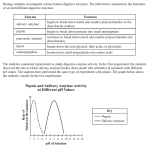
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Gene encoding the group B streptococcal protein R4, its
Clinical neurochemistry wikipedia , lookup
Community fingerprinting wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Paracrine signalling wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Multilocus sequence typing wikipedia , lookup
Gene nomenclature wikipedia , lookup
Biosynthesis wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Biochemistry wikipedia , lookup
Gene expression wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Expression vector wikipedia , lookup
Genetic code wikipedia , lookup
Metalloprotein wikipedia , lookup
Magnesium transporter wikipedia , lookup
Interactome wikipedia , lookup
Homology modeling wikipedia , lookup
Protein purification wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Point mutation wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Indian J Med Res 119 (Suppl) May 2004, pp 213-220 Gene encoding the group B streptococcal protein R4, its presence in clinical reference laboratory isolates & R4 protein pepsin sensitivity B.L. Smith*†, A. Flores*, J. Dechaine†, J. Krepela†, A. Bergdall† & P. Ferrieri* *Departments of Laboratory Medicine & Pathology & Pediatrics, University of Minnesota Medical School & Department of Biology, Luther College, USA † Received August 7, 2003 Background & objectives: R proteins were first identified by Lancefield in group B Streptococcus (GBS) as resistant to trypsin at pH8 and sensitive to pepsin at pH2. The R4 protein found predominantly in type III and some type II and V invasive isolates conforms to these criteria. The Rib protein, although structurally and epidemiologically similar to R4, was reported as resistant to both proteases. We report here the gene encoding the R4 protein from a type III group B streptococcal isolate (76-043) well characterized in our laboratory. Methods: Trypsin extracted GBS proteins were assayed for protease sensitivities by double-diffusion Ouchterlony using varying conditions for the enzyme pepsin. Standard haemoglobin assay was used to examine pepsin enzymatic activity. Thirty clinical isolates of varying protein profiles identified by double-diffusion from our reference strain laboratory were screened by PCR and Southern technique. SDS-PAGE gel purified R4 amino acid sequences were determined and used to design oligonucleotide primers for screening a 76-043 genomic library. Results: R4 was sensitive to pepsin at pH2 but appeared resistant at pH4, the reported pH used for Rib. By standard haemoglobin assay and trypsin extract studies of R4 protein, pepsin was shown to be active at pH2, yet easily inactivated; assays of GBS surface proteins are critical at pH2. Of the amino acids initially sequenced from R4, 88 per cent (61/69) showed identity to Rib; the r4 nucleotide sequence was identical to that of rib. All isolates with strong positive protein reactions for R4 were positive in both PCR and Southern technique, whereas isolates expressing alpha, beta, R1/R4, and R5 (BPS) protein profiles were not. Interpretation & conclusion: Sequenced PCR products aligned with identity to the R4 and Rib nucleotide sequences and confirmed the identity of these proteins and their molecular sequences. Key words a-like proteins - group B Streptococcus - pepsin sensitive - R4 - Rib - surface protein - trypsin resistant Group B streptococcus is a leading cause of infant mortality and studies of protein and genetic profiles are invaluable for detection and identification of virulent clinical isolates. Historically, proteins from Streptococcus were identified by double-diffusion Ouchterlony and their reactivity to the enzymes trypsin and pepsin. Lancefield’s classic experiments first identifying R proteins in group A Streptococcus (GAS), group B Streptococcus (GBS) and other streptococcal species defined R proteins as resistant to trypsin at pH8 and sensitive to pepsin at pH2. Although the first pepsin studies varied pH2-pH3 and pepsin concentration 0.5-1.0 per cent 1-4, standard protocols now test pepsin sensitivity at pH2 and 0.5 per cent pepsin5, 7 and characterization of various R proteins has continued in several laboratories6-8. The R4 protein which has been well characterized in our laboratory, conforms to these criteria, is found predominantly in type III as well as some type II and V invasive isolates that cause disease, and is useful in serotyping and characterization of invasive isolates5. Numerous studies 213 214 INDIAN J MED RES (SUPPL) MAY 2004 in our reference laboratory and elsewhere have well characterized the prevalence of R4 in invasive disease, but information on the gene sequence of R4 is not known. and the homology of the R4 and Rib amino acid and molecular sequences. Material & Methods Another GBS surface protein, Rib, was named primarily for its resistance to both proteases, and identified at that time as a novel GBS surface protein that conferred protective immunity9. Rib has been well characterized9-11. Both R4 and Rib are highly associated with type III and some type II GBS invasive strains, both demonstrate similarity in type III isolates of different geographic origin with some strain to strain variation5,7,9, and both R4 and Rib are trypsin resistant and highly repetitive in structure, but do not crossreact with alpha proteins7,9, yet we believed that Rib resembled R4 protein in more than simply structural and epidemiological association. In addition, Bevanger et al 6 using a monoclonal antibody raised against a purified R protein initially termed Ra, demonstrated that both R4 and Rib showed crossreactivity between reference GBS isolates and anti-R4 protein serum. This suggested that Rib was not only a member of the R protein family, but that Rib resembled R4, one of the alpha-like GBS surface proteins6. Although Lancefield initially believed that R proteins in GAS were not immunogenic3, more recent studies have shown immunogenicity of GBS R proteins in rabbits and in humans, thus both R4 and Rib are immunogenic12-14. Although serological crossreactivity was shown to exist between the R28 protein of GAS and Rib, Rib was not believed to be an R protein based on early data suggesting inability of R proteins to confer immunity and pepsin sensitivity results at pH49,15. Because the standard assay for pepsin sensitivity is performed at pH2, and Rib was identified as pepsin resistant at pH4, we predicted that R4 and Rib were the same. We have previously reported that R4 was sensitive to pepsin at pH2 but appeared resistant to pepsin at pH47. In this study, we sought to determine the nature of pepsin activity with respect to pepsin sensitivity testing of GBS surface proteins, and to determine if loss of pepsin activity at higher pH was responsible for R4’s apparent resistance to pepsin at pH4. We report that pepsin sensitivity testing of GBS surface proteins is critical at pH2. We also report the identification of the gene sequence for the R4 protein from a type III invasive isolate 76-043, the presence of the R4 gene in clinical reference laboratory isolates with R4 protein profiles, Bacterial isolates and growth: A variety of Streptococcus agalactiae isolates have been obtained from epidemiological studies spanning many years and 30 different GBS clinical isolates were obtained from the University of Minnesota Medical School GBS Reference Laboratory. Each previously serotyped isolate was assayed by double-diffusion Ouchterlony and identified by its hot HCl- and trypsin-extracted surface proteins from GBS5,7. Isolates were identified by serotype and by protein profile as R4, R1/R4, R5 (BPS), alpha, beta, or negative in which no associated protein was detected (Table I). A type III group B streptococcal isolate (76-043) identified with antisera in our reference laboratory and well characterized in our laboratory with respect to polysaccharide and protein profiles, opsonic requirements, and virulence in animal models of infection was extracted by both hot HCl- and trypsin-extracted methods5,7. Surface protein R4 was identified, isolated, and amino acid sequences and nucleotide sequences for R4 were determined from this isolate. S. agalactiae isolates were grown to stationary phase (THB OD600 ~0.5-0.7) in Todd-Hewitt Broth (THB) (Fisher, Pittsburg, PA) at 37oC without shaking. Escherichia coli strain DH5a used for transformation, XL1-Blue MRF’ used as the bacterial host strain for the lambda phage, and SOLR cells and ExAssist helper phage used for in vivo excision were provided by Stratagene (LaJolla, CA). E. coli was grown in Luria Broth (LB) at 37oC with aeration or on NZCYM media (Life Technologies, Grand Island, NY) as specified by Stratagene. Pepsin sensitivity testing of trypsinized extracts in immunodiffusion: Double diffusion Ouchterlony was performed as described previously using antiserum that detects R45,16. To examine R4 protein sensitivity to pepsin, Lancefield’s standard assay using trypsinized cell extracts was followed1,5: extraction of GBS isolate 76043 surface proteins with 0.1 per cent trypsin in Sorensen’s phosphate buffer/30 min/37oC, reaction stopped at pH6/ice/5 min, and extracts kept at -80oC until pepsin treatment. 1.0 per cent Pepsin (Sigma, USA) SMITH et al: SEQUENCE OF GBS PEPSIN-SENSITIVE R4 PROTEIN was prepared by resuspension in acidified water at varying pH. Equal volumes of GBS trypsinized cell extracts were incubated with 1.0 per cent pepsin (final concentration 0.5% pepsin), or were not treated with pepsin as controls maintaining equivalent volumes and protein concentrations with acidified water. Cell extracts tested in immunodiffusion included a non-pepsin/non-pH control, a non-pepsin/pH2 control, and incubation with pepsin at pH2, pH4, pH6, pH8. After incubation for 2 h at 37oC, the reaction was stopped at neutral pH and the extracts were kept at -80oC until ready for use in immunodiffusion. Standard haemoglobin assay: To examine pepsin enzymatic activity, a standard spectrophometric pepsin assay was used with haemoglobin as a substrate. Pepsin was dissolved in acidified water at the same varying pH conditions as above and incubated with 2.5 per cent (w/v) haemoglobin at 37oC for 10 min. After digestion, the resulting peptide fragments soluble in trichloroacetic acid (TCA) were quantified by A280 (Sigma, Communication) 17,18. Activity of pepsin was also observed after freeze-thaw and after irreversible denaturation of pepsin at pH > 819. Determination of R4 amino acid sequence: GBS trypsinized cell extracts were electrophoresed on 7.5 per cent SDS-PAGE under reducing conditions, and transferred and immobilized on PVDF membranes or maintained in SDS-PAGE gel slices and subjected to amino-terminal and internal peptide sequencing at Harvard Microchemistry Facility, Cambridge, MA. The N-terminal sequence and five internal amino acid sequences were obtained and a BLASTp search was performed using http://www.ncbi.nlm.nih.gov/blast/. Generation of R4 sequence-probe and screening of isolates by polymerase chain reaction and southern screening: From amino acid sequences determined above, oligonucleotide primers were derived from the amino-terminal sequence of R4, R4Nterm 5’ –ACAGTATTGAAGACAGATGGA- 3’, and internal peptide sequences designated R4peak31U 5’GTTTTGATAAGTTTAGAT-3’, R4peak31D 5’ – ATCTAAACTTATCAAAAC- 3’, and R4peak18 5'TTCTAAATTAGTAACTTCAGTTAT -3’ (Integrated DNA technologies, Inc., Coralville, IA) (Fig. 1). (2µM)Primers amplified 75ng of 76-043 genomic 215 DNA by PCR in a 100µl reaction: 1x buffer/ 2.5mM MgCl 2 / 0.1mM dNTP’s/ 2.4U taq polymerase (Promega, Madison, WI) at 94 oC for 5 min, 30 consecutive cycles at 94oC for 1 min, 40oC for 1 min, and 72oC for 1 min with a final extension 72oC for 7 min. The amplified products were cleaned by WizardTM PCR Preps DNA Purification System (Promega, Madison, WI), and sequenced by the University of Minnesota Microchemical Facility and Advanced Genetic Computing Facility. A 262bp digoxygeninlabeled r4 probe was generated by the addition of digoxygenin-11-dUTP (alkali labile) per manufacturer’s instructions using R4Nterm and internal R4peak18 primers. Probe concentration was assayed by dot blot and used to screen the library and to perform Southern blot using digoxygenin protocol per manufacturer’s instructions (Roche Molecular Biochemicals, Indianapolis, IN). Preparation of genomic DNA: Genomic DNA for PCR template DNA, Southern, and for construction of the genomic library was isolated from all GBS isolates using a method previously described20 and adapted for Qiagen DNeasyTM kit as per manufacturer’s protocols (Qiagen, Valencia, CA) or were phenol:chloroform extracted. For Southern blot screening of clinical isolates, approximately 1 µg of DNA from isolates 1-30 and positive control isolate 76-043 were digested either with EcoRV which cut 1x within the r4 gene sequence or HindIII which cut outside the r4 sequence. Construction and screening of GBS genomic library: A genomic library of randomly sheared 76-043 DNA in Lambda ZapII vector was created by Stratagene (Stratagene, LaJolla, CA). The digoxygenin-labeled 262bp r4 probe was used to screen replica filters of about 11,500 plaques from the library after incubation with XL1-Blue MRF’ bacterial host strain and plated on NZCYM bottom agar in 0.7 per cent top agarose as specified by Stratagene. Digoxygenin-detection protocol was followed as per manufacturer’s instructions (Roche Molecular Biochemicals, Indianapolis, IN). Plaques identified in the initial screening were re-hybridized with the 262bp r4 probe in a secondary screen. Plaques with the strongest signal were chosen for in vivo excision of the pBluescript SK(-) phagemid by co-infection with helper phage according to procedure provided by Stratagene. INDIAN J MED RES (SUPPL) MAY 2004 216 Results Because of similarities between R4 and Rib, and the classification of Rib as a pepsin resistant protein at pH4, it was important to first study the activity of pepsin in relation to pepsin sensitivity testing; second, and more importantly, to determine the gene sequence encoding the R4 protein; and third to determine if the gene sequence could be detected only in isolates previously determined to be R4 expressing isolates. Pepsin sensitivity of trypsin-extracted proteins and pepsin activity by standard haemoglobin assay: Because pepsin is an enzyme with an optimal pH range for activity, above which the enzyme is denatured18,19,21, we examined if the loss of pepsin activity at higher pH was responsible for R4’s apparent resistance to pepsin at pH4. Here we directly examined pepsin sensitivity and pepsin activity concurrently using two assays. First, we tested trypsin-extracted R4 protein under various pH conditions for pepsin sensitivity using two different prototypic reference antisera for R4 in double-diffusion (results not shown). The trypsin-extracted R4 showed a precipitin result with anti-R4 antiserum for both controls. As the classic example of pepsin sensitivity for pepsin at pH2, no precipitin result was shown. At pH4, pH6 and pH8, however, precipitin results were positive confirming our previous results that R4 appears resistant to pepsin at higher pH. In order to test directly the activity of the pepsin used in these experiments, a second assay was used. We tested pepsin activity using a standard haemoglobin assay17 that determined pepsin enzymatic activity under these different conditions. We first examined pepsin digestion of haemoglobin at pH2, pH > 8, and pH > 8 returned to pH2. A280 measurements were significantly decreased with A 280 =0.198, A 280 =0.003, and A280=0.000, respectively, indicating that pepsin was irreversibly denatured above pH8 in concordance with other studies of pepsin activity18,19,21. In addition, we observed that with repeated freeze-thaw of the pepsin, loss of pepsin activity occurred (data not shown). To examine directly pepsin activity of the acidified pepsin used in the R4 pepsin sensitivity tests, we then employed the haemoglobin assay with these pepsin reagents. In duplicate experiments (Fig.2), pepsin activity was highest at pH2 and pepsin activity was significantly decreased for pepsin at pH4, pH6, and pH8. R4 amino acid and nucleotide sequence determination: To determine if R4 and Rib were the same, it was necessary to obtain amino acid and nucleotide sequences for R4. SDS-PAGE gel purified R4 protein was sequenced and N-terminal and internal amino acid sequences were aligned by BLAST to the Rib protein. Six different amino acid sequences were obtained by MALDI-mass spectrophotometry: Nterm, peak31, peak18, peak 28, peak 23, and peak15. Of the 69 amino acids sequenced from R4, 61 showed identity (88%) to the Rib amino acid sequence. The sequences highlighted in Fig.1 indicate identity and alignments. Amino acid sequences were used to design PCR oligonucleotide primers as stated above. The amplified product of R4Nterm/R4peak31D was 181bp; the amplified product of R4peak31U/R4peak18 was 81bp and the amplified product of R4Nterm/R4peak18 primers was 262bp. These products were sequenced and aligned with the rib nucleotide sequence in size and order. Primers R4Nterm and R4peak18 shown in Fig.1 were used to generate the 262bp digoxygenin-labeled r4 PCR product used to screen the 76-043 GBS genomic library. Using the R4 protein-generated probe, two clones were obtained and used for sequencing the r4 gene. The entire open reading frame for the gene encoding R4 was subcloned. The resulting clones were sequenced from the 5’ nucleotide sequences encoding the N-terminus through the first three repeats, internal identical tandem repeats of 237bp, through the last three repeats, and the 3’ nucleotide region encoding the carboxyl terminus. The predicted translations were consistent with the known amino acid sequence of the 120kDa R4 protein containing approximately 12 tandem repeats (Fig.1). Although the sequence was not determined for all internal DNA repeat regions, the repeats of 237bp appeared to be identical tandem repeats, consistent with other ladder-like proteins, and were identical to the repeats found in the gene encoding Rib10. Using NCBI BLASTn, r4 nucleotide sequences were 99 per cent identical (3682/3693) to the gene encoding Rib. A single nucleotide difference in 11 of the 12 repeats of the rib gene was the sole difference. The r4 nucleotide sequence was highly homologous (3576/3577) to the recently published genome of a type V/R GBS strain. The r28 gene sequence encoding the R protein of GAS, as well as gene sequences for other alpha-like proteins also showed homology. Little homology was detected with a GBS genome from a nonR4 producing type Ia strain. Fig.1 shows the deduced SMITH et al: SEQUENCE OF GBS PEPSIN-SENSITIVE R4 PROTEIN 217 Table. Isolate 76-043 and 30 different GBS clinical isolates were identified by their double-diffusion protein profiles, verified by PCR using R4-generated oligonucleotide primers and sequence identity to the r4 probe, and confirmed by Southern blot detection using the digoxygenin-labeled R4-generated probe Isolate No. Serotype/protein profile PCR results Sequence results Southern 76-043 III/R4 + + + 1 V/c (a only) - ND - 2 V/R5 - ND - 3 Ib/c (b only) - ND - 4 Ib/c (a and b) - ND - 5 V/R4 (weak) - ND - 6 III/R4 + + + 7 III/R4 + + + 8 III/R4 + + + 9 III/R4 + + + 10 V(R1/R4) - ND - 11 V/negative - ND - 12 III/R4 + + + 13 Ia/c (a only) - ND - 14 V/negative - ND - 15 II/R4 + + + 16 Ib/c (a and b) - ND - 17 III/R4 + + + 18 II/c (a only) - ND - 19 III/R4 + + + 20 Ia/negative - ND - 21 III/R4 + + + 22 V/ (R1/R4) - ND - 23 III/R4 + + + 24 Ia/c (a only) - ND - 25 III/R4 + + + 26 III/R4 + + + 27 Ia/c (a only) - ND - 28 V/negative - ND - 29 III/R4 + + + 30 V/R4 + + + PCR and Southern results were done in replicates and duplicates, respectively. Sequences were determined only for samples that yielded a PCR product. ND, not determined. 218 INDIAN J MED RES (SUPPL) MAY 2004 Fig.1. Amino acid and nucleotide sequence of R4. The nucleotide sequence of R4 is shown with its translated amino acid sequence. Highlighted amino acid sequences were obtained by amino acid sequencing of R4 protein and aligned with identity to Rib protein sequences. Sequences for the N-terminus and peak 18 were used to design the primers to create the R4 digoxygenin- labeled PCR probe for screening of the genomic library of isolate 76-043 and for PCR and Southern screening of clinical isolates. (R4 Genbank Accession #AY179867). SMITH et al: SEQUENCE OF GBS PEPSIN-SENSITIVE R4 PROTEIN nucleotide sequence, the translated R4 protein and amino acid sequences used to generate primers (R4 Genbank Accession # AY179867). PCR, sequencing, and southern screening of 30 different isolates using R4-generated primers and probe: As a final determination that R4 and Rib were the same and that clinical isolates identified as R4 isolates could be detected using the R4-generated probe, PCR and Southern blot were used to screen 30 clinical isolates of varying protein profiles as identified by doublediffusion from a reference laboratory. All isolates with strong positive protein reactions for R4 were positive by PCR, whereas isolates expressing alpha, beta R1/R4 and R5 (BPS) protein profiles were not detected. In addition, R4 producing serotypes II, III and V were all detected. Products were consistent in size with expected r4 and rib sequences. Positive reactions with PCR products were sequenced and aligned with identity to the r4 and rib sequences. In Southern blot, the R4-generated digoxygenin probe hybridized to EcoRV digested genomic DNA as two bands as predicted based on sequence restriction digest patterns for all PCR-positive/R4-positive isolates. This confirmed that oligonucleotides generated from the R4 protein in PCR, and the R4-generated probe in Southern identified only R4-positive isolates from the reference laboratory isolates (Table I). Discussion The R4 protein has been identified and characterized as a pepsin sensitive protein. R4 is similar to Rib epidemiologically, in size and in ladder-like repeats. Both are trypsin resistant, do not crossreact with alpha, but Fig.2. Pepsin activity tested by standard haemoglobin test. Activity of pepsin at varying pH used concurrently in pepsin sensitivity tests were tested by standard haemoglobin assay n=2, duplicates done on separate days with no deviation P<0.05. 219 crossreact with each other and with anti-R4 antisera, and are immunogenic, yet the Rib protein was described as pepsin resistant at pH4. We previously reported R4 to be sensitive to pepsin at pH2 but appeared resistant to pepsin at pH47. Therefore, we examined pepsin enzymatic activity using a standard haemoglobin assay correlating it with pepsin sensitivity tests of R4. By Lancefield’s standard trypsin extract studies, we determined that R4 was pepsin sensitive at pH2 and appeared resistant at pH4, pH6, and pH8 likely due to corresponding loss of pepsin activity at these higher pH values. In concurrent tests, the pepsin used in the sensitivity tests was determined to be inactive under various conditions. By standard haemoglobin assay, pepsin was shown to be active at pH2, yet inactive at higher pH, inactive if irreversibly denatured at high pH but restored to pH2, and decreased in activity with repeated freeze-thaw usage. This supported our results observed in double-diffusion pepsin sensitivity tests, revealing that pepsin was inactive at pH4, and not that R4 or Rib were pepsin resistant at this pH. These results indicated that careful handling of pepsin is critical; fresh pepsin, careful pH methods, and pH2 should be used with each experiment to prevent irreversible denaturation and promote optimal pepsin activity. Together these results demonstrated that R4 was pepsin sensitive at pH2, and that R4 and Rib appeared to be resistant due to loss of pepsin activity at pH4. This proved to play an important role in our later determination of R4 and Rib similarities. After determining the gene and amino acid sequence of R4, confirming R4 sensitivity to pepsin at pH2 and examining loss of pepsin activity with increasing pH, our prediction that R4 and Rib protein are the same was confirmed. The R4 protein amino acid and nucleotide sequence was identified and aligned with identity to the amino acid and nucleotide sequence of the Rib protein, thereby confirming identity. It is of interest to note that previous epidemiological studies of R4 as well as the current PCR and Southern screening of clinical isolates confirmed that R4 was found in serotypes II, III and V, and that identity between the r4 gene of a type III isolate in this study and the recently published genome of a type V isolate further supported these data, whereas little homology was detected in a type Ia genome. R4-generated primers yielded PCR products while the R4-generated digoxygenin-labeled probe detected INDIAN J MED RES (SUPPL) MAY 2004 220 only R4-positive isolates in Southern blot, and not isolates with alpha, beta, R1/R4 or R5 (BPS) proteins. PCR product sequences were identical to nucleic acid sequences encoding R4 and Rib. It is interesting to note that some isolates serotyped as R1/R4 were not detected by the r4 probe. Further sequence determination of the R1 protein and its relationship to R4 are needed. One cannot exclude that R1, when expressed with R4, is a dual protein, but with epitopic distinction from R4. Sequencing of R4 from strains expressing both R1 and R4 might elucidate why these strains were PCR negative. After many years of inference that R4 and Rib proteins were identical, elucidation of the nucleotide sequence verifies that these two genes and proteins are the same. Much valuable reference information is known for R4 and many valuable immunogenicity and protectivity studies have been done for Rib. Taken together, much more is now known for the R4/Rib protein. Acknowledgment This work was partially funded by Minnesota Medical Foundation grant and by the NIH training grant in Pediatric Infectious Diseases (HD07381). Authors thank Luther College Honors Research students for PCR screening studies and for travel support provided by Luther College. References 1. Lancefield R. Studies on the antigenic composition of group A hemolytic streptococci. J Exp Med 1943; 78 : 465-76. 2. Lancefield R, Dole V. The properties of T antigens extracted from group A hemolytic streptococci. J Exp Med 1946; 84 : 449-71. 3. Lancefield R, Perlmann G. Preparation and properties of a protein (R Antigen) occurring in streptococci of group A, type 28 and in certain streptococci of other serological species. J Exp Med 1952; 96 : 83-97. 7. Flores AE, Ferrieri P. Molecular diversity among the trypsin resistant surface proteins of group B streptococci. Zbl Bakt 1996; 285 : 44-51. 8. Wilkinson H. Comparison of streptococcal R antigens. Appl Microbiol 1972; 24 : 669-70. 9. Stålhammer-Carlemalm M, Stenberg L, Lindahl G. Protein Rib: a novel group B streptococcal cell surface protein that confers protective immunity and is expressed by most strains causing invasive infections. J Exp Med 1993; 177 : 1593-603. 10. Wastfelt M, Stålhammer-Carlemalm M, Dellisse A, Cabezon T, Lindahl G. Identification of a family of streptococcal surface proteins with extremely repetitive structure. J Biol Chem 1996; 271 : 18892-7. 11. Larsson C, Stålhammer-Carlemalm M, Lindahl G. Experimental vaccination against group B Streptococcus, an encapsulated bacterium, with highly purified preparations of cell surface proteins Rib and a. Infect Immun 1996; 64 : 3518-23. 12. Fasola EL, Flores AE, Ferrieri P. Immune responses to the R4 protein antigen of group B streptococci and its relationship to other streptococcal R4 proteins. Clin Diagn Lab Immunol 1996; 3 : 321-5. 13. Hordnes K, Tynning T, Kvam AI, Bevanger L, Brown TA, Jonsson R, et al. Cervical secretions in pregnant women colonized rectally with group B streptococci have high levels of antibodies to serotype III polysaccharide capsular antigen and protein R. Scand J Immunol 1998; 47 : 179-88. 14. Linden V, Kvist K, Christensen P. Correlation between low levels of maternal IgG antibodies to R protein and neonatal septicemia with group B streptococci carrying R protein. Int Arch Allergy Appl Immun 1983; 71 : 168-72. 15. Stålhammer-Carlemalm M, Areschoug T, Larsson C, Lindahl G. The R28 protein of Streptococcus pyogenes is related to several group B streptococcal surface proteins, confers protective immunity and promotes binding to human epithelial cells. Mol Microbiol 1999; 33 : 208-19. 16. Wilkinson H, Moody M. Serological relationships of type I antigens of group B streptococci. J Bacteriol 1969; 97 : 629-34. 17. Anson ML. The estimation of pepsin, trypsin, papain, and cathepsin with hemoglobin. J Gen Physiol 1938; 22 : 79-89. 18. Ryle A. The porcine pepsins and pepsinogens. Biochem J 1966; 98 : 485-7. 4. Lancefield R. Differentiation of group A streptococci with a common R antigen into three serological types, with special reference to the bactericidal test. J Exp Med 1957; 106 : 525-44. 19. Richter S, Tanaka T, Yada R. Mechanism of activation of the gastric aspartic proteinases: pepsinogen, progasticsin, and prochymosin. Biochem J 1998; 335 : 481-90. 5. Flores A, Ferrieri P. Molecular species of R-protein antigens produced by clinical isolates of group B streptococci. J Clin Microbiol 1989; 27 : 1050-4. 20. Suvorov AN, Flores AE, Ferrieri P. Cloning of the glutamine synthetase gene from group B streptococci. Infect Immun 1997; 65 : 191-6. 6. Bevanger L, Kvam A, Maeland J. A Streptococcus agalactiae R protein analysed by polyclonal and monoclonal antibodies. APMIS 1995; 103 : 731-6. 21. Cottrell T, Harris L, Tanaka T, Yada R. The sole lysine residue is porcine pepsin works as a key residue for catalysis and conformational flexibility. Glycobiology 1995; 270 : 19974-8. Reprint requests: Dr B.L. Smith, Department of Pediatrics, University of Minnesota, 420, Delaware St SE No.196 Minneapolis MN 55455, USA e-mail: [email protected]