* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Secondary structure
Genetic code wikipedia , lookup
Circular dichroism wikipedia , lookup
Expanded genetic code wikipedia , lookup
Self-assembling peptide wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Biosynthesis wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Biochemistry wikipedia , lookup
Peptide synthesis wikipedia , lookup
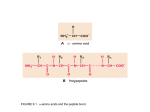
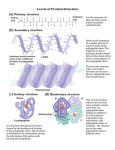
Secondary Structure of Proteins The term secondary stricture refer to the common regular folding pattern of polypeptide backbone. Some physicochemical principles which act as guide lines for the determination of three dimensional conformation of proteins: 1. Oxygen has the highest electronegativity. 2. Availability of lone pair of electron on Nitrogen. 3. The oxygen and hydrogen atom of the peptide bonds have the potential for hydrogen bonding. 4. Bulky groups on side chains of amino acid residues in polypeptide cause steric hindrance when present close to gather. 5. Hydrophobic residues try to be together away from water. 6. +ive or –ive charge on side chains can stabilize or destabilize the higher order structures. Nature of the Peptide Bond: In 1930s and 1940s Linus Pauling and Robert Corey analyzed amino acids and dipeptide structures by X-ray crystallography. Their main observations were: 1. The C-N bond in peptide group is 0.13 Ao shorter than the regular C-N single bond (indicating partial double bond character). 2. The C=O bond is 0.02 Ao longer than the regular C=O double bond in aldehydes and ketones (partial single bond character) These observations suggested that there is a resonance or partial sharing of two pairs of electrons between the carbonyl oxygen and the amide nitrogen. Oxygen has partial –ive charge and Nitrogen has partial +ive charge. Thus the peptide bond is a rigid planer structure stabilized with – 85kJ/mole resonance energy. With only a few exception, all the peptide groups assume trans conformation i.e. Ca atoms are on the opposite sides of the peptide bond. Polypeptide backbone conformation: Based on the fact that peptide bond is a planer and rigid structure, the poly peptide backbone conformation will depend mainly on the torsion angle or rotation angles about Ca-N bond ( f angle) and Ca-C bond ( j angle) Whenever there is a possibility of free rotation of groups across a bond, amongst all the possibilities only those conformations are most stable where there is minimum steric hindrance. Ramchandran Plot: •Using the Van Der Waals distances for various inter-atomic contacts to asses the steric hindrance Ramchandran calculated the rotation angles which would allow the most stable conformation with little or no hinderance. •Both rotation angles can be platted against each other, and stable conformations can be predicted based on the amino acid residues. •When crystal structures of many proteins were determined by X-ray crystallography, the rotation angles obtained from these studies were found to be matching with the predicted allowed rotation angles. Ramchandran Plot for L-Ala residues Helical Structures With the information already obtained such as ; The rigid and planer nature of peptide bond, Allowed conformations using torsion angles and X-ray result of keratin protein obtained by Willium Astbury, who observed that a regular structure was occurring in keratin that reapeated everu 5.15 to 5.2 Ao. Pauling and Corey proposed the helical conformation which they called a helix. In this structure the polypeptide backbone is is tightly wound around an imaginary axis. The side chain R groups of AA residues protrude outward from the helical backbone. The repeating unit is a turn of helix (3.6 amino acids) with the periodicity of 5.4 Ao Characteristics of a helics: Polypeptide chains containing L-aminoacid residues make righr handed helices. Therefore all the a helical conformations in living systems are right handed. No of AA per turn (n)……3.6 Distance traveled in one turn (pitch) p 5.4 Ao Torsion angles f= -57 o and j= -47o Hydrogen bonding between the C=O group of nth AA residue to the NH group of the n+4th AA residue in peptide. Polypeptide with D-AA residue make left handed a helics f= +57 o and j= +47o Rest of the parameters are same as for right handed helics. Effect of amino acid sequence on the stability of a helix Constrains that affect the stability of a helix 1. The electrostatic repulsion or attraction between the successive AA residues with charged R groups 2. The bulkiness of the adjecent R group. 3. Interaction between the side chains spaced 3 or 4 AA apart 4. Occurrence of Pro and glycine. 5. The interaction of the amino acid residues at the ends of the helical segment and the electrical dipole inherent to the a helix Interaction between the R group of AA three residue apart in an a helix The electric dipole of a helix The b-conformation or b-sheets Pauling and Corey (1951) proposed a second type of repititive structure they called “b-conformation”. This is more extended conformation of polypeptide chains. In this structure, the polypeptide chain is extended in a zigzag fashion rather than in helical form. The zigzag polypeptide chains are arranged side by side like pleats and hydrogen bonds are formed between the adjacent segment of the chain. The R group of the side chain protrude in opposite side creating an alternating pattern. Parallel b-conformation: with adjacent polypeptide chains Having same amino to carboxy terminus orientation. Anti-parallel b-conformation: with adjacent polypeptide chains having opposite amino to carboxy terminus orientation. When two or more beta sheets are layered closely together within a Protein the R groups of amino acid residues on touching surfaces must be small for example silk fibroin or spider web fibroin which Have beta sheet structure have very high content of Gly and Ala.