon usage seen in coding regions Gene prediction algorithms attempt to determine whether a particular DNA sequence constitutes a working gene. The parameters distinguishing genes from non-genes are not well understood. Although certain features, such as splice sites and ORFs, are fairly well defined other features, defined, features such as regulatory regions regions, are still very difficult to predict. predict Even identifying ORFs is not straightforward, particularly in mammalian genomes that are characterized by small exons and large introns. A further problem in gene prediction is that our knowledge of identifying features in genes is constantly expanding. Computer scientists would classify gene prediction as a problem in pattern recognition. Machine learning algorithms are good for solving problems in pattern recognition because they can be trained on a sample data set to “learn” what defines the pattern in question when well-defined rules are not available. Arial A Markov chain, model or process refers to a series of observations in which the probability off an observation b ti depends d d on a number b off previous i observations. b ti M Markov k processes describe many biological phenomena, including base-pair substitutions resulting from mutations. In some cases, the states in a Markov process are not known with certainty. The example of a dishonest gambler is often used to illustrate this point. The gambler may carry a loaded die that he or she occasionally substitutes for a fair die, but not so often that the other players would notice. The fair die has a one-in-six chance of showing any particular number. When using the loaded die, a player will have a 50% chance of rolling a one and a 10% chance of rolling any other number. It is in these types of situations that stochastic models called hidden Markov models (HMMs) are useful, because they take into account unknown (or hidden) states. For example, exactly when the cheating gambler is using a fair or loaded die is hidden from the other players, but insight may still be gained by looking at the outcome of the cheater’s rolls. If he or she rolls three ones in a row, it is more likely (a 12.5% chance) that the loaded die is being used than the fair one, which would have only a 0.5% chance of generating three ones in a row. Hidden Markov models describe the probability of transitions between hidden states, as well as the probabilities associated with each state. In the example of the cheating gambler, an HMM would describe the probabilities of rolling particular numbers given the loaded or fair die and the probability that the dishonest gambler would switch from one die to another. Hidden Markov models can be used to answer three types of questions. The first type is the likelihood question: Given a particular HMM, what is the probability of obtaining a particular outcome (e.g., rolling three ones)? The second type is the decoding question: Given a particular HMM, what is the most likely sequence of transitions between states for a particular outcome? In the case of the cheating gambler, this sequence would be the order in which he or she transitioned from one die to another. The third type yp is the learning g question: q Given a p particular outcome and set of assumptions about possible transition states, what are the best model parameters (e.g., probabilities between transition states)? This third question allows HMMs to be used for machine learning. Every HMM has a start and end state, denoted by the S and E, respectively. Hidden states lie between the start and end states. In the figure, the squares are states, and the lines between them indicate the probability of one state transitioning to another. The loops on the upper and lower states show the probabilities associated with the state remaining the same. States transition back and forth until the HMM reaches the end state. HMM can be used to assess the probability that a particular sequence is a gene or Arial Arial Arial Dynamic programming: the solution of a general problem is obtained by the recursive solution of smaller versions of the problem. The main difficulty in exon assembly is that the number of possible assemblies grows exponentially with the number of predicted exons. Dynamic programing allows to find an efficient solution without having to enumerate all possible combinations Genomscan: http://genes.mit.edu/genomescan.html Arial GRAIL and GRAIL-EXP predictions of URO-D (U30787). The top track displays the annotated exonic structure. Below GRAIL ab initio prediction. Below the alternative splicing forms identified by GRAIL-EXP by using an external EST database. The output gives the exon number, strand (forward + and reverse -), start and stop point of the exon, exon type (start, internal, terminal),its length, its raw score and qualitative description. Here GRAIL corrected predicted 5 of th tten kknown exons as wellll as partt off a one off the the th internal i t l exons Arial Arial Output: Gene, exon number, type, strand, begin end of prediction, reading frame, , scoring values and a Probability value. In this example GENSACAN correctly predicted 9 of the ten exons. Miss the first exon even if the GenomeScan version was used In Genome SCAN you enter the Protein you suspect similar using a previous BLASTX search. Arial FGENESH predictions on the URO-D sequence. The tabular format of the output gives the gene number, the strand (+ or -), the exon number within the gene, the exon type, the start and stop positions for the exon, an exon score, ORF start and stop positions, and exon length. The amino acid sequence of the predicted protein product is given below the gene prediction. The method can also predict TATA boxes and polyA tails. Here. FGENESII correctly pre-dicted seven of the ten exons, two exons were partially detected detected, and the initial exon was missed altogether altogether. The method detected the presence of a polyA tail as well. Arial Transposons are DNA sequences with the ability to jump about a host’s genome. Class I transposons use an RNA intermediate and are capable of copying themselves many times over. Retroviruses are believed to have evolved from this class of transposable elements. Class II transposons use a DNA intermediate and are restricted mainly to extracting themselves from one genomic location and reinserting themselves in another. MITEs are a type of class II transposon that has a very high copy number in plants, which is atypical for this class of transposons. transposons Arial Transposons can be broadly defined as DNA sequences flanked by repeats that have the ability to insert themselves throughout the genome. In some cases, transposableelement (TE) insertion can be faster than chromosome replication, allowing such sequences to rapidly accumulate within an organism’s genome. For example, 44% of the human genome is composed of sequences derived from TEs. Because transposons are able to occupy random positions within the genome, they can have deleterious effects by disrupting gene function; in some rare cases cases, a TE can even evolve into a beneficial gene. A TE can also alter gene expression by inserting itself near the regulatory region of a preexisting gene. In eukaryotic genomes with large amounts of noncoding DNA, TEs will mostly have a neutral effect. Transposable-element content varies greatly between eukaryotic species. Approximately one half of the human and maize genomes consists of transposable elements. Drosophila, on the other hand, has only 15% of its genome dedicated to TEs, and Arabidopsis, C. elegans, and S. cerevisiae all have less than 5%. TE content does not appear to be evenly distributed in genomes. For example, in humans, certain types of TE sequences are found in much higher concentrations in the X chromosome. Transposable elements can be classified as autonomous, nonautonomous, and inactive. Autonomous elements possess all of the genes that are required for transposition, which allows the sequence to move about the genome with only the help of host enzymes. Nonautonomous elements are degraded versions of autonomous sequences that require the proteins of autonomous elements for transposition. In contrast, inactive elements Arial Transposable elements can be further categorized as class I and class II. Two kinds of class I transposons are LTR and non-LTR retrotransposons. Class I transposable elements (also called retrotransposons, or retroelements) move via an RNA intermediate. Reverse transcriptase is then used to generate a cDNA sequence that can reinsert itself into the genome. This mode of action allows class I elements to copy themselves many times over, thereby significantly expanding an organism’s genome. A large portion (42%) of the human genome comes from retrotransposon sequences sequences. Interestingly, while a reversetranscriptase gene has been found in almost all eukaryotic genomes examined to date, it is found in only a minority of Bacteria and in almost no Archaea. Arial LTR retrotransposons are flanked by Long Terminal repeats that are identical at the time of insertion. The start and end of the LTR include a few inverted base pairs (sometimes not exact inversion). When the retroelement inserts in the host it generates a duplication of the host sequence, which can be use to identify pairs of LTRs from the same element. This is not part of the transposon and is characteristic of the insertion site. The region between the LTR code for the proteins. Arial Several features differentiate class II transposons from class I transposons. Class II transposons are able to move about the genome using DNA only, whereas class I transposons require an RNA intermediate to do so. Another major difference is that class II or DNA transposons are typically unable to copy themselves. A DNA transposon will extract itself from one region of the genome in order to insert itself into another. There are, however, some conditions under which a class II transposon can be replicated. For example if a class II transposon moves shortly after its sister DNA example, strand is copied, a second copy of the transposon will exist. There are approximately 200,000 copies of sequences derived from class II transposons in the human genome. P-elements in Drosophila and activation–dissociation (Ac/Ds) elements in maize are also examples of DNA-based transposons. The figure in the slide shows the structure of the Tc1-mariner class II transposons found in a variety of animal species. The transposase protein is solely responsible for the excision of the TE from one region and its integration into another. This protein is flanked by inverted terminal repeats and target-site duplications on each side. Arial Miniature inverted transposable elements (or MITEs) are an unusual group of TEs found in a variety i t off organisms. i They Th are especially i ll common iin plants l t ((e.g. 6% off th the rice i genome). ) For many years, MITEs were a mystery to biologists, because their high copy number implied active (or autonomous) elements, but none had been found. After the sequencing of the rice genome three separate groups of researchers published proof that MITEs can move about the rice genome. Plant MITEs fall into two categories: stowaway-like and tourist-like. Stowaway-like MITEs are moved about the genome by autonomous class II transposons similar to the TC1-mariner element. Tourist-like elements have as their autonomous partners a newly described class of active MITE called PIF/Pong. The figure in the slide compares the sequences of Pong and a nonautonomous MITE called Ping. The red and green regions represent homologous terminal regions among a variety of degraded Ping elements (mPings in the figure) and Pong elements. Unlike active Pong sequences, Ping elements have degraded ORF-1 genes. While the function of ORF-1 is not fully understood, the ORF-2 gene is believed to code for the transposase that is responsible for making Ping elements mobile. The high copy number of MITEs is very unusual for a class II transposon. The small size of MITEs (typically less than 500 bp) is believed to be the reason that these transposons are so common. Insertions of MITEs in the genome would presumably be less disruptive than insertions of larger TEs and hence would not be selected against as strongly. Science 2009 Vol. 325. no. 5946, pp. 1391 – 1394: We performed a genome-wide screen of functional interactions between Stowaway MITEs and potential transposases in the rice genome and identified a transpositionally active MITE that possesses key properties that enhance transposition The MITE contains internal sequences that enhance transposition. MITEs achieve high transposition activity by scavenging transposases encoded by di t tl related distantly l t d and d self-restrained lf t i d autonomous t elements. l t Arial Arial Arial Arial Transposons are DNA sequences with the ability to jump about a host’s genome. Class I transposons use an RNA intermediate and are capable of copying themselves many times over. Retroviruses are believed to have evolved from this class of transposable elements. Class II transposons use a DNA intermediate and are restricted mainly to extracting themselves from one genomic location and reinserting themselves in another. MITEs are a type of class II transposon that has a very high copy number in plants, which is atypical for this class of transposons. transposons Arial