* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Parameterization for solvent molecules around a
Computational chemistry wikipedia , lookup
Plateau principle wikipedia , lookup
Theoretical ecology wikipedia , lookup
General circulation model wikipedia , lookup
Numerical weather prediction wikipedia , lookup
History of numerical weather prediction wikipedia , lookup
Data assimilation wikipedia , lookup
Biology Monte Carlo method wikipedia , lookup
Joint Theater Level Simulation wikipedia , lookup
Solvent models wikipedia , lookup
Implicit solvation wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
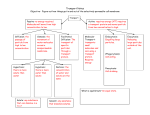
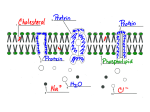
PARAMETERIZATION OF SOLVENT MOLECULES AROUND A SOLUTE Adnan Khan1 Peter Kramer1 & Rahul Godawat2 1Department of Mathematical Sciences 2Department of Chemical Engineering Rensselaer Polytechnic institute 1.THE PROBLEM 6. THE DD-I MODEL Two important lines of research in the behavior of proteins are As a first model (following Garde et.al), we guess the correct form to be - Given the sequence : predict the structure of the protein - Given the structure : predict the function of the protein dX D dt 2D dW kT bulk bulk B Want to study the functioning of a protein given the structure We use Monte Carlo methods to simulate this SDE The behavior depends on the surrounding molecules To explicitly deal with sufficient number of solvent molecules is difficult due to computational costs We calculate the diffusivity and compare it with the diffusivity obtained directly from the MD data Initial configuration for the Buckyball Water system 2. OUR APPROACH 5. BENCHMARKING We want to parameterize the state of water molecules near a protein To benchmark our code we use two exactly solvable processes As a basic model we will use dX A(X, ) B(X, )dW The Ornstein Uhlenbeck process governed by dX Xdt 2dW A is the drift vector and B is the diffusion tensor; these need to be determined, dW is white noise The Langevin process with quadratic potential, which is governed by the equation There are two approaches we use - We guess the correct form of the coefficients - We use MD data to determine the coefficients 7. THE DD-II MODEL dX Vdt dV aVdt aXdt adW We next use a more general drift-diffusion parameterization We are interested in the overdamped case which corresponds to a>>1 dX U dt D long The OU process can be considered a coarse-grained version of the Langevin process 3. GENERAL PROGRAM A measure of diffusivity which is used in the literature is We start by considering a simple solute: specifically, we consider a Buckyball (C60), as this has approximate spherical symmetry Once we have a good parameterization for water around this simple solute, we shall consider more complex solutes that mimic the geometric and chemical properties of a protein We shall then use the appropriate parameterization when simulating the dynamics of an actual protein We use inputs obtained from MD simulations to develop our parameterization X(t 2 ) X(t) X(t) x 2 D(X, ) X(t ) X(t) X(t) x 2 6 2 The parameters Ulong Dlong & Dlat are calculated directly from the MD data We then calculate the diffusivity, and compare it to the value obtained directly from the MD data We are interested in the regime where the V dynamics are coarse-grained, i.e. we want a value of such that v<< <<X where is the momentum relaxation time and is the characteristic time for the X dynamics We also use the two aformentioned exact systems to understand how to calculate the best value of for our models 8. CONCLUSIONS & FUTURE WORK We note that the DD-II model does better than DD-I model in capturing the features of the diffusivity The MD simulation was performed with a Buckyball (C60) hydrated by 1380 water molecules A 3.5 x 3.5 x 3.5 cubic box was used, which was periodic in all dimensions The simulation parameters were –Integrator time step 2fs –Temperature 300K (using Nose Hoover thermostat) lat X X I dW X We calculate this for our test models and compare the results to the exact solutions 4. MD SIMULATION The interactions were –LJ (Lennard Jones) cutoff 1nm –Particle Mesh Ewald (PME) long X X dW D X X We shall now use the DD-II model with input from MD simulation of water around more complex solutes Numerical vs Exact Diffusivity for OU and Langevin systems We hope to construct effective models for the dynamics of water around different types of proteins, using these simulations