* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Effect of shRNA knockdown of protein complex subunits on complex
Survey
Document related concepts
Transcript
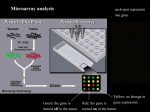
Effect of shRNA knockdown of protein complex subunits on complex formation and quantitation using SILAC technique. Mahbod R. Hajivandi; John F. Leite; Xiquan Liang; Antje Taliana; Marieke Svoboda; Marshall Pope; WP512 Invitrogen, Carlsbad, CA Results and Discussion Anti-Arp3 IgG 25.00 Time 35.00 40.00 45.00 50.00 Figure 5. shRNA knockdown of native Arp3 subunit in 293 cells stably transfected with TAPtagged Arp3 results in ~55% knockdown of native Arp3 subunit. Western blots of lysates from 293 cells stably transfected with TAP-tagged Arp3 show a knockdown of the Arp3 subunit that is consistent with SILAC experiments. LALETTVLVESYTLPDGR, 1976.05 Da DITYFIQQLLR, 1408.77 Da 705.4034 100 shRNA 989.0305 B % 707.3888 % C 990.9956 Arp3 #1 Target cleavage Figure 2. shRNA in the RNA interference pathway. 705 707 709 711 m/z 985 987 989 991 993 995 m/z 997 Methods DNTINLIHTFR, 1342.70 Da TLESSIQGLR, 1102.62 Da 552.3014 554.2891 % Figure 6. shRNA knockdown of native Arp3 subunit in 293 cells stably transfected with TAPtagged Arp3 results in knockdown of ArpC2 subunit. Western blots of lysates from 293 cells stably transfected with TAP-tagged Arp3 show a knockdown of the ArpC2 subunit resulting from shRNA directed at Arp3. Controls demonstrate that the knockdown is not the result of shRNA binding to exogenous genes non-specifically or due to non-specific transfection effects. E D % 674.4113 m/z 678 680 ELLLQPVTISR, 1267.77 Da 605.8733 634.8792 100 100 G F Anti-Arp3 IgG 636.8659 % % 607.8644 604 Light Arg 14 N Arg 4 Heavy Arg 15 N -Arg 4 Cells must be propagated for at least six doubling times to achieve 100% incorporation For example, the total cell number would be 64 x 105 cells if started with 105 cells Take small aliquot of cells (such as 108) to examine 100% incorporation in MALDI-TOF Mix cells at 1:1 ratio Figure 1. Molecular model of filament extension by the Arp 2/3 complex. (A) A model of the actin filament branches (in the pink color) was superimposed over the 2D images shown in (B). An additional representation of the Arp2/3 complex at the filament branch is shown in (C) so that the protein backbone is exposed. RNA interference (RNAi) is the preeminent gene silencing technology currently employed in mammalian reverse genetics experiments, in which a loss-of-function phenotype of a gene of known sequence is sought. In RNAi, the primary effector molecules are double-stranded short interfering RNAs (siRNAs)1. One strand of each siRNA molecule is incorporated into a cytoplasmic, multiprotein RNA-Induced Silencing Complex (RISC) and serves as a guide for locating complementary target RNAs. A RISC nuclease cleaves the target RNA within the region basepaired to the siRNA guide. The target is then subject to degradation by cytoplasmic exonucleases. Volkmann N, Amann KJ, Stoilova-McPhie S, Egile C, Winter DC, Hazelwood L, Heuser JE, Li R, Pollard TD, Hanein D. Structure of Arp2/3 complex in its activated state and in actin filament branch junctions. Science. 2001 Sep 28;293(5539):2456-9. Cell mix after control shRNA (-) Cell mix after shRNA treatment (+) TAP-Affinity enrichment of Arp 2/3 complex shRNA KO the endogenous Arp subunit to enrich complex with biotintagged transfected subunit - + SDS-PAGE/ Western tryptic digest + LCMS ShRNA Quantification via the ratio of isotopic peptide pairs in MS spectrum Figure 3. Experimental strategy of SILAC for quantification. Identical pools of cells stably transfected with biotin-tagged ArpC2 or Arp3 are differentially labeled with isotope-enriched lysine. One of the sets of cells is also treated with shRNA targeted to either native expressed ArpC2 or Arp3. After the complex is enriched by streptavidinagarose, the components are separated by SDS-PAGE followed by MS analysis for quantitation. Knockdown of the endogenous Arp subunit by shRNA should result in enhanced efficiency of Arp 2/3 complex affinity-enrichment due to competitive substitution with biotin-tagged Arp subunit. m/z Mock 556 676 ArpC2#4 554 674 LacZ (-) 552 672 pENTRU6(-) 550 LIGNMALLPIR, 1209.73 Da 670 Arp3#2 shRNA 668 Stably Transfected Cells (Biotin-tagged Arp2 or Arp3) SILAC Anti-Arp2 IgG Arp3 #1 Tandem affinity tagged (6xHis coupled to biotin)ArpC2 or Arp3 subunits were stably expressed in mammalian cells and the tag was used to purify the Arp2/3 complexes from lysates. Cell lines were transfected with shRNAs U6 Entry clones specifically targeting a second subunit of the same complex in medium containing 15N-Arg (Ajinomoto). Another culture was treated with lacZ shRNA U6Entry clones in non-labeled control medium. The cultures were combined and protein complexes purified using Streptavidin agarose. The complex subunits were digested with trypsin and digested peptides were analyzed by Q-TOF. 672.3736 100 100 Actin related proteins Arp2 and Arp3, along with 5 other proteins, form a complex (Arp2/3 complex) that binds to the side of an actin filament and nucleates the formation of a new filament branch. Filament extension by Arp2 and Arp3 spearheads the molecular mechanism that supports a wide variety of cellular processes such as cell motility, cell shape and structure. The mechanism of filament extension is shown in the figure below from the Pollard laboratory publication of the high-resolution crystal structure of the Arp 2/3 complex. 30.00 Mock 20.00 100 703 Introduction 15.00 Arp3#2 RISC loading siRNA unwinding Target recognition 10.00 ArpC2#4 5.00 Target mRNA LacZ (-) % siRNA (21-23 bp) pENTRU6(-) A Mock 100 LacZ (-) shRNA expression Nuclear export Endogenous Dicer activity ArpC2#4 shRNA RNA Arp3 #1 DNA Arp3#2 Pol III pENTRU6(-) Overview In combination with mass spectrometry, recent advances in protein tagging and purification have made it possible to isolate and characterize native complexes in high throughput. Stable isotopic labeling in cell cultures (SILAC) has been used successfully to study the dynamics of cell signal-dependent protein-protein interactions. Our goal was to develop short-hairpin RNA (shRNA) knockdown as an investigative means to study the role of the individual subunits on the activity and stoichiometry of complexes. Because many of these knockdown effects are anticipated to be subtle, we reasoned that conventional methods may not have sufficient dynamic range to discern small changes in protein expression. Instead we have employed metabolic labeling, SILAC, to precisely quantify stoichiometric changes in complex formation caused by shRNA knockdown perturbations. 606 608 610 m/z 633 635 637 639 m/z Figure 7. shRNA knockdown of native ArpC2 subunit in 293 cells stably transfected with TAPtagged ArpC2 results in knockdown of Arp3 subunit. Western blots of lysates from 293 cells stably transfected with TAP-tagged ArpC2 show a knockdown of the Arp3 subunit resulting from shRNA directed at Arp3. Controls demonstrate that the knockdown is not the result of non-specific shRNA or transfection effects. Figure 5. shRNA knockdown of native Arp3 subunit in 293 cells stably transfected with biotintagged Arp3. Total Ion Chromatogram of digested affinity enriched Arp2/3 complex from a SILAC labeling experiment (A). Trypsin digestion and RP-LC separation were performed in the presence of Invitrosol™ LC/MS. MS spectra of peaks correspond to digested peptides from the different subunits of the complex (B) ARP3 (C) ARP2 (D) AR1A (E) ARPC2 (F) AR21 (G) AR20, unexpectedly suggest that that recovery of the Arp 2/3 complex via the tagged Arp3 was knocked down ~55%. Subunit APR3. APR2. AR1A. AR34. AR21. AR20 Molecular Weight Da 47341 44732 41557 34311 20533 19654 Calculated PI 5.61 6.3 8.6 6.84 8.8 8.53 Accession Number P32391 O15142 Q92747 O15144 O15145 P59998 Sequence Coverage 44% 50% 27% 55% 38% 66% Table 1. showing the Arp 2/3 complex subunits identified and sequence coverage. Figure 8. Position of 6xHIS Biotin tag in the Arp 2/3 molecular model. Molecular model representations (left and right images are 90º to eachother) of the Arp 2/3 complex was generated based on the PDB file and using SPDB program. Both TAP tags (N-terminus for Arp3 and C-terminus for ArpC2) are located on the protein surface which could lead to interference of complex formation. Conclusions We present a new method for the analysis of protein expression knockdown and how it may be applied to determine protein complex assembly, stoichiometry and monomer turnover. Specifically, the marriage of two powerful technologies, shRNA and SILAC, combine to produce a method capable of detecting significant stoichiometric effects under even relatively subtle knockdown conditions. In this study, we focus on the Arp 2/3 complex assembly. Using differential expression analysis by SILAC, while the introduction of targeted shRNA serves as an experimental stimulus, we have quantified the degree of knockdown of specific complex subunits. We observe an unexpected reduction in complex affinity enrichment after introducing an affinity-tagged Arp complex subunit followed by an shRNA treatment to knockdown the native isomer. We confirm this phenomenon is specific to the Arp complex and validate the effect by SILAC analysis and Western blot analysis. We hypthesize the charge density associated with the TAP tag disrupts complex formation. We plan to validate this hypothesis using SILAC analysis to monitor subunit turnover upon shRNA treatment, and a modification in the affinity enrichment design. Invitrogen Corporation • 1600 Faraday Avenue • Carlsbad, California 92008 USA • Telephone: 760 603 7200 • FAX: 760 602 6500 • Toll Free Telephone: 800 955 6288 • E-mail: [email protected] • www.invitrogen.com