* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Structural Knowledge Base Development for Metal Complexes
Oxidation state wikipedia , lookup
Hydroformylation wikipedia , lookup
Cluster chemistry wikipedia , lookup
Metal carbonyl wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Stability constants of complexes wikipedia , lookup
Metalloprotein wikipedia , lookup
Spin crossover wikipedia , lookup
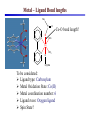
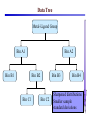
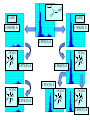
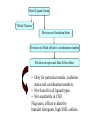
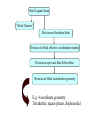
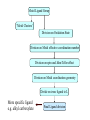
Development of Molecular Geometry Knowledge Bases from the Cambridge Structural Database Stephanie Harris Crystal Grid Workshop Southampton, 17th September 2004 Cambridge Structural Database Stored geometric information for ~300,000 structures Search using Conquest Substructure search, user input required Molecular Geometry Knowledge Bases Library of chemically well-defined geometric information Limited user input Rapid retrieval of statistical data Molecular Geometry Knowledge Base: Mogul Bond lengths, valence angles and torsion angles Compiled from the CSD Applications Model building Refinement restraints Structure validation Comparative values Published bond length tables: Organic and metal containing structures Published late 1980s Compiled from CSD of ~50,000 structures Cannot be accessed by computer programs Mogul 1.0 Whole molecule input Graphical (cif, SHELX, mol2 files) or command-line interface Integration with client applications, e.g. Crystals Quick, automatic retrieval of statistical data, histogram distributions, CSD structures Search Algorithm All non-metal fragments in the CSD coded Set of keys code chemical environments Fragments with identical keys are chemically identical Use hierarchical search tree Generalised searching if insufficient hits Mogul Search .S1 .C7 Search O pTol O S N N O N CN Metal – Ligand Bond lengths Me C O Co-O bond length? O N OH2 Co N OH2 O C(O)Me To be considered: Ligand type: Carboxylate Metal Oxidation State: Co(II) Metal coordination number: 6 Ligand trans: Oxygen ligand Spin State? Method Analysis of M-L bond lengths. For a range of metal and ligand types identify factors which influence M-L bond lengths and evaluate their importance. For a defined Metal-Ligand group sub-divide bond length distribution to produce ‘chemically meaningful’ datasets: • Unimodal distributions. • ‘Reasonably small’ sample standard deviations. From hand-crafted examples develop an algorithm to produce a molecular geometry knowledge base for metal complexes. Data Tree Metal-Ligand Group Bin A1 Bin B1 Bin A2 Bin B2 Bin C1 Bin B3 Bin C2 Bin B4 Sharpened distributions Smaller sample standard deviations Criteria Influencing M-L Bond Lengths 1. Ligand, L 2. Coordination mode of ligand 3. Effective Metal Coordination Number 4. Metal Oxidation State 5. Metal clusters and cages 6. Spin state 7. Jahn-Teller effect 8. Metal coordination geometry 9. Ligand trans to L M =6 M =6 Ligand Template Library B M A B B Ligand • Non-metal atom or fragment bonded to a metal. • Two ligands are the same if they have same connectivity (topology) and stereochemistry. OO- O O Method • All ligands in CSD to be classified. • Classify according to contact atom coordinated to metal. • Ligands with multiple contact atoms can be present in more than one ligand group. e.g. SCN- Cambridge Structural Database Approximately 22,000 formulae Approximately 780,000 ligands No. of occurrences of unique formulae in CSD Total Number of Ligands Number of formulae 550,000 (70%) 70 100 – 999 109,263 (14%) 394 10 – 99 76,000 (10%) 3000 1–9 45,700 (6%) 18,937 Ligand Template Hierarchy • Exact ligand templates (724) • R-substituted templates (H’s replaced with ‘innocent’ R groups) • Generic templates (ALL ligands classified) Cobalt Carboxylate Bond Lengths Co O 3 C C sp No. of Frags. O Co-O: 1.929(62) Å 619 Fragments Co-O (Å) Co O 3 C C sp O Co(II) Co(III) 2.049(58) Å 1.904(20) Å 1.929(62) Å OC(O)C L L Co II L L L 2.073(42) Å 1.904(20) Å OC(O)C L L Co III L L L 1.910(15) Å OC(O)C L L Co II L L O 2.074(32) Å OC(O)C L L Co III L L N OC(O)C L L Co III L L O 1.895(17) Å Fe-Cl Chlorides 2.242(68) Å Cl III Fe L L L 2.189(24) Å Pyridines e.g. Fe (spin state) Fe N Fe(II)L5py High Spin 2.166(84) Å 2.225(29) Å Tertiary phosphines, Carbon-ligands Copper complexes (Jahn-Teller effect) Standardisation of Cu connectivity Cu(II)-OH2 2.232(225) Å Metal-Ligand Knowledge Base 1. CSD data adjustment: Standardisation of metal connections Assignment of metal as part of a metal cluster Assignment of metal oxidation state 2. Classification of ligands by ligand template library 3. Perform algorithm on all possible M-L fragments to produce knowledge base Algorithm: Metal-Ligand Group From ligand template library: Generic or more specific e.g. Carboxylates: C O O O O C C O 3 sp C C O Et Metal-Ligand Group ‘Metal Clusters’ Division on Oxidation State Division on Metal effective coordination number Division on spin and Jahn-Teller effect • Only for particular metals, oxidation states and coordination numbers. • Not found for all ligand types. • Not searchable in CSD. Flag users, effects evident by: bimodal histogram, high SSD, outliers. Metal-Ligand Group ‘Metal Clusters’ Division on Oxidation State Division on Metal effective coordination number Division on spin and Jahn-Teller effect Division on Metal coordination geometry E.g. 4-coordinate geometry: Tetrahedral, square planar, disphenoidal Metal-Ligand Group ‘Metal Clusters’ Division on Oxidation State Division on Metal effective coordination number Division on spin and Jahn-Teller effect Division on Metal coordination geometry Divide on trans ligand to L More specific ligand e.g. alkyl carboxylate Final Ligand division Generalised Searching • No hits or insufficient number of hits. • Allows the retrieval of data on related fragments. • Hierarchical search tree structure • Move up to a higher, less specific level of data tree. • Order of algorithm important. Should order of criteria be changed? Should order depend on M-L group? E.g. Should oxidation state always be the first main division? Conclusions • Pre-processing of structural data from the CSD to construct molecular geometry knowledge bases. • Knowledge bases to contain chemically well-defined datasets. • Limited user input required. • Quick, automatic retrieval of statistical data, distributions. • Efficient analysis of large number of chemical fragments. • Outliers, high SSD? Further Analysis – Computational Chemistry. • Further development to include extra chemical information e.g. computational data. Acknowledgements Bristol University: Guy Orpen Natalie Fey X-Ray Crystallography Group Cambridge Crystallographic Data Centre: Robin Taylor Frank Allen Ian Bruno Greg Shields