* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download TITLE : BLAST
Protein (nutrient) wikipedia , lookup
Protein adsorption wikipedia , lookup
Community fingerprinting wikipedia , lookup
Genetic code wikipedia , lookup
Molecular evolution wikipedia , lookup
Protein structure prediction wikipedia , lookup
Point mutation wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
TITLE : BLAST
1
2
Table of Contents
1.0 Introduction to Blast ------------------------------------------------------------------------------ 4
2.0 Types of Blast --------------------------------------------------------------------------------------11
3.0 Common Databases for Use with BLAST available at NCBI ----------------------------21
4.0 How Blast Work? -------------------------------------------------------------------------------- 29
5.0 Interpretation of Blast Result ------------------------------------------------------------------38
3
Basic Local Alignment Search Tool
(BLAST)
1.0 Introduction
Basic Local Alignment Search Tool (BLAST) is an algorithm used for comparing
primary biological sequence information such as nucleotides sequence and amino-acids
sequence in order to find regions of local similarity.
BLAST algorithm having the same function with FASTA algorithm . However, BLAST
works faster and more time-efficient than FASTA.
FASTA will compared it’s query sequence though out all database sequences while BLAST
will search for the high number of similar local regions and gives the result after a threshold
value reached.
BLAST also widely used among bioinformatics researchers due to its availability on the
World Wide Web through a large server at the NCBI (National Center for Biotechnology
Information). Some other site that may use BLAST as well are GenomeNet, ExPasy and
FlyBase. There still other sites uses BLAST.
4
1.1 Background
Designed by Eugene Myers at the University of Arizona, Stephen Altschul, Warren
Gish, and David J.Lipman at the U.S. National Center for Biotechnology Information
(NCBI) and Webb Miller at the Pennsylvania State University.
It was published in the Journal of Molecular Biology in 1990.
The earlier version of BLAST algorithm is Smith-Waterman algorithm.
BLAST is faster than Smith-Waterman algorithm.
1.2 Uses of BLAST
There are some uses of BLAST underlined by Wikipedia website.
1.2.1 Identifying species
BLAST can identify a species and find homologous species correctly. This is very useful
when we are working with a DNA sequence from an unknown species.
1.2.2 Locating domains
BLAST can locate known domain within the sequence of interest when we are working with
a protein sequence.
1.2.3 DNA mapping
When working with a known species, and looking to sequence a gene at an unknown
location, BLAST can compare the chromosomal position of the sequence of interest, to
relevant sequences in the database(s).
1.2.4 Comparison
When working with genes, BLAST can locate common genes in two related species, and can
be used to map annotations from one organism to another.
5
1.3 Input
Input sequences are either in FASTA or Genbank format.
1.3.1 FASTA
FASTA is also known as Pearson format
Advantages:
Easy to manipulate and parse sequences using text-processing
processing tools and scripting languages
like Python, Ruby, and Perl.
FASTA format is a text-based
based format for representing either nucleotide sequences or peptide
sequences, in which nucleotides or amino acids are represented using single
single-letter codes.
Nucleic acids code supported (Nucleotide Sequence):
Amino acids code supported (Peptide Sequence):
6
*Noted that the degenerate nucleotide codes in red are treated as mismatches in nucleotide
alignment.
Restrictions of FASTA format input:
all lines of text be shorter than 80 characters in length.
The description line is distinguished from the sequence data by a greater
greater-than (">")
symbol at the beginning.
Blank lines are not allowed in the middle of FASTA input.
lower-case
case letters are accepted and are mapped into upper-case
a single hyphen or dash (‘ - ’) can be used to represent a gap of indeterminate length.
In amino acid sequences,
sequences, U and * are acceptable letters (see above).
U is replaced by X first before the search since it is not specified
specified in any scoring
matrices.
Before submitting a request, any numerical digits in the query sequence should either
be removed or replaced by appropriate letter codes (e.g., N for unknown nucleic acid
residue or X for unknown amino acid residue).
Too many degenerate
enerate codes within an input nucleotide query will cause blast.cgi to
reject the input.
To represent gaps, use a string of N or X instead of ‘ - ’.
7
Example of input in FASTA format:
Example of barely sequence input :
Example of Multi Sequence FASTA
FAS
file:
8
1.3.2 Genebank
Genbank sequence database is an open access.
It is an annotated collection of all publicly available nucleotide sequences and their protein
translation.
Contained over 65 billion nucleotide bases in more than 61 million sequences in its database.
Identifier such as accession, accession.version or gi's usually used as an input .
Example of acceptable Genebank input :
CAA89576
CAA89576.1
1015707
gi|1015707
9
1.4 Output
BLAST output can be delivered in a variety of formats include HTML, plain text, and XML
formatting.
For NCBI's web-page, the default format for output is HTML.
Result on NCBI are given in a graphical format showing the hits found, a table showing
sequence identifiers for the hits with scoring related data, as well as alignments for the
sequence of interest and the hits received with corresponding BLAST scores for these. The
easiest to read and most informative of these is probably the table.
10
2.0 Types of BLAST
Basic Local Alignment Search Tool (BLAST) is one of bioinformatics tool that hosted by
National Center for Bioinformatics Information (NCBI) that allows similarity searches
against the databases of proteins or DNA which has been constantly updated. In BLAST,
there are different program and tools that help anyone that are doing research or study in this
area. There are five types of BLAST tools.
1. Nucleotide BLAST
In nucleotide BLAST tools, user must be using the nucleotide query as the sequence, and
then NCBI will search the inserted query against the nucleotide database.
-blastn
In blastn algorithm, nucleotide query will be compared against the nucleotide database.
11
Figure 2.1: Nucleotide BLAST interface
- Megablast
Megablast is one types of nucleotide BLAST algorithm. This algorithm specifies to identify
an unknown query/sequence that has been inserted by user whether the query/sequence
already exists in other public database. Furthermore, megablast used to compare two large
sets of sequences swiftly. Besides that, this algorithm will efficiently search long alignments
between similar query/sequence. In correspond to use this algorithm, users must choose
in program selection. Megablast algorithm is
specifically designed to efficiently find long alignments between very similar sequences and
thus is the best tool to use to find the identical match to your query sequence.
Figure 2.2 Megablast algorithm interface
Example of nucleotide query:
acatgggattatcaatcaccagttaacaacaatcttcagtcttccaccataactcagtgtaaaaccgagcccagacacacaaatggcttc
ggttgaagaaattagaaatgcccaacgtgctcaaggtccagccaccattctagccataggcacagccaccccagctcattttatcaacc
aggctgagtatcctgattactactttcgtatcacaaacagtgagcacaaaacagagttaaaagaaaaattcaagcgcatgtgtgataaat
ccatgataaacaaac
12
2. Protein BLAST
In protein BLAST tools, user must use the protein query as the sequence, and then NCBI will
search the inserted query against the protein database.
Figure 2.3 Protein BLAST interface
Example of protein query:
MSINIRDPLIVSRVVGDVLDPFNRSITLKVTYGQREVTNGLDL
13
3. BLASTX
Blastx is another tool in BLAST. Firstly, user must insert a nucleotide query as the sequence
then blastx will convert the query into six-reading frames protein sequences. After that, the
translated query will be compared against NCBI protein databases and return the results if
there are hits. This tool is advantageous when user trying to search homologous protein in a
nucleotide coding region.
Figure 2.4 Blastx interface
14
Figure 2.5: Sample blastx output
* six-reading frames - (in sequence analysis) translation of a DNA sequence taking into
account the three possible reading frames in each direction of the strand, giving rise to three
forward (positive strand) and three reverse (negative strand) translations.
Example:
Input sequence: attgttgctacttct
Reading frame:
123
at tgttgctacttct
1st reading frame:
I
V
A
T
S
123
at tgttgctacttct
2nd reading frame:
L L
L
L
123
at tgttgctacttct
3rd reading frame:
C
C
Y
F
15
4. Translated BLAST: tblastn
Tblastn is the tools that used protein query that have been translated into six-reading frames
then compares it against NCBI nucleotide database. Uses of this tools is to find homologous
protein coding regions in nucleotide sequences that are not annotate such as the expressed
sequence tags (ESTs) and draft genome record (HTG). Both EST and HTG are located in
BLAST database respectively.
EST
EST is the short and single-read complementary DNA (cDNA) sequences which consist of
biggest pool of sequence data for many organisms and portions of transcripts of genes that
have not been characterized.
HTG
HTG is the draft sequences from many genome projects or biggest genomic clones.
Figure 2.6 Translated BLAST: tblastn interface
16
Figure 2.7: Sample tblastn output
17
5. Translated BLAST: tblastx
Tblastx is the tools that converts nucleotide query that have been inserted into six-reading
frames protein sequence then compares it against NCBI nucleotide database. This tool detects
potential frame-shift and ambiguities that may prevent open-reading frame (OFR) and also to
identify potential proteins that are encoded by ESTs.
Figure 2.8 Translated BLAST: tblastx interface
Figure 2.9: Sample tblastx output
18
Summary/Comparison:
Types of
Types of
Types of
BLAST
query /
database
sequences
use
Blastn
Nucleotides
Purpose
Nucleotide - Normally used.
Function
- Useful tool for primer or short
sequence search.
- Identify similar query
sequence
- Directly compare from nucleotide
query against nucleotide database.
Blastp
Peptides
Protein
- Normally used.
- Directly compare from protein query
against protein database.
- Identify similar query
sequence
Blastx
Translated
Protein
nucleotide
- Identify similar protein
sequence
- Useful when identifying homologous
protein (protein that having similar
primary, secondary and tertiary
structure) in a nucleotide coding
region.
- Useful for identifying of the
unknown reading frame sequence.
- More sensitive than blastn.
Tblastn
Peptides
Translated
- Identify similar protein
nucleotide
- Useful for identified homologous
protein coding regions in
unannotated nucleotide sequences.
19
Tblastx
Translated
Translated
nucleotide
nucleotide
- Identify very distant
- Useful tool for identified novel gene
relationships between
(pieces of DNA that have not been
nucleotide sequences
identified before as being genes).
- Identify similar protein.
- More sensitive than blastp.
- Take long time to search because of
it sensitivity and batch searching is
not recommended.
- Long query/sequences also not
recommended.
Importance of translation BLAST (blastx, tblastn and tblastx).
1. Firstly, protein sequences are better conserved than nucleotide sequences.
2. Results produced are more reliable and accurate when dealing with coding DNA.
3.
Able to directly see the function of the protein sequence, since by translating the
sequence of interest before searching often gives the annotated protein hits.
20
2.1 Advantage(s) and Disadvantage(s) of BLAST tools on net or computer
1. On net
Advantage(s)
•
Disadvantage(s)
User can freely use the database that has
•
been remotely hosted by NCBI.
•
User totally cannot use the customized
database.
Completely no setup or only little setup
•
Requires internet connection
is required to manage the tools.
2. On computer
Advantage(s)
•
Disadvantage(s)
•
Can Use a Customized Database.
Requires some setup and computer
expertise
•
•
For UNIX user it is better suited to
scripting or automation when performed
large number of queries.
•
no internet connection
21
Expensive
Additional note:
http://pga.mgh.harvard.edu/Parabiosys/education/seminars/blast.pdf
Useful links:
1. http://blast.ncbi.nlm.nih.gov/Blast.cgi?CMD=Web&PAGE_TYPE=BlastHome
2. http://www.ncbi.nlm.nih.gov/blast/html/BLASThomehelp.html
3. http://www.ncbi.nlm.nih.gov/blast/Why.shtml
22
3.0 BLAST Database Content
A BLAST search has four components: query, database, program, and search purpose/goal.
To discuss effective BLASTprogram selection, we first need to know what databases are
available and what sequences these databases contain. in thissection, we will first take a look
at the common BLAST databases. According to their content, they are grouped into
nucleotideand protein databases. These databases and their detailed compositions are listed in
the two tables below.NCBI also provides specialized BLAST databases such as the vector
screening database, variety of genome databases fordifferent organisms, and trace databases.
23
24
25
Creating custom databases
there are 3 step to make custom database;database;
1. fasta file is convert into a binary blast database.
2. change the blast.html so that the database can be selected at drop-down
drop down menu
3. modified the blast.rc
26
1.Converting
Converting a fasta file into a binary BLAST database
A binary BLAST database is a collection of multiple files
1. .nhr
2. .nin
3. .nsq files
They must be created from a fasta file in a terminal, using the BLAST+ toolkit, The legacy
BLAST toolkit can be used to achieve the same goal, though the command line syntax
differs.
Enter the following into the terminal;terminal;
2.Adding
Adding the custom database to the drop-down
drop
box
To add a database to the drop-down
drop down box, modify blast.html in a text editor
After the HTML has been modified correctly, the custom database should be able to be found
when viewing the blast page with an internet browser
27
3.Modifying blast.rc
change the blast.rc file like below
As many databases can be added space separated as required after each program.
28
4.0 How Blast Work
4.1 INTRODUCTION
The BLAST programs improved the overall speed of searches while retaining good
sensitivity.This is important as databases continue to grow.Blast first breaking the
query sequence and database sequences into fragments of "words".Then Blast
seeking matches between fragments of query and database. Whenever the algorithms
find the “hit” (a match between a “word” and a database entry) ,the hit are extend in
either direction in an attempt to generate an alignment with a score exceeding the
threshold of "S".
4.2 BLAST Algorithm
The BLAST programs are the comparison algorithms that are used to search sequence
databases for optimal local alignments to a query.The algorithms :
Scoring of matches done using scoring matrices
Sequences are split into words
•
For protein sequence(default n=3)
BLAST algorithm extends the initial “seed” hit into an HSP
HSP = high scoring segment pair
3 Step Process of Blast
1. Make lookup (hash) table of query sequence
2. Scan database for hits
3. Extend hits that meet certain scoring criteria
Figure of how blast work:
29
lookup table
PROTEIN
Query Sequence: MLTNSEFVSMWSAESCRTPLCSVNNSYFPGAL
MLT NSE FVS…..
LTN SEF VSM…
TNS EFV SMW…
BLAST STEP PROCESS
30
After making words for the sequence of interest.The neighborhood words are also
assembled and these words must having a score at least the threshold T or greater than
T.
How to calculate neighborhood score threshold?
By using Blossum 62 scoring Matrices
Query Word: GTW
Subject Word: GTW = 6 + 5 + 11 = 22
Query Word: GTW
Subject Word: GSW = 6 + 1 + 11 = 18
Query Word: GTW
Subject Word: ATW = 0 + 5 + 11 = 16
By comparing the each character in query words and Subject word,we get the value
and sum up for the total score.For Example:
Query Word= GTW
and Subject Word=GTW
31
The first character in query word which is G are compared to the first character in subject
word which is also G.Where are the value 6 are coming from?.The answer is coming from the
Blossum 62 scoring matrix(refer figure 1 below).As you can see,the value where the
intersection of two square which is in red colour in figure 1 are taken.The same step are apply
to the second character and third character.The value which is 6,5 and 11 are sum and get the
total of 22.
The same step are also apply to the other words.
Figure 1
Step 2:
Scan the database for entries that match the lookup table
fast and relatively easy.
32
Step 3:
when manage to find a hit (a match between a “word” and a database entry), extend
the hit in either direction.
Keep track of the score (use Blossum 62 scoring matrix)
Each time the alignment is extended, an aligntment score is increases or decreased.
When the alignment score drops below a predefined threshold, the extension of the
alignment stops
Neighborhood Words
Neighborhood can be Words with a score over a predefined threshold
33
Protein Blast
http://blast.ncbi.nlm.nih.gov/Blast.cgi
Click the protein blast from BLAST program
34
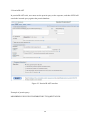
Page for protein Blast.
Four components to a BLAST search:
(1) Choose the sequence (query)
(2) Select the BLAST program
(3) Choose the database to search
(4) Choose optional parameters
Then click “BLAST”
(1) Choose the sequence (query)
35
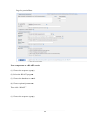
(2) Select the BLAST program
36
(3) Choose the database to search
(4) Choose optional parameters Then click “BLAST
37
5.0 Interpretation of Blast Result
NCBI blast can accept the inputs query sequence in form of FASTA format, GI or accession
number. User should get a result page from NCBI blast after running the blast. So, this
section will discuss about the interpretation of results from NCBI blast. The structure of
result page will consist of summary, graphical overview, descriptions table and alignments
section.
Note : To explain the structure of result page in details, GI:17529185 is used as an example
of blast in following sections.
5.1 Summary
Figure 5.0 : Summary of Query Input
This is the first part shown on the top of result page. Summary gives the overall descriptions
of the input which are fasta sequence or accession number or GI numbers and database
information. Information of fasta consists of RID, query ID, description, molecular type and
query length. RID is the request ID that can be kept by user and use it to access the result
page again whenever they need within the valid period. Query ID is an ID will be assign to
the each input that user entered. Description is the description of input and will be shown if
any. Molecule type is the type of molecule of input. Query length is the total length of the
input. Database information indicates what database and program will be used in running the
blast of the input entered. There will be consisting of database name, description and
program.
38
5.2 Graphical Overview
Figure 5.1 : Graphical Overview of Blast Result
The graphical overview show the distribution of blast hits on query sequence. The numbered
red bar at the top of the figure 5.1 is represent the query sequence while the number attached
to the red bar is query coordinates. The alignment scores are defined by using color key. As
can be seen from figure 5.1, alignment scores less than 40 is represented by black color key
while alignment scores equal or more than 200 is represented by red color key. The most
similar aligned sequence are shown closet to the query sequence. In this case, there are three
aligned sequence are the most similar to the query sequence due to the high alignment score
since its colored bar has red color key from query 1 to the end. In other words, the next
following colored bars indicate the aligned sequence from database that match the query with
lower score. Mouse over the bars displays the definition line which consist definition and
39
score for that aligned sequence to be shown in the window above the graphic as figure 5.2
shown. It will also show the alignment for that sequence if click on it.
Figure 5.2 : Mouse Over On Graphical Overview
40
5.3 Description Table
Figure 5.3 : Description Table for Blast Result
The descriptions table provides a summary of the aligned sequence from database which
identified by Blast to be similar to the input query. As can be seen from figure 5.3, from left
to right, the descriptions table columns display the description, max score, total score, query
cover, ident and accession. In traditional report format, this is known as one-line descriptions
because each line in the table is composed of those seven fields which are description, max
score, total score, query cover, ident and accession.
Definition for each field
Description
Title of database sequence that matched to the query sequence.
Max Score
The maximum alignment score from that matched database sequence.
Total Score
The sum of all alignment scores of alignment segments.
Query Cover
The coverage (%) ofthe query sequence being aligned to the matched
database sequence.
E - value
The lowest expect value from that matched database sequence.
Ident
The highest percentage of identityof all pairwise alignments between
query and database sequences.
Accession
Accession number of database sequence that matched to the query
sequence.
41
5.4 Alignments
The alignments section contains the detailed pairwise alignments between query and subject
sequence in database or we know as (Sbjct). Alignment section provides statistic line which
composed of score, expect, identities, gap and strand for each pairwise alignment.
Figure 5.4 : Alignment of Blast Result
42
Figure 5.5 : Alignment of Blast Result (2)
Definition on Statistics Line
Score
Summed HSP (High Scoring Pair) score (S)
Bit Score
A normalized scorein bits(S’)
Expect (E)
Expected number of chance HSP aligns
Identities
Number and percentage of exact residue matches
Gaps
Number and percentage of gaps introduced
43
5.5 Score, bit Score and E-value Calculation
1.5.1
Score
Score (S) is a number which can be used to evaluate the relevance of a finding in biology.A
score is a numerical value that can tell the overall quality of an alignment in terms of
sequence alignments. Higher numbers of score indicate higher similarity of alignment. The
score scale for score calculation is relying on scoring system used which included
substitution matrix and gap penalty. Overall, the score indicates that the higher the score, the
best of the alignment.
For nucleotides blast,same score (positive) is given to each identical match, and a penalty
(negative) score is assigned to all mismatches. Since the scoring system may be varied from
default according to different situation, but for the purpose of highlighting the occurrence of
gaps and mismatches, the scoring system as below is used :
Match (Positive) = +1 point
Mismatch (Negative) = -2 points
Gap opening (Negative) = -2 points
Gap extension (Negative) = -1 point
Example 1 :
AAC GTT TCC AGT CCA AAT AGC TAG GC
| | | .. | | |
| .| | | . | | . | || | | |
AAC CGT TCCAGT ACA ATT ACC TAG GC
Matches (+1): 18
Mismatches (-2): 5
Gaps (opening -2, extension -1): 1, 2
Score (S) = 18 * (+1) + 5 * (-2) +1 * (– 2) + 2 * (-1) = 4
44
For amino acid (protein) blast, blosum62 substitution matrices are used to calculate score.
BLOSUM is known as Blocks Substitution Matrix.This matrix claims that score (positive) or
penalty score (negative) is given for each identical amino acid or substitution between two
amino acids. As can be seem from figure 7, identities or substitutions are not allhave equal
value. This is because blosum62 give a signification of the likelihood that a specific
substitution may occur between many proteins. So, BLOSUM 62 is used as the default matrix
in BLAST algorithm to calculate score for protein alignment. Gap opening scoring is -4 and
extension is -1.
Figure 7 : Blosum62 Substitution Matrix
45
Example 2 :
Consider this pairwise sequence alignment:
Query LENTFFVQANC
Sbjct YENITIIQSNC
The score is calculated by total up all the numbers from left to right as follows:
Query L E N T F F V Q A N C
Sbjct Y E N I T I I Q S N C
Score -1 5 6 -1 -2 0 3 5 1 6 9
Score (S) = (-1) + 5 + 6 + (-1) + (-2) + 0 + 3 +5 +1 +6 + 9
= 31
How to get score form blosum62 ?
46
5.5.2 Bit-score
Bit-score (S’) is a score(S) in log-scaled version. In BLAST, the bit-score (S')is a score
being normalized and expressed in bits.
Formulae to calculate bit-score :
WhereS = score, λ and K = constant parameters depend on the scoring system used.
Example 3 :
As referred from Example 2, we know that the score is 31. For BLOSUM62 ,λ = 0.318 and
K= 0.14. So, let substitutes the values into the equation to get the bit-score.
S' = (λ S - ln K) / ln 2
= (0.318 * 31 - ln 0.14) / ln 2
= 17.0586(4s.f)
5.5.3 E – Value (E)
E-value is an expectation value which reveals the expectation number of BLAST alignments
with Score to be seen as a result of chance. It is efficient for searching large databases to
know how easily (or rather uneasily)that an alignment could arise by chance. The higher the
Score (more significant), the lower the E-value is. E-value and Score are related, but E-value
contains more information.
47
Formulae to calculate E-value (E):
E = mn 2-s'
where m is the length of the subject sequence in database, n is the length of the query
sequence and S' is the normalized score from above.
Example 4 :
As referred from Example 3, we know that the bit-score is 17.0586. Let assume that the
length of the database sequence, m is 11 while the length of query sequence, n is 11. So, let
substitutes the values into the equation to get the E-Value.
E = mn2-s’
= 11 * 11 * 2-17.0586
= 8.86 x 10-04
There is another way to calculate E-value without having bit-score.
E = K m n e-λS
Where S is the score, λ and K are parameters that characterize the expected distribution of S
for the scoring system used, m is the length of thesubject sequence indatabase andn is the
length of the query sequence.
Example 5 :
As referred from Example 2, Score, S is 31, For BLOSUM62 ,λ = 0.318 and K= 0.14. Let
assume the length ofsubject sequence indatabase, m is 11 and the length of the query
sequence, n is 11.
E = Kmne- λS
= 0.14 * 11 * 11 * e-(0.318 * 31)
= 8.86 x 10-04
48
5.5.4 Exercise
1. For nucleotides blast, you are given the substitution matrix scoring system as below :
Gap opening = -2
Gap extension = -1
Calculate the score for each alignment pair :
a) Query AATCGTGCCTTGGACCCCTCA
Sbjct AATCCTGCCTTGGACCCGTCC
b) Query TTACGCGCTCCGGAAAGATGG
Sbjct TTACGC _CTCCGGACAGATG_
c) Query CGGGAGGCCAAAGATCTAAGC
Sbjct C_ GGAGGCC__ _ GACCTAAGC
Answer :
a) 12
b) 12
c) 8
49
2. For protein blast, based on the blosum62 substitution matrix, find out the score and E-value
for questions below.
For BLOSUM62,λ = 0.318 and K= 0.14while m and n depend on the questions below.
a) Query NLYENFVQATF
Sbjct NYAENTIQSII
b) Query LNCQEFVDTPG
Sbjct VWCGFFADTPG
C) Query CLASV-ETPMWP
Sbjct CLTSLAQTPL-P
Answer :
a)score = 20
E-value = 0.0293
b) score = 31
E-value = 8.86 x 10-04
c) score = 33
E-value = 4.69 x 10-04
50