AASK Additional Activities
... (native structure) following their complete unfolding (denaturation) by heating. 1. Have each student document the native shape of their folded protein with a digital photo. 2. Ask the students to unfold their protein and then re-fold it. 3. Check the refolded protein against the photo of the native ...
... (native structure) following their complete unfolding (denaturation) by heating. 1. Have each student document the native shape of their folded protein with a digital photo. 2. Ask the students to unfold their protein and then re-fold it. 3. Check the refolded protein against the photo of the native ...
here - BioGeometry
... fanfare last month – researchers now face an even more daunting task of figuring out how the 30,000 or so genes give rise to the biological protein machinery that makes humans uniquely humans. A central problem in this field, called “proteomics,” is how to mathematically describe the intricate foldi ...
... fanfare last month – researchers now face an even more daunting task of figuring out how the 30,000 or so genes give rise to the biological protein machinery that makes humans uniquely humans. A central problem in this field, called “proteomics,” is how to mathematically describe the intricate foldi ...
Symmetry
... Horse liver alcohol dehydrogenase exists as a homodimer or heterodimerthere are two different types of monomer, denoted E and S. However these two forms are near-identical (both consisting of 374 amino acids), as there are only six residues which are different. None of these six occur in the inter ...
... Horse liver alcohol dehydrogenase exists as a homodimer or heterodimerthere are two different types of monomer, denoted E and S. However these two forms are near-identical (both consisting of 374 amino acids), as there are only six residues which are different. None of these six occur in the inter ...
Prob_Set_2_2007
... - due Wednesday Feb 21 in class 1) Pick out a protein of known structure that is central to your research or is connected to your research interests. Download the coordinates from the Protein Data Bank (www.rcsb.org) and use a rendering program such as Chimera, Rasmol, Pymol, Molmol, iMol (Macs) etc ...
... - due Wednesday Feb 21 in class 1) Pick out a protein of known structure that is central to your research or is connected to your research interests. Download the coordinates from the Protein Data Bank (www.rcsb.org) and use a rendering program such as Chimera, Rasmol, Pymol, Molmol, iMol (Macs) etc ...
Table S9.
... This domain, found in various prokaryotic proteins, has no known function. This family consists of several proteins of uncharacterised function. This family of proteins with unknown function appear to be restricted to Cyanobacteria. This family of proteins with unknown function appears to be restric ...
... This domain, found in various prokaryotic proteins, has no known function. This family consists of several proteins of uncharacterised function. This family of proteins with unknown function appear to be restricted to Cyanobacteria. This family of proteins with unknown function appears to be restric ...
more details
... contrary to standard models, that the amino acid propensities at a given location vary widely due to these epistatic interactions, with the changes in propensities occurring at a wide range of timescales. We also observe that, following a substitution at a given location, the rest of the protein und ...
... contrary to standard models, that the amino acid propensities at a given location vary widely due to these epistatic interactions, with the changes in propensities occurring at a wide range of timescales. We also observe that, following a substitution at a given location, the rest of the protein und ...
Supplementary data 1,2,3,4,6,7,8,9 include N, Total (ProtScore)
... Supplementary data 1,2,3,4,6,7,8,9 include N, Total (ProtScore), %Cov(95), Accession, Name, Conf, Sequence,Modifications for the identified proteins. The definitions of the table fields are described as follows: N is the rank of the specified protein relative to all other proteins in the list of det ...
... Supplementary data 1,2,3,4,6,7,8,9 include N, Total (ProtScore), %Cov(95), Accession, Name, Conf, Sequence,Modifications for the identified proteins. The definitions of the table fields are described as follows: N is the rank of the specified protein relative to all other proteins in the list of det ...
A Novel Scoring Function for Predicting the Conformation of Pairs of
... Many pairs of helices in transmembrane (TM) proteins are tightly packed. We present a scoring function and a computational methodology for predicting the tertiary fold of a pair of α-helices, such that its chances of being tightly packed are maximized. Since the number of TM protein structures solve ...
... Many pairs of helices in transmembrane (TM) proteins are tightly packed. We present a scoring function and a computational methodology for predicting the tertiary fold of a pair of α-helices, such that its chances of being tightly packed are maximized. Since the number of TM protein structures solve ...
20 Proteins - mrhortonbiology
... Drew has been buying protein shakes from his gym to gain muscle, but they do not have nutrition facts. How could he determine whether they actually have much protein in them? (Be specific about what he would do, what he would measure, etc) ...
... Drew has been buying protein shakes from his gym to gain muscle, but they do not have nutrition facts. How could he determine whether they actually have much protein in them? (Be specific about what he would do, what he would measure, etc) ...
tutorial10_3D_structure
... •Class: The secondary structure composition: mainlyalpha, mainly-beta and alpha-beta. • Architecture: The overall shape of the domain structure. Orientations of the secondary structures : e.g. barrel or 3layer sandwich. • Topology: Structures are grouped into fold groups at this level depending on b ...
... •Class: The secondary structure composition: mainlyalpha, mainly-beta and alpha-beta. • Architecture: The overall shape of the domain structure. Orientations of the secondary structures : e.g. barrel or 3layer sandwich. • Topology: Structures are grouped into fold groups at this level depending on b ...
ECS 189K - UC Davis
... does that string form a knot? (i.e. if you were to hold the two extremities of the string and pull, would it result in the formation of a knot, or would the string become linear?) Some proteins do form knots, and it remains unclear at this time if these knots play a role in defining the functions of ...
... does that string form a knot? (i.e. if you were to hold the two extremities of the string and pull, would it result in the formation of a knot, or would the string become linear?) Some proteins do form knots, and it remains unclear at this time if these knots play a role in defining the functions of ...
biochemistry/docs/Protein structure 1
... exclusively by hydrogen bonds between peptide bond elements. Supersecondary structure- Recurring structural feature of proteins composed of two or more secondary structural elements. Domain- A segment of protein structure that is autonomously stable. Tertiary structure- A stable, independent protein ...
... exclusively by hydrogen bonds between peptide bond elements. Supersecondary structure- Recurring structural feature of proteins composed of two or more secondary structural elements. Domain- A segment of protein structure that is autonomously stable. Tertiary structure- A stable, independent protein ...
Organic chemistry and Biological chemistry for Health Sciences
... The heme molecule is held in the folded globin molecule by electrical attraction between two electrically charged side chains and the Fe2+ ion in heme. For many proteins native form only emerges only as two or more polypeptides assemble into a quaternary structure. Individual molecules of polypeptid ...
... The heme molecule is held in the folded globin molecule by electrical attraction between two electrically charged side chains and the Fe2+ ion in heme. For many proteins native form only emerges only as two or more polypeptides assemble into a quaternary structure. Individual molecules of polypeptid ...
... RESULTS: Scores of the four domains (physical, psychological, social relationships and environment) were very similar. There were statistically signifi cant differences in mean scores for the environment domain according to skin color, with blacks and pardos having lower scores. Women also had the l ...
Protein Aggregation in High-Protein Caramel
... Here, the caramel takes on a tapioca-like structure, with large visible aggregates of protein structures (Figure 1), as it loses its desirable smooth texture. There are two general categories of proteins in milk — the caseins (≈80%) and the serum proteins (≈20%). The various casein proteins form int ...
... Here, the caramel takes on a tapioca-like structure, with large visible aggregates of protein structures (Figure 1), as it loses its desirable smooth texture. There are two general categories of proteins in milk — the caseins (≈80%) and the serum proteins (≈20%). The various casein proteins form int ...
Gene Section STARD13 (star-related lipid transfer (START) domain containing 13)
... deleted in HCC. DLC2 is flanked by microsatellite markers D13S171 and D13S267. Loss of heterozygosity on these two markers is frequently found in HCC. Allelic losses at markers D13S171 and D13S267 are detected in 33.3% and 40.7% of the informative cases, respectively. RT-PCR analysis of DLC2 mRNA in ...
... deleted in HCC. DLC2 is flanked by microsatellite markers D13S171 and D13S267. Loss of heterozygosity on these two markers is frequently found in HCC. Allelic losses at markers D13S171 and D13S267 are detected in 33.3% and 40.7% of the informative cases, respectively. RT-PCR analysis of DLC2 mRNA in ...
Name: Date: Kingdoms and Domains – Section 15.4 Worksheet The
... The Tree of Life Evolves (pages 325) 1. Is the following sentence true or false? The scientific view of life was more complex in Linnaeus’s time. _____________________ 2. What fundamental traits did Linnaeus use to separate plants from animals? _____________________ _________________________________ ...
... The Tree of Life Evolves (pages 325) 1. Is the following sentence true or false? The scientific view of life was more complex in Linnaeus’s time. _____________________ 2. What fundamental traits did Linnaeus use to separate plants from animals? _____________________ _________________________________ ...
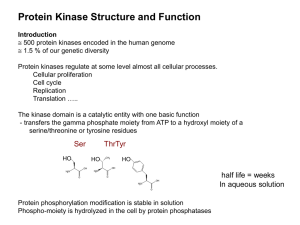
Protein domain

A protein domain is a conserved part of a given protein sequence and (tertiary) structure that can evolve, function, and exist independently of the rest of the protein chain. Each domain forms a compact three-dimensional structure and often can be independently stable and folded. Many proteins consist of several structural domains. One domain may appear in a variety of different proteins. Molecular evolution uses domains as building blocks and these may be recombined in different arrangements to create proteins with different functions. Domains vary in length from between about 25 amino acids up to 500 amino acids in length. The shortest domains such as zinc fingers are stabilized by metal ions or disulfide bridges. Domains often form functional units, such as the calcium-binding EF hand domain of calmodulin. Because they are independently stable, domains can be ""swapped"" by genetic engineering between one protein and another to make chimeric proteins.