Increased transcription rates correlate with increased reversion rates
... Starving an auxotroph for its required amino acid will derepress/deattenuate the operon (including the defective gene) and reversion rates to wild-type should increase if transcription enhances mutations. Cells with the ability to activate transcription in the presence of ppGpp (relA+) would be expe ...
... Starving an auxotroph for its required amino acid will derepress/deattenuate the operon (including the defective gene) and reversion rates to wild-type should increase if transcription enhances mutations. Cells with the ability to activate transcription in the presence of ppGpp (relA+) would be expe ...
Testing the ABC floral-organ identity model: expression of
... Neither AP3 nor PI are expressed past stage 4 in either Ap3 or Pi lossof-function mutants. Expression of PI throughout the flower leads to expression and function of AP3 in whorls 1+2+3 and expression of AP3 throughout the flower leads to expression and function of PI in whorls 2+3+4. ...
... Neither AP3 nor PI are expressed past stage 4 in either Ap3 or Pi lossof-function mutants. Expression of PI throughout the flower leads to expression and function of AP3 in whorls 1+2+3 and expression of AP3 throughout the flower leads to expression and function of PI in whorls 2+3+4. ...
Bicoid associates with the 5 -cap-bound complex of caudal mRNA
... Figure 1. BCD copurifies with 5⬘-cap-bound proteins. Cytoplasmic protein extracts of young embryos were affinity-purified using a cap-analog m7GTP-sepharose resin (Edery et al. 1988). (a) Silver-stained SDS-PAGE of affinity-purified proteins contained within cytoplasmic extracts of wild-type and bcd ...
... Figure 1. BCD copurifies with 5⬘-cap-bound proteins. Cytoplasmic protein extracts of young embryos were affinity-purified using a cap-analog m7GTP-sepharose resin (Edery et al. 1988). (a) Silver-stained SDS-PAGE of affinity-purified proteins contained within cytoplasmic extracts of wild-type and bcd ...
Xdbx inhibits neurogenesis - Development
... high homology with the homeodomain region of the zebrafish hlx1 gene (Fig. 1B; Fjose et al., 1994), the only domain of hlx1 for which published sequence is available. In contrast to these extensive homologies, the homology between Xdbx and other members of the hlx class of homeodomain genes is speci ...
... high homology with the homeodomain region of the zebrafish hlx1 gene (Fig. 1B; Fjose et al., 1994), the only domain of hlx1 for which published sequence is available. In contrast to these extensive homologies, the homology between Xdbx and other members of the hlx class of homeodomain genes is speci ...
Regulation of Na/K-ATPase β1 -subunit gene
... Ca2+ and in the force of contraction. This effect on cardiac contractility is the basis of the major therapeutic use of these drugs in the treatment of congestive heart failure. Recently, using cultured neonatal rat cardiac myocytes, we showed that partial inhibition of Na/K-ATPase by ouabain also a ...
... Ca2+ and in the force of contraction. This effect on cardiac contractility is the basis of the major therapeutic use of these drugs in the treatment of congestive heart failure. Recently, using cultured neonatal rat cardiac myocytes, we showed that partial inhibition of Na/K-ATPase by ouabain also a ...
Powerpoint template for scientific posters (Swarthmore College)
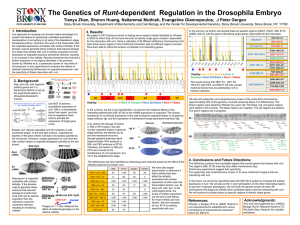
... Our approach for studying runt function takes advantage of a system that allows for genetically controlled quantitative manipulation of runt activity in all cells of the blastoderm stage Drosophila embryo. Central to this work is the observation that the engrailed expression correlates with embryo l ...
... Our approach for studying runt function takes advantage of a system that allows for genetically controlled quantitative manipulation of runt activity in all cells of the blastoderm stage Drosophila embryo. Central to this work is the observation that the engrailed expression correlates with embryo l ...
Novel regulatio pendage transformation
... developmental stages, respectively. The maxillipeds are false-coloured in red and the legs). Therefore, the originally established second thoracic appendages/ first pereopods are false-coloured in green. The first thoracic tagma boundary between the second maxilla limbs (A) can be seen in their tran ...
... developmental stages, respectively. The maxillipeds are false-coloured in red and the legs). Therefore, the originally established second thoracic appendages/ first pereopods are false-coloured in green. The first thoracic tagma boundary between the second maxilla limbs (A) can be seen in their tran ...
Adrenomedullin Gene Expression Is Developmentally Regulated and
... was used as the DNA template for PCR mutagenesis to produce mutated Adm promoter constructs, which lack one (pMutA or pMutB) or both of the potential HIF-1 binding sites (pMutAB). In pMutA, the core HIF-1 site (59-ACGT-39) at -1095 upstream of the transcription binding site was replaced with 59-AAAT ...
... was used as the DNA template for PCR mutagenesis to produce mutated Adm promoter constructs, which lack one (pMutA or pMutB) or both of the potential HIF-1 binding sites (pMutAB). In pMutA, the core HIF-1 site (59-ACGT-39) at -1095 upstream of the transcription binding site was replaced with 59-AAAT ...
VegT activates Bix4 to specify endodermal
... within the oocyte. Prominent among these are the RNAs encoding Vg1, a member of the TGF-β family (Rebagliati et al., 1985; Weeks and Melton, 1987), and VegT (Horb and Thomsen, 1997; Lustig et al., 1996; Stennard et al., 1996; Zhang and King, 1996) a member of the T-box family of transcription factor ...
... within the oocyte. Prominent among these are the RNAs encoding Vg1, a member of the TGF-β family (Rebagliati et al., 1985; Weeks and Melton, 1987), and VegT (Horb and Thomsen, 1997; Lustig et al., 1996; Stennard et al., 1996; Zhang and King, 1996) a member of the T-box family of transcription factor ...
Whole-transcriptome RNAseq analysis from minute amount of total
... RNA-seq methods, some of which are described in literature (10): library complexity, the number of unique reads, ribosomal RNA read-count in comparison to total reads, reproducibility, evenness of coverage at annotated transcripts, performance at 50 - and 30 -ends and cross-platform consistency. In ...
... RNA-seq methods, some of which are described in literature (10): library complexity, the number of unique reads, ribosomal RNA read-count in comparison to total reads, reproducibility, evenness of coverage at annotated transcripts, performance at 50 - and 30 -ends and cross-platform consistency. In ...
Molecular Biology of Transcription and RNA Processing
... nuclear RNA (snRNA) of various types is found in the nucleus of eukaryotic cells, where it participates in mRNA processing. Certain snRNAs unite with nuclear proteins to form ribonucleoprotein complexes that are responsible for intron removal. We discuss these activities in later sections of this c ...
... nuclear RNA (snRNA) of various types is found in the nucleus of eukaryotic cells, where it participates in mRNA processing. Certain snRNAs unite with nuclear proteins to form ribonucleoprotein complexes that are responsible for intron removal. We discuss these activities in later sections of this c ...
Full-Text PDF
... of allergens from their sources [8]. Obtaining the allergen-encoded cDNAs, three different methods are applied. The first is based on immunoscreening, the second is based on amino acids determination, and the third usesthe sequence similarity at the protein and nucleic acid level. The workflow for t ...
... of allergens from their sources [8]. Obtaining the allergen-encoded cDNAs, three different methods are applied. The first is based on immunoscreening, the second is based on amino acids determination, and the third usesthe sequence similarity at the protein and nucleic acid level. The workflow for t ...
Centipede Hox genes - Development
... developmental genetics facilitates this, as it provides some basis for speculating about the developmental processes of other arthropods. The body plan of Drosophila is encoded in part by the patterned expression of a set of transcription factors called the Hox proteins, which divide the embryo into ...
... developmental genetics facilitates this, as it provides some basis for speculating about the developmental processes of other arthropods. The body plan of Drosophila is encoded in part by the patterned expression of a set of transcription factors called the Hox proteins, which divide the embryo into ...
PPT
... but rare in population; may not be reflective of common disease. Also, hard to collect family data. ...
... but rare in population; may not be reflective of common disease. Also, hard to collect family data. ...
RIBOZYMES
... Peptidyl transferase 23S rRNA,RNase P, Group I and Group II introns, G1R1 branching ribozyme,Leadzyme, Hairpin ribozyme, Hammerhead ribozyme, HDV ribozyme, Mammalian CPEB3 ribozyme, VS ribozyme, glmS ribozyme, CoTC ribozyme Artificial ribozymes are synthesised in the laboratory based on the dual ...
... Peptidyl transferase 23S rRNA,RNase P, Group I and Group II introns, G1R1 branching ribozyme,Leadzyme, Hairpin ribozyme, Hammerhead ribozyme, HDV ribozyme, Mammalian CPEB3 ribozyme, VS ribozyme, glmS ribozyme, CoTC ribozyme Artificial ribozymes are synthesised in the laboratory based on the dual ...
New Construct Approaches for Efficient Gene Silencing in Plants
... Figure 2. RNA analysis of gus plants containing a terminator-free silencing construct. A, Diagrams of expression cassettes for the gus gene and the silencing constructs of pSIM717 and 715. The position of introns is indicated with horizontal striped bars. Primers (pr1–pr4) used for RT-PCR are shown ...
... Figure 2. RNA analysis of gus plants containing a terminator-free silencing construct. A, Diagrams of expression cassettes for the gus gene and the silencing constructs of pSIM717 and 715. The position of introns is indicated with horizontal striped bars. Primers (pr1–pr4) used for RT-PCR are shown ...
The RNA world meets behavior: AfiI pre
... destined to be modified in a glutamine codon (CAG). (b) Further transcription unveils the intronic editing site complementary sequence (ECS), which somehow base-pairs and forms a helix with the coding region, looping out intervening sequences. Exonic sequences are in green, and intronic sequences ar ...
... destined to be modified in a glutamine codon (CAG). (b) Further transcription unveils the intronic editing site complementary sequence (ECS), which somehow base-pairs and forms a helix with the coding region, looping out intervening sequences. Exonic sequences are in green, and intronic sequences ar ...
MOL WS 2016 Handout T3 Metabolism RNA world
... RNase P is unique from other RNases in that it is a ribozyme – a ribonucleic acid that acts as a catalyst in the same way that a protein based enzyme would. Its function is to cleave off an extra, or precursor, sequence of RNA on tRNA molecules. Bacterial RNase P has two components: an RNA chain, ca ...
... RNase P is unique from other RNases in that it is a ribozyme – a ribonucleic acid that acts as a catalyst in the same way that a protein based enzyme would. Its function is to cleave off an extra, or precursor, sequence of RNA on tRNA molecules. Bacterial RNase P has two components: an RNA chain, ca ...
Counting Small RNA in Pathogenic Bacteria
... key insights into how messenger RNA (mRNA) are produced,1 the regulatory role of long noncoding RNA,2 and determination of cell fate by spatial patterning. One key method used to obtain single cell expression data is singlemolecule fluorescence hybridization (smFISH), a technique first introduced for ...
... key insights into how messenger RNA (mRNA) are produced,1 the regulatory role of long noncoding RNA,2 and determination of cell fate by spatial patterning. One key method used to obtain single cell expression data is singlemolecule fluorescence hybridization (smFISH), a technique first introduced for ...
Kallikrein-like prorenin-converting enzymes in inbred
... Tissue preparation. Part of the surgically removed submandibular gland was frozen in liquid nitrogen for further use, whereas the rest of it was fixed in chilled phosphate buffered saline (PBS) containing 3% paraformaldehyde and 0.02% diethylpyrocarbonate for 2–3 h. Preparation of tissue extract. One ...
... Tissue preparation. Part of the surgically removed submandibular gland was frozen in liquid nitrogen for further use, whereas the rest of it was fixed in chilled phosphate buffered saline (PBS) containing 3% paraformaldehyde and 0.02% diethylpyrocarbonate for 2–3 h. Preparation of tissue extract. One ...
Editorial Noncoding RNAs
... the microRNAs (miRNAs), controls intricate networks of gene expression via post-transcriptional mechanisms. Originally described in animals by V. Ambros and G. Ruvkin in 1993, microRNAs are now recognized to impact human development and health in many ways. Another distinct and rapidly growing class ...
... the microRNAs (miRNAs), controls intricate networks of gene expression via post-transcriptional mechanisms. Originally described in animals by V. Ambros and G. Ruvkin in 1993, microRNAs are now recognized to impact human development and health in many ways. Another distinct and rapidly growing class ...
Probing the evolution of appendage specialization by
... appendage morphology, we used PhHS to misexpress each of the 2 identified splice variants of Parhyale Ubx (PhUbx isoforms I and II), which differ in their first N-terminal amino acids (21). Misexpression in embryos from stable transgenic lines that express uniform low levels of PhUbx-I or PhUbx-II u ...
... appendage morphology, we used PhHS to misexpress each of the 2 identified splice variants of Parhyale Ubx (PhUbx isoforms I and II), which differ in their first N-terminal amino acids (21). Misexpression in embryos from stable transgenic lines that express uniform low levels of PhUbx-I or PhUbx-II u ...
PDF file
... a comparable concentration of poly(Glu4:Tyr1), the best phosphoprotein substrate tested. With artificial protein substrates, the specific activity of GST-PIR1 is about 5 orders of magnitude lower than that of PTP 1B (31), a typical tyrosine-specific phosphatase and at least 10-fold lower than that o ...
... a comparable concentration of poly(Glu4:Tyr1), the best phosphoprotein substrate tested. With artificial protein substrates, the specific activity of GST-PIR1 is about 5 orders of magnitude lower than that of PTP 1B (31), a typical tyrosine-specific phosphatase and at least 10-fold lower than that o ...
Genome-wide characteristics of sequence coverage by next
... “The most likely explanation for why genes for common diseases have not been found is that, with few exceptions, they do not exist. …., if inherited genes are not to blame for our commonest illnesses, can we find out what is? “ ...
... “The most likely explanation for why genes for common diseases have not been found is that, with few exceptions, they do not exist. …., if inherited genes are not to blame for our commonest illnesses, can we find out what is? “ ...
MicroRNA
A micro RNA (abbreviated miRNA) is a small non-coding RNA molecule (containing about 22 nucleotides) found in plants, animals, and some viruses, which functions in RNA silencing and post-transcriptional regulation of gene expression.Encoded by eukaryotic nuclear DNA in plants and animals and by viral DNA in certain viruses whose genome is based on DNA, miRNAs function via base-pairing with complementary sequences within mRNA molecules. As a result, these mRNA molecules are silenced by one or more of the following processes: 1) cleavage of the mRNA strand into two pieces, 2) destabilization of the mRNA through shortening of its poly(A) tail, and 3) less efficient translation of the mRNA into proteins by ribosomes. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1000 miRNAs, which are abundant in many mammalian cell types and appear to target about 60% of the genes of humans and other mammals.miRNAs are well conserved in both plants and animals, and are thought to be a vital and evolutionarily ancient component of genetic regulation. While core components of the microRNA pathway are conserved between plants and animals, miRNA repertoires in the two kingdoms appear to have emerged independently with different primary modes of action. Plant miRNAs usually have near-perfect pairing with their mRNA targets, which induces gene repression through cleavage of the target transcripts. In contrast, animal miRNAs are able to recognize their target mRNAs by using as little as 6–8 nucleotides (the seed region) at the 5' end of the miRNA, which is not enough pairing to induce cleavage of the target mRNAs. Combinatorial regulation is a feature of miRNA regulation in animals. A given miRNA may have hundreds of different mRNA targets, and a given target might be regulated by multiple miRNAs.The first miRNA was discovered in the early 1990s. However, miRNAs were not recognized as a distinct class of biological regulators until the early 2000s. Since then, miRNA research has revealed different sets of miRNAs expressed in different cell types and tissuesand has revealed multiple roles for miRNAs in plant and animal development and in many other biological processes. Aberrant expression of miRNAs has been implicated in numerous disease states, and miRNA-based therapies are under investigation.Estimates of the average number of unique messenger RNAs that are targets for repression by a typical microRNA vary, depending on the method used to make the estimate, but several approaches show that mammalian miRNAs can have many unique targets. For example, an analysis of the miRNAs highly conserved in vertebrate animals shows that each of these miRNAs has, on average, roughly 400 conserved targets. Likewise, experiments show that a single miRNA can reduce the stability of hundreds of unique messenger RNAs, and other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold).