* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download No Slide Title
Survey
Document related concepts
Transcript
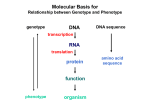
Regulation of eukaryotic genes Gene silencing Enhancers Activators Functional domains of activators Idea for another extra credit project Explore DNA binding domains of proteins. 1. Go to a web site with a Chime tutorial, e.g. GAL4 or Cro 2. Or use Kinemages 3. Write a roughly 2 page report on how a particular protein recognizes a DNA sequence States of eukaryotic genes • Inactive: – Closed chromatin – Open chromatin, but repressors or lack of activators keep frequency of initiation low. – Open chromatin, transcription has initiated, but polymerases will not elongate. • Active: – Open chromatin, basal transcription: requires TATA + Inr – Open chromatin, activated transcription: requires enhancer or upstream activator sequences Silent and open chromatin Transcription initiation and pausing Basal and activated transcription Silencing Mechanism Silencer • Cis-acting sequences that cause a decrease in gene expression • Similar to enhancer but has an opposite effect on gene expression • Gene repression - inactive chromatin structure (heterochromatin) • Examples – Telomeric silencing – a or genes - silent loci of mating type switching in yeast Silencer binding proteins • Silencer binding protein serve as anchors for expansion of repressed chromatin • Rap1 protein binds to silencer elements • SIR proteins (Silent Information Regulators) • Nucleates assembly of multi-protein complex – hypoacetylated N-terminal tails of histones H3 and H4 – methylated N-terminal tail of H3 (Lys 9) • Experiments: Condensed chromatin – Resistant to DNaseI digestion – Delete silencer - genes are derepressed Gene Silencing Silencing Mechanism Enhancers • Cis-acting sequences that cause an increase in expression of a gene • Act independently of position and orientation with respect to the gene. • Can act to: – Increase the rate of initiation at a promoter – Increase the fraction of cells in which a promoter is active SV40 Control region • Origin of replication • Promoter and upstream activator sequences for early transcription • Promoter for late transcription • Enhancer SV40 map Many regulatory DNA sequences in SV40 control region Stimulation of transcription by enhancer is independent of orientation and position SV40: Early Late wt T-Ag + pos T-Ag + orien T-Ag + Enhancer Enh- T-Ag - Enhancers also regulate cellular genes Enhancer contains multiple binding sites for transcriptional activators SV40: Early Late Enhancer T-Ag wt A C B high level deletion C B low level revertant C C B high level An enhanson Enhancers can occur in a variety of positions with respect to genes Enhancer Upstream Enhancer P Transcription unit Adjacent Downstream Internal Distal Ex1 Ex2 Activator proteins Modular nature of activator proteins • DNA binding domain: recognition and binding to specific DNA sequences • Multimerization domain: allows formation of homo- or hetero-multimers • Activation domain: – Needed for increase in expression of responding gene – Targets are still under investigation • General transcription factors • Histone modifying enzymes • Nucleosome remodeling complexes, etc Modular structure of GAL4 1 98 148 196 N DNA Activation binding Dimerization 768 881 C Activation GAL80 binding Induction by galactose exposes an activation surface • In the presence of galactose, GAL4 activates several genes whose products are required for galactose metabolism. • GAL4 binds to a DNA sequence called UASG. • In the absence of galactose, GAL80 blocks GAL4 activation. • Binding of the sugar causes GAL80 to move. • This exposes the activation domain of GAL4. Induction of GAL4 Domain swap experiments show the domains are interchangeable • Fuse an DNA-binding domain (DBD) from one transcription factor to the activation domain (AD) of a different one. – DBD from LexA (E. coli) – AD from GAL4 (yeast) • Now a target gene can be placed under control of the DNA binding site for the first factor – GAL1 gene with oLex (LexA binding sites) can be activated by the fusion protein. • Basis for 2-hybrid screen for any interacting proteins Domain swap experiments: Diagram 1 Domain swap experiments: Diagram 2 Two Hybrid Screens (Interaction Cloning), part 1 Two Hybrid Screens (Interaction Cloning), part 2