* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Horizontal Gene Transfer - Oxford Academic
Hospital-acquired infection wikipedia , lookup
Community fingerprinting wikipedia , lookup
Human microbiota wikipedia , lookup
Bacterial cell structure wikipedia , lookup
Marine microorganism wikipedia , lookup
Carbapenem-resistant enterobacteriaceae wikipedia , lookup
Antibiotics wikipedia , lookup
Bacterial morphological plasticity wikipedia , lookup
Staphylococcus aureus wikipedia , lookup
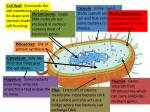
Evolution, Medicine, and Public Health [2015] pp. 193–194 emph clinical briefs: foundational doi:10.1093/emph/eov018 Horizontal Gene Transfer Alita R. Burmeister1,2,3 1 Department of Microbiology and Molecular Genetics, 2Department of Ecology, Evolutionary Biology, and Behavior Program, and 3BEACON Center for the Study of Evolution in Action, Michigan State University DEFINITION AND BACKGROUND Horizontal gene transfer (HGT) is the movement of genetic information between organisms, a process that includes the spread of antibiotic resistance genes among bacteria (except for those from parent to offspring), fueling pathogen evolution. Many resistance genes evolved long ago in natural environments with no anthropogenic influence but these genes are now rapidly spreading to and among human pathogens. HGT occurs by three well-understood genetic mechanisms (Fig. 1): . Transformation: Bacteria take up DNA from their environment . Conjugation: Bacteria directly transfer genes to another cell . Transduction: Bacteriophages (bacterial viruses) move genes from one cell to another EXAMPLES IN HUMAN BIOLOGY AND PUBLIC HEALTH EXAMPLES IN CLINICAL MEDICINE Antibiotic use in human medicine and agriculture continually selects for resistant bacteria [2, 6]. For example, tetracycline and lactams commonly fed to animals provide a selective environment for tetracycline and methicillin resistance. Genes conferring resistance to these antibiotics have horizontally transferred into a sensitive human-associated Staphylococcus aureus strain, resulting in methicillin-resistant strain CC398 [7]. After a strain gains resistance by HGT, the bacteria proliferate and continue to evolve as they move among patients and hospitals [1]. This process occurs in many bacterial lineages, resulting in diverse populations of a variety of strains, such as USA300 [5]. Ongoing HGT poses a problem for clinical surveillance and treatment. Bacterial populations evolve rapidly, resulting in diversity that necessitates individual screening to determine effective treatments and to detect new strains, such as methicillin and high level vancomycin resistant S.aureus (MRSA and VRSA) [2]. Even when new drugs and diagnostic tools become available, the persistence of HGT will require ongoing surveillance for newly resistant pathogens, leaving practitioners and researchers racing with evolution. funding This material is based upon work supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. DGE1424871. Once transferred, the genes and pathogens continue to evolve, often resulting in bacteria with greater resistance [1, 2, 3, 4]. All genes—not just those causing drug resistance—may be horizontally transferred and proliferate by natural selection, including virulence determinants [5]. references 1. Harris SR, Feil EJ, Holden MTG et al. Evolution of MRSA during hospital transmission and intercontinental spread. Science 2010;327:469–74. 2. Lindsay, J. A. Hospital-associated MRSA and antibiotic resistance-what have we learned from genomics? Int J Med Microbiol 2013;303:318–23 3. McCarthy AJ, Loeffler A, Witney AA et al. Extensive horizontal gene transfer during Staphylococcus aureus co-colonization in vivo. Genome Biol Evol 2014;6:2697–708. 4. Stanczak-Mrozek KI, Manne A, Knight GM et al. Figure 1. Mechanisms of bacterial horizontal Within-host diversity of MRSA antimicrobial gene transfer resistances. J Antimicrob Chemother 2015;70: 2191–2198. 5. McDougal LK, Steward CD, Killgore GE et al. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates ß The Author(s) 2015. Published by Oxford University Press on behalf of the Foundation for Evolution, Medicine, and Public Health. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted reuse, distribution, and reproduction in any medium, provided the original work is properly cited. 194 Evolution, Medicine, and Public Health | from the United States: establishing a national database. J Clin Microbiol 2003;41:5113–20. 6. US Department of Health and Human Services. 7. Price LB, Stegger M, Hasman H et al. Antibiotic Resistance Threats in the United States. Staphylococcus aureus CC398: host adaptation Atlanta: CDC, 2013; p, 11–14. and emergence of methicillin resistance in livestock. MBio 2012;3.