Lecture 34: NMR spectroscopy
... Instead of quoting precise frequency position, we use a common reference such TMS (Tetra-methyl silane) proton signal position and calculate the relative positions of other types of nuclei by defining the chemical shift scale as follows: ...
... Instead of quoting precise frequency position, we use a common reference such TMS (Tetra-methyl silane) proton signal position and calculate the relative positions of other types of nuclei by defining the chemical shift scale as follows: ...
BY 330 Spring 2015Worksheet 3 Draw a protein made up of two
... 2. Explain why the cell only makes a small number of the possible proteins, despite the fact that many options are available. ...
... 2. Explain why the cell only makes a small number of the possible proteins, despite the fact that many options are available. ...
Proteins and Nucleic Acids Proteins (pp.46-48) Monomer
... Outline of Information to pull out of pp. 46-50 in Text book ...
... Outline of Information to pull out of pp. 46-50 in Text book ...
1. Overview
... – Heavy atom replacement – make a landmark – Ex: Selenomethionine • Plenty of computer algorithms now ...
... – Heavy atom replacement – make a landmark – Ex: Selenomethionine • Plenty of computer algorithms now ...
Knuffke Prezi- Macromolecules
... Organic Compounds Substances which contain Carbon. All living things contain Carbon, it is known as the backbone of life. ...
... Organic Compounds Substances which contain Carbon. All living things contain Carbon, it is known as the backbone of life. ...
A1980JQ46200001
... Neuberger’s suggestion that some of ovalbumin’s tyrosyl residues were Hbonded to carboxylate sidechains in the protein. 1 My results appeared to rule out all but very weak associations. “When I moved on to work in John Edsall’s laboratory at Harvard it was inevitable that, in addition to my primary ...
... Neuberger’s suggestion that some of ovalbumin’s tyrosyl residues were Hbonded to carboxylate sidechains in the protein. 1 My results appeared to rule out all but very weak associations. “When I moved on to work in John Edsall’s laboratory at Harvard it was inevitable that, in addition to my primary ...
ProteinChipâ technology is one of the most exciting advancements
... ProteinChip technology is one of the most exciting advancements in protein analysis in the last 5 years. The Protein Biology SystemTM (PBS) combines the power of mass analysis with chromatography surfaces on an integrated platform. The PBS can easily be used by biologists, biochemists, and clinicia ...
... ProteinChip technology is one of the most exciting advancements in protein analysis in the last 5 years. The Protein Biology SystemTM (PBS) combines the power of mass analysis with chromatography surfaces on an integrated platform. The PBS can easily be used by biologists, biochemists, and clinicia ...
Chemistry 160 Protein Structure Homework
... 4. Describe the 4 levels of protein structure. 5. Describe 3 types of interactions that stabilize protein structure. 6. What drives protein folding? 7. Give two ways amino acid sequences are determined. 8. A small protein was cleaved in two separate experiments by chymotrypsin and by trypsin. The ch ...
... 4. Describe the 4 levels of protein structure. 5. Describe 3 types of interactions that stabilize protein structure. 6. What drives protein folding? 7. Give two ways amino acid sequences are determined. 8. A small protein was cleaved in two separate experiments by chymotrypsin and by trypsin. The ch ...
Hands-on Exercise: Locating Protein Information
... Find the homologous mouse and fruit fly orthologs of this human protein and report the % protein shared identity. ...
... Find the homologous mouse and fruit fly orthologs of this human protein and report the % protein shared identity. ...
Test 2 - HCC Learning Web
... 16. What is meant for protein to be denatured? 17. What type of linkages join glucose, amino acids, fatty acids and nucleotides together? 18. The label on a container of margarine lists "hydrogenated vegetable oil" as the major ingredient. What is the result of adding hydrogens to vegetable oil? 19. ...
... 16. What is meant for protein to be denatured? 17. What type of linkages join glucose, amino acids, fatty acids and nucleotides together? 18. The label on a container of margarine lists "hydrogenated vegetable oil" as the major ingredient. What is the result of adding hydrogens to vegetable oil? 19. ...
Slide 1 - AccessMedicine
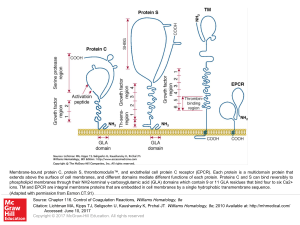
... Membrane-bound protein C, protein S, thrombomodulin™, and endothelial cell protein C receptor (EPCR). Each protein is a multidomain protein that extends above the surface of cell membranes, and different domains mediate different functions of each protein. Proteins C and S can bind reversibly to pho ...
... Membrane-bound protein C, protein S, thrombomodulin™, and endothelial cell protein C receptor (EPCR). Each protein is a multidomain protein that extends above the surface of cell membranes, and different domains mediate different functions of each protein. Proteins C and S can bind reversibly to pho ...
Nuclear Magnetic Resonance (NMR)
... the nuclear Eigenstates split in energy. The energy difference between these states is proportional to the magnetic field strength and depends also on the so-called gyromagnetic ratio. Transitions between those states can be resonantly induced by application of electromagnetic radiation of the appro ...
... the nuclear Eigenstates split in energy. The energy difference between these states is proportional to the magnetic field strength and depends also on the so-called gyromagnetic ratio. Transitions between those states can be resonantly induced by application of electromagnetic radiation of the appro ...
No Slide Title
... acceptor atom is (the C=O oxygen). If we miss-place one of these we screw up big time - Since we are basically cyclizing the peptide, there is no way we can get the right structure. • If we decide that it’s reasonable to use a H-bonding energy penalty, we can put it into the force field more or less ...
... acceptor atom is (the C=O oxygen). If we miss-place one of these we screw up big time - Since we are basically cyclizing the peptide, there is no way we can get the right structure. • If we decide that it’s reasonable to use a H-bonding energy penalty, we can put it into the force field more or less ...
Student worksheet for Proteins
... Student—please print this worksheet and complete it as you interact with the tutorial. The completed worksheet should be turned in to your assigned teacher. Tutorial: Proteins Protein Definition: (PRO-teen) A molecule made up of amino acids that are needed for the body to function properly. 1. Prote ...
... Student—please print this worksheet and complete it as you interact with the tutorial. The completed worksheet should be turned in to your assigned teacher. Tutorial: Proteins Protein Definition: (PRO-teen) A molecule made up of amino acids that are needed for the body to function properly. 1. Prote ...
For complex multicellular organisms to function, individual
... immune system, and allows for wound healing. All of the contacts involved in these processes are made through specialized molecules on the surface of cells known as adhesion proteins. These are particularly interesting from the perspective of statistical physics, since protein bonds are never perman ...
... immune system, and allows for wound healing. All of the contacts involved in these processes are made through specialized molecules on the surface of cells known as adhesion proteins. These are particularly interesting from the perspective of statistical physics, since protein bonds are never perman ...
Nuclear magnetic resonance spectroscopy of proteins
Nuclear magnetic resonance spectroscopy of proteins (usually abbreviated protein NMR) is a field of structural biology in which NMR spectroscopy is used to obtain information about the structure and dynamics of proteins, and also nucleic acids, and their complexes. The field was pioneered by Richard R. Ernst and Kurt Wüthrich at the ETH, and by Ad Bax, Marius Clore and Angela Gronenborn at the NIH, among others. Structure determination by NMR spectroscopy usually consists of several phases, each using a separate set of highly specialized techniques. The sample is prepared, measurements are made, interpretive approaches are applied, and a structure is calculated and validated.NMR involves the quantum mechanical properties of the central core (""nucleus"") of the atom. These properties depend on the local molecular environment, and their measurement provides a map of how the atoms are linked chemically, how close they are in space, and how rapidly they move with respect to each other. These properties are fundamentally the same as those used in the more familiar Magnetic Resonance Imaging (MRI), but the molecular applications use a somewhat different approach, appropriate to the change of scale from millimeters (of interest to radiologists) to nano-meters (bonded atoms are typically a fraction of a nano-meter apart), a factor of a million. This change of scale requires much higher sensitivity of detection and stability for long term measurement. In contrast to MRI, structural biology studies do not directly generate an image, but rely on complex computer calculations to generate three-dimensional molecular models.Currently most samples are examined in a solution in water, but methods are being developed to also work with solid samples. Data collection relies on placing the sample inside a powerful magnet, sending radio frequency signals through the sample, and measuring the absorption of those signals. Depending on the environment of atoms within the protein, the nuclei of individual atoms will absorb different frequencies of radio signals. Furthermore the absorption signals of different nuclei may be perturbed by adjacent nuclei. This information can be used to determine the distance between nuclei. These distances in turn can be used to determine the overall structure of the protein.A typical study might involve how two proteins interact with each other, possibly with a view to developing small molecules that can be used to probe the normal biology of the interaction (""chemical biology"") or to provide possible leads for pharmaceutical use (drug development). Frequently, the interacting pair of proteins may have been identified by studies of human genetics, indicating the interaction can be disrupted by unfavorable mutations, or they may play a key role in the normal biology of a ""model"" organism like the fruit fly, yeast, the worm C. elegans, or mice. To prepare a sample, methods of molecular biology are typically used to make quantities by bacterial fermentation. This also permits changing the isotopic composition of the molecule, which is desirable because the isotopes behave differently and provide methods for identifying overlapping NMR signals.