PDF
... development from nutrition In flies and other insects, starvation delays metamorphosis from larval to pupal stages, but once a certain size or weight – the critical weight – is reached, development proceeds independently of nutrition. What regulates this switch? On p. 2345, Christen Mirth and collea ...
... development from nutrition In flies and other insects, starvation delays metamorphosis from larval to pupal stages, but once a certain size or weight – the critical weight – is reached, development proceeds independently of nutrition. What regulates this switch? On p. 2345, Christen Mirth and collea ...
PDF
... development from nutrition In flies and other insects, starvation delays metamorphosis from larval to pupal stages, but once a certain size or weight – the critical weight – is reached, development proceeds independently of nutrition. What regulates this switch? On p. 2345, Christen Mirth and collea ...
... development from nutrition In flies and other insects, starvation delays metamorphosis from larval to pupal stages, but once a certain size or weight – the critical weight – is reached, development proceeds independently of nutrition. What regulates this switch? On p. 2345, Christen Mirth and collea ...
Word of the Day
... One-stranded Instead of Thymine, RNA has Uracil Three types of RNA: mRNA –carries copies of DNA from nucleus to the cytoplasm .(messenger RNA) tRNA – folded RNA that connects to mRNA to connect amino acids. (transport) ...
... One-stranded Instead of Thymine, RNA has Uracil Three types of RNA: mRNA –carries copies of DNA from nucleus to the cytoplasm .(messenger RNA) tRNA – folded RNA that connects to mRNA to connect amino acids. (transport) ...
A Bioinformatics Tool for Analyzing G
... Gene expression control (since not all mRNA become proteins) Internal Ribosomal Entry Sites (IRES) • Allows entry of ribosomes to start translation not at the beginning of the 5’ UTR ...
... Gene expression control (since not all mRNA become proteins) Internal Ribosomal Entry Sites (IRES) • Allows entry of ribosomes to start translation not at the beginning of the 5’ UTR ...
Document
... • Modified bases arise from chemical changes made to the four standard bases after transcription. (tRNA-modifying enzymes) • Common secondary structure – the cloverleaf structure ...
... • Modified bases arise from chemical changes made to the four standard bases after transcription. (tRNA-modifying enzymes) • Common secondary structure – the cloverleaf structure ...
Dicer-Like
... RNA interference • Dicer and Dicer-Like (DCL) enzymes are involved in RNA interference (RNAi) • Nontranslated RNA fragments bind to mRNA and prevent translation into a protein ...
... RNA interference • Dicer and Dicer-Like (DCL) enzymes are involved in RNA interference (RNAi) • Nontranslated RNA fragments bind to mRNA and prevent translation into a protein ...
7 - Nature
... 17p13.3 between markers D17S1866 and D17S1574 in cancers. (b) Genomic organization of the human miR-22 locus. (c) RNA editing of miR-22 precursor. Bold bases form mature miR22; boxed sequence is miR-22 seed region; red bases with arrows are prone to editing. (d) Phylogenetic conservation of the non- ...
... 17p13.3 between markers D17S1866 and D17S1574 in cancers. (b) Genomic organization of the human miR-22 locus. (c) RNA editing of miR-22 precursor. Bold bases form mature miR22; boxed sequence is miR-22 seed region; red bases with arrows are prone to editing. (d) Phylogenetic conservation of the non- ...
Proteome and Gene Expression Analysis
... • First, we’ll talk about how to find out what genes are being transcribed in the cell. – This is often referred (somewhat misleadingly) to gene “expression”. ...
... • First, we’ll talk about how to find out what genes are being transcribed in the cell. – This is often referred (somewhat misleadingly) to gene “expression”. ...
7.5 Eukaryotic Genome Regulation
... 4. Regulation of mRNA degradation • Life span of mRNA determines amount of protein synthesis – mRNA can last from hours to weeks ...
... 4. Regulation of mRNA degradation • Life span of mRNA determines amount of protein synthesis – mRNA can last from hours to weeks ...
Regulation of Gene Expression
... Prokaryote gene expression typically is regulated by an operon, the collection of controlling sites adjacent to polycistronic proteincoding sequences. ...
... Prokaryote gene expression typically is regulated by an operon, the collection of controlling sites adjacent to polycistronic proteincoding sequences. ...
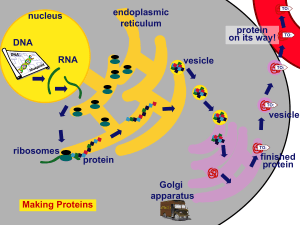
Protein Synthesis Notes
... A. What Is It? 1. RIBO-NUCLEIC ACID - The “Middle-Man” between DNA (nucleus) & the ribosomes (cytoplasm). 2. Structure a. Ribose (Sugar) b. Single-stranded, not double. c. Thymine is replaced by URACIL. - Adenine binds with Uracil. ...
... A. What Is It? 1. RIBO-NUCLEIC ACID - The “Middle-Man” between DNA (nucleus) & the ribosomes (cytoplasm). 2. Structure a. Ribose (Sugar) b. Single-stranded, not double. c. Thymine is replaced by URACIL. - Adenine binds with Uracil. ...
No Slide Title
... Transfer RNA (tRNA) leaves the nucleus, binds to the amino acid specified by it’s anticodon and transfers it to the ribisome where it meets up with mRNA to assemble a protein. ...
... Transfer RNA (tRNA) leaves the nucleus, binds to the amino acid specified by it’s anticodon and transfers it to the ribisome where it meets up with mRNA to assemble a protein. ...
Protein Synthesis
... The mRNA is single stranded. The mRNA uses U instead of T. (A,U,G,C) The DNA zips back together. ...
... The mRNA is single stranded. The mRNA uses U instead of T. (A,U,G,C) The DNA zips back together. ...
ch 19 gene expression in eukaryotes
... – folding, cleaving, adding sugar groups, targeting for transport ...
... – folding, cleaving, adding sugar groups, targeting for transport ...
Schol Biol: Genetics
... Genes within DNA are a code for proteins (proteins do the actual work in our bodies) In cells, genes are copied into a message form (messenger RNA/mRNA) to then be used by the protein making factories (ribosomes) The copying for any particular gene is switched on and off as required Specific target ...
... Genes within DNA are a code for proteins (proteins do the actual work in our bodies) In cells, genes are copied into a message form (messenger RNA/mRNA) to then be used by the protein making factories (ribosomes) The copying for any particular gene is switched on and off as required Specific target ...
Discovery of Muscle Atrophy Gene Regulatory Network Using
... Brandon King Gilberto Hernandez, M.D. ...
... Brandon King Gilberto Hernandez, M.D. ...
MBch15
... Perceiving order in the makeup of the code The genetic code might have evolved in a way to minimize deleterious effects of mutations. 1. Codons with pyrimidines in the 2nd position mostly specify hydrophobic amino acids; while those with purines in the 2nd ...
... Perceiving order in the makeup of the code The genetic code might have evolved in a way to minimize deleterious effects of mutations. 1. Codons with pyrimidines in the 2nd position mostly specify hydrophobic amino acids; while those with purines in the 2nd ...
BioSc 231 2001 Exam5
... A. is determined by genetically programmed changes in the cells B. is imposed by the position of cells in the embryo C. is mediated by physical interaction between cells D. is mediated by morphogens _____ Mutations that cause cells to undergo developmental fates characteristic of other types of cell ...
... A. is determined by genetically programmed changes in the cells B. is imposed by the position of cells in the embryo C. is mediated by physical interaction between cells D. is mediated by morphogens _____ Mutations that cause cells to undergo developmental fates characteristic of other types of cell ...
Non-coding RNA | Principles of Biology from Nature Education
... the RNA molecule folds back on itself and forms a number of small, doublestranded hairpin loops. The RNA molecule is cleaved into shorter doublestranded segments by an enzyme complex known as the Microprocessor complex. Each segment then leaves the nucleus and enters the cytoplasm, where another enz ...
... the RNA molecule folds back on itself and forms a number of small, doublestranded hairpin loops. The RNA molecule is cleaved into shorter doublestranded segments by an enzyme complex known as the Microprocessor complex. Each segment then leaves the nucleus and enters the cytoplasm, where another enz ...
RNA interference 1. The central dogma 3. The RNAi mechanism
... mRNA is cleaved and destroyed. No protein can be synthesized. ...
... mRNA is cleaved and destroyed. No protein can be synthesized. ...
MicroRNA
A micro RNA (abbreviated miRNA) is a small non-coding RNA molecule (containing about 22 nucleotides) found in plants, animals, and some viruses, which functions in RNA silencing and post-transcriptional regulation of gene expression.Encoded by eukaryotic nuclear DNA in plants and animals and by viral DNA in certain viruses whose genome is based on DNA, miRNAs function via base-pairing with complementary sequences within mRNA molecules. As a result, these mRNA molecules are silenced by one or more of the following processes: 1) cleavage of the mRNA strand into two pieces, 2) destabilization of the mRNA through shortening of its poly(A) tail, and 3) less efficient translation of the mRNA into proteins by ribosomes. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1000 miRNAs, which are abundant in many mammalian cell types and appear to target about 60% of the genes of humans and other mammals.miRNAs are well conserved in both plants and animals, and are thought to be a vital and evolutionarily ancient component of genetic regulation. While core components of the microRNA pathway are conserved between plants and animals, miRNA repertoires in the two kingdoms appear to have emerged independently with different primary modes of action. Plant miRNAs usually have near-perfect pairing with their mRNA targets, which induces gene repression through cleavage of the target transcripts. In contrast, animal miRNAs are able to recognize their target mRNAs by using as little as 6–8 nucleotides (the seed region) at the 5' end of the miRNA, which is not enough pairing to induce cleavage of the target mRNAs. Combinatorial regulation is a feature of miRNA regulation in animals. A given miRNA may have hundreds of different mRNA targets, and a given target might be regulated by multiple miRNAs.The first miRNA was discovered in the early 1990s. However, miRNAs were not recognized as a distinct class of biological regulators until the early 2000s. Since then, miRNA research has revealed different sets of miRNAs expressed in different cell types and tissuesand has revealed multiple roles for miRNAs in plant and animal development and in many other biological processes. Aberrant expression of miRNAs has been implicated in numerous disease states, and miRNA-based therapies are under investigation.Estimates of the average number of unique messenger RNAs that are targets for repression by a typical microRNA vary, depending on the method used to make the estimate, but several approaches show that mammalian miRNAs can have many unique targets. For example, an analysis of the miRNAs highly conserved in vertebrate animals shows that each of these miRNAs has, on average, roughly 400 conserved targets. Likewise, experiments show that a single miRNA can reduce the stability of hundreds of unique messenger RNAs, and other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold).