* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Statistics Workshop Introduction to statistics using R Tarik C. Gouhier June 17, 2013
Survey
Document related concepts
Psychometrics wikipedia , lookup
Bootstrapping (statistics) wikipedia , lookup
Linear least squares (mathematics) wikipedia , lookup
Degrees of freedom (statistics) wikipedia , lookup
Analysis of variance wikipedia , lookup
Foundations of statistics wikipedia , lookup
Transcript
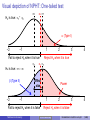
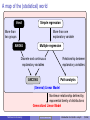
Statistics Workshop Introduction to statistics using R Tarik C. Gouhier [email protected] Department of Marine and Environmental Sciences Northeastern University June 17, 2013 Statistical distributions Statistical distributions are characterized by their moments, which quantify location and dispersion: Location anchors the distribution Location: E[X] Dispersion measures the spread The first moment is the expected value P or mean E[X] = N1 N i=1 Xi The second central moment is thei h variance Var[X] = E (X − E[X])2 Dispersion The nth central moment is E (X − E[X])n −2 The standard deviation σX = Northeastern University √ −1 0 X 1 2 Var[X] Statistics workshop Introduction to statistics using R (3/66) Bivariate expectations: covariance and correlation Covariance measures the covariation between two variables: Cov[X, Y] = E [(X − E[X]) (Y − E[Y])] Note that the variance is merely the covariance between X and X Pearson product-moment correlation is the standardized covariance between two variables: ρ(x, y) = Cov[X][Y] E [(X − E[X]) (Y − E[Y])] = σX σY σX σY With −1 ≤ ρ(x, y) ≤ 1 In R, univariate and bivariate expectations of variables x and y can be computed via mean(x), var(x), sd(x), cov(x,y), cor(x,y) and cor.test(x,y) Northeastern University Statistics workshop Introduction to statistics using R (4/66) The Central Limit Theorem The sum or mean of a set of independent random variables drawn from the same statistical population (i.e., identically and independently distributed) will be normally distributed regardless of their parent distribution See web demo I built using R: http://spark.rstudio.com/synchrony/climit Northeastern University Statistics workshop Introduction to statistics using R (5/66) Neyman-Pearson Hypothesis Testing (NPHT) Embraces Karl Popper’s perspective that theories/hypotheses should be falsifiable in order to be regarded as scientific Test a null hypothesis H0 vs. alternate hypothesis Ha Compute the p-value: probability of obtaining a test statistic at least as extreme as the one observed assuming that H0 is true Reject H0 hypothesis if the p-value is smaller than a predetermined significance level α and accept Ha Note: failure to reject H0 does not mean your accept H0 Process leads to four possible outcomes: H0 is true H0 is false Northeastern University Reject H0 Type I error (α) Correct (1 − β) Statistics workshop Fail to reject H0 Correct (1 − α) Type II error (β) Introduction to statistics using R (7/66) Visual depiction of NPHT: Two-tailed test αc H0 is true: µT = µC µC µT αc α 2 (Type I) −2 α 2 (Type I) −1 Fail to reject H0 1 2 3 4 when it is true Reject H0 when it is true µC µT αc αc Ha is true: µT ≠ µC β (Type II) Effect size Power −2 −1 Fail to reject H0 when it is true Northeastern University Power 1 2 3 4 Reject H0 when it is false Statistics workshop Introduction to statistics using R (8/66) Visual depiction of NPHT: One-tailed test H0 is true: µT ≤ µC µC αc µT α (Type I) −2 −1 1 2 3 4 3 4 Fail to reject H0 when it is true Reject H0 when it is true µC αc µT Ha is true: µT > µC β (Type II) −2 Effect size −1 1 Fail to reject H0 when it is false Northeastern University Power 2 Reject H0 when it is false Statistics workshop Introduction to statistics using R (9/66) NPHT in R p-value: area under the curve Right or upper tail P(X > x): 1-pf(fval, df1, df2) Left or lower tail P(X ≤ x): pf(fval, df1, df2) Critical value at α level: Right or upper tail: qf(1-alpha/2, df1, df2) Left or lower tail: qf(alpha/2, df1, df2) Power at α level: conduct power analysis using power.anova.test, power.t.test by specifying any four of the following parameters Effect size Sample size Variance α Power (1 − β) Note: The example above was provided for the F-distribution. Similar functions are available for many other distributions (?Distributions) Northeastern University Statistics workshop Introduction to statistics using R (10/66) Output from NPHT NPHT generates three types of metrics: 1 2 3 p-value: statistical significance Effect size: a measure of biological significance Absolute: µT − µC in treatment units µ −µ Relative: 100 TµC C µ −µ Normalized: d = T σ C to allow comparisons across studies Explanatory power: proportion of variation explained (e.g., R2 , η2 ) Many ecological and evolutionary studies have very low explanatory power and biological significance but high statistical significance (Moller and Jennions, 2002) This is partly because ANOVA tables generally only present p-values Many are calling for a shift in focus from statistical significance (p-values) to biological significance (effect size; e.g., Cohen, 1988) Northeastern University Statistics workshop Introduction to statistics using R (11/66) Notes about the p-value Lower p-values don’t necessarily mean stronger results because the p-value is a complex combination of sample size and effect size Trivial to get a significant p-values with gigantic datasets, a big issue in bioinformatics Use various correction factors to reduce false positives (type I error) due to multiple tests Choice of critical level in biology α = 0.05 is completely arbitrary: “[...] surely, God loves the 0.06 nearly as much as the 0.05" (Rosnow and Rosenthal, 1989) Selecting proper α depends on question: is it more important to reduce false positives (type I error) or false negatives (type II error)? Northeastern University Statistics workshop Introduction to statistics using R (12/66) A map of the (statistical) world t-test Simple regression More than two groups More than one explanatory variable ANOVA Multiple regression Discrete and continuous explanatory variables ANCOVA Relationship between explanatory variables Path analysis (General) Linear Model Nonlinear relationship defined by exponential family of distributions Generalized Linear Model Northeastern University Statistics workshop Introduction to statistics using R (14/66) General Linear Model General formula 1 x11 x12 y1 y2 = 1 x21 x22 .. .. .. .. . . . . 1 xN1 xN2 yN Y = X . . . x1K 1 β0 . . . x2K β1 2 + × .. . .. .. .. . . . N βK . . . xNK × β + Where: Y: vector of N observations from response or dependent variable X: N × K + 1 design matrix containing K explanatory or independent variables and a column for the intercept β: vector of K coefficients + 1 intercept ∼ N(0, σ2 ): vector of N identically and independently distributed residuals Northeastern University Statistics workshop Introduction to statistics using R (15/66) General Linear Model assumptions General formula 1 x11 x12 y1 1 x y x22 21 2 .. = .. .. .. . . . . 1 xN1 xN2 yN Y = X β0 . . . x1K 1 . . . x2K β1 2 .. × .. + .. .. . . . . βK . . . xNK N × β + Structural assumptions: 1 Y is a single continuous response variable 2 X contains 1 to K continuous or discrete variables 3 Y are X are linearly related Data assumptions: 1 Normality: ∼ N(0, σ2 ): 2 Independence: Cov[i , j ] = 0 3 Homoscedasticity: σ2i = σ2j 4 Non-collinearity: Cov[xi , xj ] = 0 Northeastern University Statistics workshop Introduction to statistics using R (16/66) General Linear Model as a unified framework General formula 1 x11 x12 y1 y2 = 1 x21 x22 .. .. .. .. . . . . 1 xN1 xN2 yN Y = X . . . x1K 1 β0 . . . x2K β1 2 + × .. . .. .. .. . . . N βK . . . xNK × β + If X matrix contains only continuous variables (i.e., covariates), then this is called regression If X matrix contains only discrete variables (i.e., factors), then this is called ANanalysis Of VAriance (ANOVA) If X matrix contains a mixture of discrete and continuous variables, then this is called ANanalysis of COVAriance (ANCOVA) Northeastern University Statistics workshop Introduction to statistics using R (17/66) Solving the General Linear Model We want to find the suite of parameter estimates β̂ that minimizes the sum 2 of squared residuals 2 = Y − Xβ̂ : Expand 2 : 2 = Y0 Y − 2X0 Yβ̂ + X0 Xβ̂2 Take the (partial) derivative with respect to β and set it to zero to find the minimum: ∂ 2 = −2X0 Y + 2X0 Xβ̂ = 0 ∂β Rearrange the terms to isolate β̂: β̂ = X0 X −1 X0 Y Note: X is a matrix so we need to transpose (X0 ), (matrix) multiply (X0 X) and then take the inverse (X0 X)−1 Northeastern University Statistics workshop Introduction to statistics using R (18/66) Fitting the General Linear Model in R R fully embraces the unified nature of the General Linear Model framework by using a single function lm to fit regressions, ANOVA and ANCOVA: Fitting via lm lm( Y = X×β + y ∼ x , data = my.data) If x contains only continuous variables (i.e, numeric): regression If x contains only discrete variables (i.e., factor): ANOVA If x contains both discrete and continuous variables: ANCOVA Utility functions for lm objects: summary(lm.fit): produces table of coefficients, R2 , p-values coef(lm.fit): produces table of coefficients anova(lm.fit): produces ANOVA table plot(lm.fit): diagnostic plots Northeastern University Statistics workshop Introduction to statistics using R (19/66) The t-test 1-sample t-test: does the mean x̄ of N samples differ from x̄0 ? Null hypothesis H0 : x̄ = x̄0 ; Alternate hypothesis Ha : x̄ , x̄0 t= x̄ − x̄0 √ s/ N Where s is the unbiased sample mean: s s= PN − x̄)2 N−1 i=1 (xi The p-value of t is obtained using a t-distribution with N − 1 degrees of freedom (2*(1-pt(t, N-1))) The (1 − α)% confidence intervals around the mean are: s CIx̄ = x̄ ± t α2 ,N−1 √ N One-tailed tests are also possible, but you must specify the direction of the the test ahead of time. Northeastern University Statistics workshop Introduction to statistics using R (20/66) One-sample t-test in R t.test(x, alternative="two.sided", mu=mu, conf=1-alpha): two-tailed test (H0 : x̄ = µ) t.test(x, alternative="less", mu=mu, conf=1-alpha): one-tailed test (H0 : x̄ > µ) t.test(x, alternative="greater", mu=mu, conf=1-alpha): one-tailed test (H0 : x̄ < µ) # Random values from normal distribution with mean=5 # and sd=1 x = rnorm(10, mean = 5) # two-tailed test at alpha = 0.01 t.test(x, mu = 4.3, conf = 0.99)$p.value ## [1] 0.03639 # one-tailed test at alpha = 0.01 t.test(x, mu = 4.3, conf = 0.99, alt = "greater")$p.value ## [1] 0.01819 Northeastern University Statistics workshop Introduction to statistics using R (21/66) Two-sample t-test 2-sample t-test, unequal sample size and equal variance: does the mean x¯1 of N1 samples from group 1 differ from the mean x̄2 of N2 samples from group 2? Null hypothesis H0 : x̄1 = x̄2 ; Alternate hypothesis Ha : x̄1 , x̄1 t= x̄1 − x̄2 q sx1 x2 N11 + 1 N2 Where sx1 x2 is the pooled standard deviation: s sx1 x2 = (N1 − 1) s2x1 + (N2 − 1) s2x2 N1 + N2 − 1 The p-value of t is obtained using a t-distribution with N1 + N2 − 1 degrees of freedom (2*(1-pt(t, N1+N2-1))) Similar t-tests exist for unequal variance and paired values. They are all performed using the same t.test function in R. Northeastern University Statistics workshop Introduction to statistics using R (22/66) Two-sample t-test in R t.test(x, alternative="two.sided", mu=mu, conf=1-alpha): two-tailed test (H0 : x̄ = µ) t.test(x, alternative="less", mu=mu, conf=1-alpha): one-tailed test (H0 : x̄ > µ) t.test(x, alternative="greater", mu=mu, conf=1-alpha): one-tailed test (H0 : x̄ < µ) # Random values from normal distribution with mean=5 # and sd=1 x = matrix(rnorm(20, mean = 5), ncol = 2) # two-tailed test assuming equal variance t.test(x[, 1], x[, 2], var.equal = TRUE)$p.value ## [1] 0.2548 # two-tailed test with paired samples across groups t.test(x[, 1], x[, 2], paired = TRUE)$p.value ## [1] 0.3222 Northeastern University Statistics workshop Introduction to statistics using R (23/66) ANOVA ANOVA is an extension of two-sample t-test for comparing the mean of more than two groups (e.g., multiple levels within a factor or multiple levels across factors), where: a factor is a categorical variable describing a type of treatment (e.g., temperature, precipitation) a level of a factor represents the “magnitude" of that factor (e.g., 5o C vs. 20o C or low vs. high) Northeastern University Statistics workshop Introduction to statistics using R (24/66) One-way ANOVA General formula yij = µ + αi + ij Or equivalently: yij = µi + ij Where: yij : vector of observations j ∈ [1, . . . , N] and group i ∈ [1, . . . , L] µ: grand mean across all observations j irrespective of group i αi : deviation from the grand mean for group i µi : mean of group i ij ∼ N(0, σ2 ): vector of identically and independently distributed errors Northeastern University Statistics workshop Introduction to statistics using R (25/66) One-way ANOVA as a multiple regression problem General formula y1 y2 = .. . yN Y = group 1 group 2 ... group K−1 x11 x21 .. . x12 x22 .. . ... ... .. . x1K−1 x2K−1 .. . xN1 xN2 ... xNK−1 X 1 α1 α 2 2 + × .. .. . . N αK−1 × α + Where: Y: vector of N observations each belonging to one of K groups X: N × K − 1 design matrix containing K − 1 dummy variables xij . If observation i belongs to group j, xij = 1, otherwise xij = 0 α: deviations from baseline group (intercept) for each group ∼ N(0, σ2 ): vector of N identically and independently distributed residuals Northeastern University Statistics workshop Introduction to statistics using R (26/66) ANOVA table calculation Null hypothesis: α1 = α2 = . . . = αk = 0 Test is based on F-test with a full model vs. reduced model (with just an intercept) To determine which pairs of groups are different, need to perform post-hoc pairwise comparisons Source df Sum of Squares Treatment L−1 SST = PL Error N−1 SSE = PL Total N−1 − µ)2 MST = SST L−1 − 1) s2i 2 PL Pni y − µ ij i=1 j=1 MSE = SSE N−1 Here, the omnibus F-test is F = F > F1−α,L−1,N−1 Northeastern University Mean Squares i=1 ni (ȳi i=1 (ni MST and we can reject H0 at level α if MSE Statistics workshop Introduction to statistics using R (27/66) Visual depiction of ANOVA y1 Grand mean: µ SSE SST (n1 − 1)s21 µ α1 = n1(y1 − µ)2 (n2 − 1)s22 y2 Group 1 α2 = n2(y2 − µ)2 Group 2 Want to maximize SST (deviations of group means from grand mean) and minimize SSE (deviations of observations from their group mean) F-value is ratio of SST normalized by the number of groups (i.e., MST or average deviation of group means from grand mean) and SSE normalized by the number of observations (i.e., MSE or average deviation of all observations from their group mean) Northeastern University Statistics workshop Introduction to statistics using R (28/66) N-way ANOVA N-way ANOVA is simple extension of One-way ANOVA where groups correspond to different levels across several factors: General formula yijk = µ + αi + βj + (αβ)ij + ijk Where: yijk : vector of observations k ∈ [1, . . . , N] belonging to level i ∈ [1, . . . , I] of factor I and level j ∈ [1, . . . , J] of factor J µ: grand mean across all observations j irrespective of factor i αi : deviation from the grand mean for level i of factor I βj : deviation from the grand mean for level j of factor J (αβ)ij : deviation from the grand mean for level i of factor I and level j of factor J ijk ∼ N(0, σ2 ): vector of identically and independently distributed errors Northeastern University Statistics workshop Introduction to statistics using R (29/66) N-way ANOVA and interactions With One-way ANOVA, focus is on the additive or independent effects of several (>2) levels within a single factor In 2+ way ANOVA, several factors can interact and lead to non-additive effects Sub-additive or antagonistic effects occur when two or more factors generate a lower response than expected based on their independent effects Super-additive or synergistic effects occur when two or more factors generate a higher response than expected based on their independent effects Overall two factors A and B interact if their effects are contingent: i.e., the effect of factor A on the group means depends on the level of factor B When two factors interact, forgo the analysis of their independent effects since they are not consistent and focus on their interactive effects Northeastern University Statistics workshop Introduction to statistics using R (30/66) Visual depiction of interactions A and B significant A x B interaction not significant A and B significant A x B interaction significant A− A+ B− A− A+ B+ B− B+ In the first panel, the effects of the “+" level of factor A on the group means is consistent across the levels of factor B and vice versa In the second panel, the effects of the “+" level of factor A on the group means depends on the levels of factor B and vice versa Northeastern University Statistics workshop Introduction to statistics using R (31/66) N-way ANOVA in R lm(y ∼ x1 + x2 + x3, data=d): 3-way ANOVA looking at only additive effects using dummy variables. This will look like a multiple regression with the intercept being the grand mean, and the the slope of each group representing the αi deviation from the grand mean. lm(y ∼ x1*x2*x3, data=d): 3-way ANOVA with all possible two-way and three-way interactions. Good luck with that! lm(y ∼ x1*x2*x3 - x2:x3, data=d): 3-way ANOVA with all possible interactions except for x2:x3 lm(y ∼ x1:x2, data=d): only look at interaction between x1 and x2 (no additive effects) summary(lm(y ∼ x, data=d)): get table of coefficients, R2 , p-values etc... anova(lm(y ∼ x, data=d)): produce ANOVA table from lm fit aov(y ∼ x, data=d): standard ANOVA table directly without going through lm Northeastern University Statistics workshop Introduction to statistics using R (32/66) Pairwise comparisons and multiple test corrections To determine which pairs of groups are significantly different from each other, we need to perform pairwise comparisons However, multiple tests will inflate your type I error beyond the desired α level Many ways of preventing inflation of type I error, including bonferroni correction, sequential bonferroni (aka Holm), FDR, etc... These range from ultra-conservative (sacrifice power or increase type II error but prevent type I error inflation) such as bonferroni correction to relatively non-conservative approaches (e.g., Tukey’s HSD provided by function TukeyHSD) All of these methods are available in function p.adjust Note: correcting for multiple tests is extremely controversial (Garcia, 2004; Gotelli and Ellison, 2004). For instance Gotelli and Ellison (2004) suggest not correcting until forced to do so by reviewers! Northeastern University Statistics workshop Introduction to statistics using R (33/66) Pairwise comparisons and multiple test corrections in R require(faraway, quiet = TRUE) data(package = "faraway", fruitfly) summary(mod.anova <- aov(longevity ~ activity, data = fruitfly)) ## ## ## ## ## Df Sum Sq Mean Sq F value Pr(>F) activity 4 12269 3067 14.3 1.4e-09 *** Residuals 119 25476 214 --Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 TukeyHSD(mod.anova)$activity ## ## ## ## ## ## ## ## ## ## diff one-isolated 1.2400 low-isolated -6.8000 many-isolated 0.9817 high-isolated -24.8400 low-one -8.0400 many-one -0.2583 high-one -26.0800 many-low 7.7817 Northeastern University -18.0400 high-low lwr upr -10.224 12.704 -18.264 4.664 -10.601 12.564 -36.304 -13.376 -19.504 3.424 -11.841 11.324 -37.544 -14.616 -3.801 19.364 Statistics workshop -29.504 -6.576 p adj 9.982e-01 4.731e-01 9.993e-01 2.147e-07 3.007e-01 1.000e+00 5.132e-08 3.441e-01 2.664e-04Introduction to statistics using R (34/66) Solving simple regression I General formula 1 1 x1 y1 " # y2 1 x2 β × 0 + 2 = .. .. .. .. β 1 . . . . N 1 xN yN yi = β0 + β1 xi + i Only one continuous explanatory variable and n observations β0 is intercept and β1 is slope We want to find the pair of parameter estimates β̂0 , β̂1 that minimizes the sum of squared residuals SSR = N X i=1 Northeastern University Statistics workshop i2 = N X 2 yi − β̂0 − β̂1 xi : i=1 Introduction to statistics using R (35/66) Solving simple regression II Expand SSR: SSR = N X y2i − 2β̂0 yi − 2β̂1 xi yi + β̂20 + 2β̂0 β̂1 x + β̂21 xi2 i=1 Take the (partial) derivative with respect to β̂0 and set it to zero to find the intercept: N X ∂SSR = −2 yi − β̂0 − β̂1 xi = 0 ∂β̂0 i=1 Northeastern University Statistics workshop Introduction to statistics using R (36/66) Solving simple regression III Extract the coefficients from the summed terms: N N X X ∂SSR xi = 0 = −2 yi − N β̂0 − β̂1 ∂β̂0 i=1 i=1 Divide all terms by N : N N 1 X ∂SSR 1 X = −2 yi − β̂0 − β̂1 xi = 0 N i=1 N i=1 ∂β̂0 Replace summed terms by means: i h ∂SSR = −2 ȳ − β̂0 − β̂1 x̄ = 0 ∂β̂0 Northeastern University Statistics workshop Introduction to statistics using R (37/66) Solving simple regression IV Rearrange the terms to isolate β̂0 : β̂0 = ȳ − β̂1 x̄ Similarly, one can obtain β̂1 by taking the partial derivative of SSR: N X ∂SSR = −2 xi yi − β̂0 − β̂1 xi = 0 ∂β̂1 i=1 Plug the estimate of the intercept β̂0 and split the summed terms: N N N X X X ∂SSR = −2 xi yi − ȳ xi + β̂1 x̄ xi − β̄1 xi2 = 0 ∂β̂1 i=1 i=1 i=1 Northeastern University Statistics workshop Introduction to statistics using R (38/66) Solving simple regression V Noting that N X xi = N x̄, divide by N and isolate β̂1 : i=1 N N 1 X 1 X 2 2 xi − x̄ = xi yi − ȳx̄ β̂1 N i=1 N i=1 h i Recall that Var(X) = E X 2 − (E [X])2 and Cov [X] = E [XY] − E [X] E [Y], where E [X] = β̂1 = 1 N P xi : 1 PN i=1 xi yi − ȳx̄ N 1 PN 2 2 i=1 xi − x̄ N PN (yi − ȳ) (xi − x̄) Cov x, y s (y) = = i=1 = ρ (x, y) PN 2 Var [x] s (x) i=1 (xi − x̄) This final equation explicitly shows the relationship between the slope β̂1 and the variance, covariance and correlation of x and y Northeastern University Statistics workshop Introduction to statistics using R (39/66) Fit in simple regression Once we have estimates for the slope β1 and the intercept β0 , we can calculate a number of important values: Predicted values: ŷi = β̂0 + β̂1 xi Residuals: i = yi − ŷi These values can be used to test whether the regression complies with the assumptions and determine the quality of the fit: Total variation: SST = XN (yi − ȳ)2 XN (ŷi − ȳ)2 = i=1 Explained variation: SSXY i=1 Residual, error or unexplained variation: SSE = XN i=1 (yi − ŷ)2 Since, SST = SSXY + SSE , we define the coefficient of determination as the proportion of the variation in y explained by x: R2 = SSXY SSE =1− SST SST Northeastern University Statistics workshop Introduction to statistics using R (40/66) Notes about the coefficient of determination The range of the coefficient of determination is 0 ≤ R2 ≤ 1 It should never be used to compare models with different levels of complexity (i.e., number of explanatory variables) because it can only increase (and never decrease) with model complexity Indeed, regressing a response variable against many unrelated explanatory variables can produce relatively high R2 values and lead to spurious conclusions about their relationship (Freedman, 1983) The adjusted R2 corrects for this by penalizing model complexity. It can thus decrease with increasing model complexity if the additional explanatory variables do not explain a sufficient amount of the previously unexplained variation: R2adj = 1 − Northeastern University Statistics workshop SSE (N − 1) SST (N − K − 1) Introduction to statistics using R (41/66) Uncertainty and significance in simple regression I If the assumptions of simple regression are met, statistical significance can be determined using a simple t-test: Null hypothesis H0 : β1 = b0 ; Alternate hypothesis Ha : β1 , b0 The slope β1 is t-distributed with N − K degrees of freedom since the numerator is normally distributed and the denominator is χ2 distributed: tβ1 = β̂1 − b0 SEβ̂1 With N observations, K parameters (including the intercept) and the standard error of the slope SEβ̂1 : s SEβ̂1 = 1 − R2 Var(y) (N − 2) Var(x) Northeastern University Statistics workshop Introduction to statistics using R (42/66) Uncertainty and significance in simple regression II SEβ̂1 can be used to generate (1 − α)% confidence intervals to quantify uncertainty about the slope estimate β̂1 : ! 1−α SEβ̂1 CIβ̂1 = β̂1 ± t α2 ,K−2 α + 2 The intercept is also t-distributed with K − 1 degrees of freedom: tβ̂0 = β̂0 SEβ̂0 With standard error SEβ̂0 : s SEβ̂0 = RMSE Northeastern University 1 x̄2 + N (N − 1) Var(x) Statistics workshop Introduction to statistics using R (43/66) Uncertainty and significance in simple regression III Where: q PN RMSE = i=1 (ŷ − y)2 N−K The equation used to compute SEβ0 can be generalized to obtain the standard error around each ŷi : s SEŷi = RMSE (xi − x̄)2 1 + N (N − 1) Var(x) Note that CBi is smaller near x̄ and larger at more extreme values of x. Since CBi is also t-distributed, one can construct (1 − α)% confidence bands around each predicted value ŷ, which describe the uncertainty associated with each prediction ŷi : CBi = ŷi ± t α2 ,N−2 SEŷi Northeastern University Statistics workshop Introduction to statistics using R (44/66) Simple linear regression in R 30 Values: y Fit: y Residuals: y − y ● ● ● ● 25 ● ● ● y ● ● 20 ● ● ● 15 ● 0 2 4 6 8 10 12 x fit <- lm(y ~ x) # Fit model plot(x, y) # Plot data p=predict(fit, interval='conf', level=0.95) # Get predictions and bands lines(x, p[, 'lwr'], col='blue', lty=2) # Plot lower band lines(x, p[, 'upr'], col='blue', lty=2) # Plot upper band abline(fit, col='blue') # Plot regression coef(fit) # Get slope and intercept confint(fit, level=0.95) # Get 95% confidence intervals for parameters summary(fit) # Get slope, intercept, p-values, R^2, etc... Northeastern University Statistics workshop Introduction to statistics using R (45/66) Diagnostics for linear regression Remember that we have to adhere to four key assumptions when fitting regressions: 1 Homoscedasticity: the variance must be constant. This is typically visible when residuals show some pattern and can be remedied via transformations. Residual vs. prediction plots are ideal here. 2 Independence of residuals: residuals must be independent or not (auto)correlated. Use ACF or lag plots to detect autocorrelation. 3 Linearity: The relationship between your explanatory and response variable should be linear. Look for patterns in residual plots to detect violation of this assumption 4 Normality: Residuals must be normally distributed. Use residual or Q-Q plots to detect violations of this assumption. Northeastern University Statistics workshop Introduction to statistics using R (46/66) Diagnostics for linear regression in R par(mfrow = c(1, 3), mar = c(4, 4, 3, 1)) plot(fit, which = 1) plot(fit, which = 2) acf(resid(fit), main = "Autocorrelation of residuals") Autocorrelation of residuals Normal Q−Q 1.0 Residuals vs Fitted 4● ● 10 ● ●6 0.5 ACF 1 ● −1 −2 ● ● ● ● ● ● 0.0 ● ● ● −0.5 ● ● 11 ● 0 2 ● 0 Residuals ● ● ● Standardized residuals 4 2 ●4 ● −4 ●6 16 18 20 22 24 Fitted values 26 28 −1.5 −0.5 0.5 1.0 Theoretical Quantiles 1.5 0 2 4 6 8 10 Lag Residual plot: lack of trend or pattern indicates independence and homoscedasticity (i.e., ρ (ŷi , i ) = 0) QQ plot: distribution of values compared to a theoretical normal distribution. Want points to be on 1:1 line. Watch for deviations in the tails, which can be remedied via transformations. Northeastern University Statistics workshop Introduction to statistics using R (47/66) Multiple regression General formula 1 x11 x12 y1 y2 = 1 x21 x22 .. .. .. .. . . . . 1 xN1 xN2 yN Y = X . . . x1K 1 β0 . . . x2K β1 2 + × .. . .. .. .. . . . N βK . . . xNK × β + Where: Y: vector of N responses X: N × K + 1 matrix for K explanatory variables and a column for intercept β: vector of K coefficients + 1 intercept ∼ N(0, σ2 ): vector of N identically and independently distributed errors Northeastern University Statistics workshop Introduction to statistics using R (48/66) Solving multiple regression I We want to find the suite of parameter estimates β̂k with k ∈ [1, . . . , K] that 2 K n X X yi − β0 − β̂k xki : minimizes the sum of squared residuals SSR = i=1 k=1 After expanding SSR, taking the partial derivatives and setting them to zero, we get the following estimate for the intercept β̂0 and the slopes β̂k : β̂0 = ȳ − K X k=1 PK β̂p x̄k β̂k = − ȳ) xki − x̂ki − x̄k − x̂¯ k 2 PK ˆ i=1 xki − x̂ki − x̄k − x̄k i=1 (yi Where x̂ki is the predicted value for xki based on a regression of xk against all other explanatory variables xj with k , j Northeastern University Statistics workshop Introduction to statistics using R (49/66) Solving multiple regression II The x̂ki terms account for the non-independence or collinearity of the K explanatory variables When the xp variables are completely independent (i.e., ρ(xk , xj ) = 0 and x̂ki = 0), we recover the slope derived for simple regression: PN β̂k = i=1 (yi − ȳ) xki − x̄k PN 2 i=1 xki − x̄k Hence, unless perfectly independent, explanatory variables will affect each others slope estimates Northeastern University Statistics workshop Introduction to statistics using R (50/66) Diagnosing multicollinearity There are many ways of identifying multicollinearity between explanatory variables (Gotelli and Ellison, 2004): Bouncing β̂ problem: coefficient estimates β̂k will be unstable and change dramatically when increasing or reducing sample size Important predictors have small coefficients and high p-values Coefficients β̂k have the wrong sign Calculate the Variance Inflation Factor (vif function in car package), which measures how much the variance of an estimated regression coefficient increases because of collinearity with other regressors: VIF = Var(β̂k ) = 1 1 − R2k Here, R2k is obtained by regressing variable xk against all other explanatory variables. VIF = 1 when there is zero collinearity and VIF ≥ 5 means significant collinearity (Kutner et al., 2004) Northeastern University Statistics workshop Introduction to statistics using R (51/66) Dealing with multicollinearity Generate a correlation matrix between all pairs of explanatory variables, identify pairs with |ρ| ≥ 0.7 and remove one of them since they are highly redundant (Dormann et al., 2013). Use Principal Components Analysis (PCA) to reduce produce K independent axes representing linear combinations of your original collinear explanatory variables xk and regress the response variable against these axes Hierarchical partitioning (Chevan and Sutherland, 1991; Mac Nally, 2000, hier.part package): Perform regressions using all combinations of explanatory variables Compare results to determine the independent and joint explanatory power of each variable Determine the significance of each explanatory variable via Monte Carlo permutations Note: This approach is only valid for K =9 or fewer explanatory variables (Olea et al., 2010) Northeastern University Statistics workshop Introduction to statistics using R (52/66) Fit, uncertainty and significance in multiple regression I The model fit is assessed via the coefficient of (multiple) determination: R2 = SSXY SSE =1− SST SST The standard error SEβ̂k of the kth coefficient is: v t SEβ̂k = 1 − R2 Var(y) 1 − R2k Var(xk ) (N − K − 1) Northeastern University Statistics workshop Introduction to statistics using R (53/66) Fit, uncertainty and significance in multiple regression II Where R2k is the coefficient of determination obtained by regressing xk against all other explanatory variables xj with k , j. The calculations for determining the confidence intervals and the significance of the coefficients are identical to those described for simple regression. The significance of the overall regression is determined via an omnibus F-test: Null hypothesis H0 : βk = 0 for all k Alternate hypothesis Ha : βk , 0 for at least one k F= R2 /K 1 − R / (N − K − 1) 2 With K (numerator) and N − K − 1 (denominator) degrees of freedom Northeastern University Statistics workshop Introduction to statistics using R (54/66) Example of the bouncing β̂ problem in multiple regression set.seed(1) x1 = 1:12 x2 = x1 + runif(12) # x1 and x2 are highly correlated cor(x1, x2) ## [1] 0.9962 y = x + rnorm(n, sd = 3) + 15 fit = lm(y ~ x1 + x2) # Get beta coefficients summary(fit)$coef fit = lm(y ~ x1) # x1 strongly related to y summary(fit)$coef ## Estimate Std. Error t v ## (Intercept) 16.403 1.8763 8 ## x1 0.883 0.2549 3 fit = lm(y ~ x2) # x2 strongly related to y summary(fit)$coef ## Estimate Std. Error t value Pr(>|t|) ## Estimate Std. Error t v ## (Intercept) 19.007 2.506 7.583 3.385e-05 ## (Intercept) 16.1044 2.106 7 ## x1 4.940 2.767 1.786 1.078e-01 ## x2 0.8636 0.271 3 ## x2 -4.144 2.815 -1.472 1.751e-01 Northeastern University Statistics workshop Introduction to statistics using R (55/66) Generalized Linear Models (Zuur et al., 2009) General formula 1 x11 x12 η1 1 x η x22 21 2 .. .. .. = .. . . . . 1 xN1 xN2 ηN η = X . . . x1K β0 β . . . x2K 1 .. × .. .. . . . . . . xNK βK × β Where: ηi = β0 + β1 xi1 + . . . + βK xiK is a vector of N linear predictors of responses Y ∼ F , where F belongs to the exponential family of distributions Link function f relates the mean of Y to predictors η: f (E[Y]) = η Variance function V relates the variance of Y to the mean of Y: Var (Y) = φV (E[Y]), with φ being the dispersion parameter X: N × K + 1 matrix for K explanatory variables and a column for intercept β: vector of K coefficients + 1 intercept Northeastern University Statistics workshop Introduction to statistics using R (57/66) The different flavors of Generalized Linear Models Generalized Linear Models come in multiple flavors (function glm in R): Model type Data type Link Family General Linear Model Continuous identity gaussian Poisson Discrete counts log poisson Logistic Continuous proportions logit binomial Gamma Continuous inverse gamma The link and family arguments come in natural pairs but you can mix-and-match different link functions and families Because the link function relates f (E[Y]) = η, must back-transform using the inverse link function f −1 to get E[Y], predictions and model coefficients Northeastern University Statistics workshop Introduction to statistics using R (58/66) Solving Generalized Linear Models Generalized linear models are fit computationally via likelihood approaches using iterative weighted least squares. Hence, there are times when no satisfactory solution will be found (the algorithm will not converge) Because model fitting is done via likelihood, generalized linear models present results in terms of partitioning deviance instead of variance (Bolker, 2008; Crawley, 2007): Null deviance: Dnull = −2 log (Lnull ) − log (Ldata ) where Ldata is the log-likelihood of a model that fits the data perfectly and Lnull is the log-likelihood of a null model with an intercept only Model deviance: Dmodel = −2 log (Lmodel ) − log (Ldata ) where Ldata is the log-likelihood of a model that fits the data perfectly Significance of explanatory variable x determined via likelihood ratio test using a full model containing x and a reduced model without x using a χ2 distribution: p-value = χ2 (Dreduced − Dmodel , dfreduced − dfmodel ) Can compute pseudo R2 = 1 − DDmodel null Northeastern University Statistics workshop Introduction to statistics using R (59/66) Dispersion in Generalized Linear Models Each distribution in the exponential family (e.g., poisson, binomial) is described by a specific dispersion or spread The data is said to be overdispersed if its dispersion is larger than that expected for the specified distribution (e.g., the spread of your count data is larger than that of the poisson distribution) Conversely, the data is said to be underdispersed if its dispersion is smaller than that expected for the specified distribution Overdispersion occurs if residual deviance is larger than residual degrees of freedom 2 i=1 wi i PN Formally, dispersion σ2D = dfresidual where dfresidual = N − K , N is the number of observations, K is the number of model parameters and wi and i are respectively the weight and residual for observation i If σ2D = 1, then your data follow the specified distribution If σ2D < 1, then your data is underdispersed If σ2D > 1, then your data is overdispersed Northeastern University Statistics workshop Introduction to statistics using R (60/66) The effect of dispersion in Generalized Linear Models Dispersion affects the p-values and standard errors (SE) of the model coefficients Hence must account for deviations from the expected dispersion σ2D = 1 to obtain robust results To do so, can use the quasi-family of distributions, which account for [over|under]dispersion when computing the p-values and SE Standard error of coefficient accounting for dispersion σ2D : SEquasi = SEnonquasi σD p-value of coefficient accounting for dispersion σ2D : − Dmodel , dfreduced − dfmodel 2 σD Dreduced 2 p − valuequasi = χ If σ2D > 1 (overdispersion), using a quasi-distribution will increase the standard error of the coefficient estimates and tend to increase the p-value (conservative approach; low type I error) If σ2D < 1 (underdispersion), using a quasi-distribution will decrease the standard error of the coefficient estimates and tend to reduce the p-value (anti-conservative approach; higher type I error) Northeastern University Statistics workshop Introduction to statistics using R (61/66) Dealing with overdispersion in Generalized Linear Models Although the quasi-family of distributions will generate more accurate standard errors and p-values for the model coefficients, they will not improve the fit Additionally, the quasi-family of distributions will not produce an AIC value making it impossible to do model selection To generate a better fit with an AIC score, must specify a different and more appropriate distribution such as the negative binomial for overdispersed count data (glm.nb in package MASS) If you have count data that is overdispersed because of excessive zeros even when using the negative binomial, then consider fitting a zero-inflated model (package pscl) If you have continuous data that is overdispersed because of excessive zeros, then consider fitting a generalized linear model with a Tweedie distribution (package statmod) Northeastern University Statistics workshop Introduction to statistics using R (62/66) Generalized Linear Models in R budworm.data <read.csv("http://www.northeastern.edu/synchrony/stats/budworm_data.csv") # Logistic regression: specify dead and total individuals as columns budworm.mod <- glm(cbind(ndead, ntotal-ndead) ~ sex + log(dose), family=binomial, data=budworm.data) # Get summary of model fit budworm.mod.summary <- summary(budworm.mod) # No overdispersion (dispersion <- sum(budworm.mod$weights* budworm.mod$residuals^2)/budworm.mod$df.residual) ## [1] 0.5896 # Calulate (pseudo)-Rsquared (budworm.mod.rsq <-1-budworm.mod$deviance/budworm.mod$null.deviance) ## [1] 0.9459 # Run ANOVA to determine significance of explanatory variables budworm.mod.anova=anova(budworm.mod, test='Chisq') Northeastern University Statistics workshop Introduction to statistics using R (63/66) References I Bolker, B. M. 2008. Ecological Models and Data in R. Princeton University Press, 1 edition. Chevan, A. and M. Sutherland. 1991. Hierarchical partitioning. American Statistician, 45:90–96. Cohen, J. 1988. Statistical power analysis for the behavioral sciences. Psychology Press. Crawley, M. J. 2007. The R Book. Wiley, 1 edition. Dormann, C. F., J. Elith, S. Bacher, C. Buchmann, G. Carl, G. Carre, J. R. G. Marquez, B. Gruber, B. Lafourcade, P. J. Leitao, T. Munkemuller, C. McClean, P. E. Osborne, B. Reineking, B. Schroder, A. K. Skidmore, D. Zurell, and S. Lautenbach. 2013. Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography, 36:027–046. Northeastern University Statistics workshop Introduction to statistics using R (64/66) References II Freedman, D. A. 1983. A note on screening regression equations. The American Statistician, 37:152–155. Garcia, L. V. 2004. Escaping the bonferroni iron claw in ecological studies. Oikos, 105:657–663. Gotelli, N. J. and A. M. Ellison. 2004. A primer of ecological statistics. Sinauer Associates, 1 edition. Kutner, M., C. Nachtsheim, J. Neter, and W. Li. 2004. Applied Linear Statistical Models. McGraw-Hill/Irwin, 5th edition. Mac Nally, R. 2000. Regression and model-building in conservation biology, biogeography and ecology: The distinction between and reconciliation of ’predictive’ and ’explanatory’ models. Biodiversity and Conservation, 9:655–671. Moller, A. and M. Jennions. 2002. How much variance can be explained by ecologists and evolutionary biologists? Oecologia, 132:492–500. Northeastern University Statistics workshop Introduction to statistics using R (65/66) References III Olea, P. P., P. Mateo-Tomas, and A. de Frutos. 2010. Estimating and modelling bias of the hierarchical partitioning public-domain software: Implications in environmental management and conservation. PLoS ONE, 5:e11698. Rosnow, R. L. and R. Rosenthal. 1989. Statistical procedures and the justification of knowledge in psychological science. American Psychologist, 44:1276–1284. Zuur, A. F., E. N. Ieno, N. Walker, A. A. Saveliev, and G. M. Smith. 2009. Mixed Effects Models and Extensions in Ecology with R. Springer, 1 edition. Northeastern University Statistics workshop Introduction to statistics using R (66/66)