* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Survey
Document related concepts
Transcript
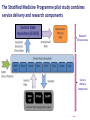
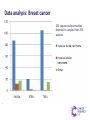
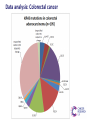
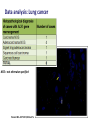
Molecular diagnosis of cancer: Making it a reality Peter Johnson On behalf of the CR UK Stratified Medicine Programme The march of technology… Single genes Gene panels Whole exomes Whole genomes Plus expression profiling, copy number variation, epigenetics, miRNA profiles… EGFR experience shows gradual test adoption even when funding is agreed Source: 12 months experience of EGFR testing in the UK, R. Butler, AstraZeneca et al, 2010 Background to the Programme SOMATIC MUTATION TESTING FOR PREDICTION OF TREATMENT RESPONSE IN PATIENTS WITH SOLID TUMOURS: – Already happening and demand predicted to increase – Funding not well established and access variable across the UK – Published data from quality assurance schemes suggest issues with the reproducibility and accuracy – Needs to work in formalin-fixed, paraffin embedded tissue – Needs to take account of multiple potential targets in each tumour – Little consensus on who to test, how to test, what to test or how to interpret and report the results The Stratified Medicine Programme pilot study combines service delivery and research components Central data repository (ECRiC) Research infrastructure Service delivery component Current gene list and technology Tumour Colorectal carcinoma Genes of interest KRAS BRAF NRAS PIK3CA TP53 mutation Breast carcinoma PIK3CA TP53 BRAF PTEN mutation + PTEN LOH by microsatellite analysis Prostate carcinoma PTEN BRAF mutation TMPRSS2-ERG fusion by FISH (moving to rt-PCR) + PTEN LOH by microsatellite analysis Lung carcinoma EGFR KRAS BRAF mutation ALK rearrangement by FISH Ovarian carcinoma Malignant melanoma TP53 PTEN PIK3CA BRAF mutation + PTEN LOH by microsatellite analysis BRAF KIT NRAS PIK3CA mutation Stratified Medicine Programme Patient Dataset Derived from the core NHS Cancer Outcomes and Services Dataset (COSD): – New NHS information standard; mandation and stepwise implementation from January 2013 – Linked to modernisation of the cancer registries with creation of a unified National Cancer Registration Service in England – Sections for demographics, diagnosis, imaging, pathology, treatment and outcomes data – Aim for automated data collection and extraction as far as possible 7229 patients consented and 5237 samples sent for testing Target for samples sent Patients consented Consented minus patient drop out Samples sent for testing Results returned Datasets sent to ECRIC 10000 Oct-12 Nov-12 Dec-12 9000 Monthly recruitment 673 708 455 Monthly samples sent 384 400 635* Monthly results returned 518 340 451 Paired samples 63 (1.5%) 8000 7000 6000 5000 68 69 (1.5%) (1.3%) 4000 3000 2000 1000 0 8 Analysis by tumour type 2000 1800 1600 1400 1200 Target for samples sent Consented 1000 Sent for testing Tested 800 600 400 200 0 Breast Colorectal Lung Melanoma Ovary Prostate 9 Programme whole gene test failure rates - Year 2 % Whole gene test 70% 60% 50% 40% 30% 20% 10% 0% 17% 4% 7% 6% 9% 6% 8% 4% 4% 3% 2% 10 6% 6% Results from the first 1000 cases Proportion of tumours with sequence variation by type Data analysis: Breast cancer 260 sequence abnormalities detected in samples from 246 patients Data analysis: Colorectal cancer Data analysis: Lung cancer NOS = not otherwise specified Pennell N A JOP 2012;8:34s-37s Findings from the Stratified Cancer Medicine programme • General acceptability to patients: over 98% consent • Key role of clinical teams for each tumour type • Critical role of pathology department in managing tissue samples • Impact of mass testing in technology hubs • Value of QA system • Highly variable sample quality, with impact on test failure rates • Complexities of NHS IT. XML messaging protocols • Automated data extraction not yet reliable 15 Challenges of obtaining the SM Programme patient dataset from routine clinical data • Data reflects the patient pathway with a multitude of one-tomany relationships: repeating identifiers required in submitted data to ensure linkage in database • Difficulty in sourcing some data items e.g. histopathology data, co-morbidity scoring, performance status • Lack of inter-operability between information technology systems used in histopathology and other parts of the electronic patient record • Use of Read codes and different versions of SNOMED • Cross-border data issues: problems sourcing some demographic details Future Priorities • Improve completeness, linkage and standardization of submitted data • Defining minimum sample requirements • Optimising test turnaround times • Moving towards routine commissioning of tests of proven utility • Development of multiplex technology for parallel analysis of many genes • Expansion of early phase therapy portfolio to exploit the molecular findings Acknowledgements • The patients who have consented to take part in the programme • Lead investigators and teams at the clinical and technology hubs • Stratified Medicine Programme team at CR-UK HQ • Dr Jem Rashbass and team, National Cancer Registration Service •Funding partners AstraZeneca and Pfizer