Supplemental Table 3
... [evidence IEA]; Catalysis of the hydrolysis of peptide bonds by a mechanism in which water acts as a nucleophile, one or two metal ions hold the water molecule in place, and charged amino acid side chains are ligands for the metal ions [goid 8237] [evidence IEA]; Interacting selectively with zinc (Z ...
... [evidence IEA]; Catalysis of the hydrolysis of peptide bonds by a mechanism in which water acts as a nucleophile, one or two metal ions hold the water molecule in place, and charged amino acid side chains are ligands for the metal ions [goid 8237] [evidence IEA]; Interacting selectively with zinc (Z ...
Perspectives in Diabetes Glucokinase Gene Structure
... insulin-producing cells. To clarify future discussion of these 5'-end). different glucokinase isoforms, herein, the liver glucokinase It is not known whether these three glucokinase isoforms isoform will be referred to as glucokinase L1, and the two have the same or different enzymatic properties. A ...
... insulin-producing cells. To clarify future discussion of these 5'-end). different glucokinase isoforms, herein, the liver glucokinase It is not known whether these three glucokinase isoforms isoform will be referred to as glucokinase L1, and the two have the same or different enzymatic properties. A ...
... abundance, the total cellular RNA levels increased eightfold (Figure 2C), suggesting that other enhancing mechanisms are involved in achieving the final RNA levels. One possible explanation for escape of silencing by the paternal Kcnq1 allele is that expression is initiated from an alternative start ...
Document
... After diffusing inside, where the H+ concentration low, it gives up the proton. So it ferries protons from regions of high concentration to regions of low concentration, thus destroying the proton gradient. Electron transport chain goes merrily on and on, but no gradient is formed and no ATP is prod ...
... After diffusing inside, where the H+ concentration low, it gives up the proton. So it ferries protons from regions of high concentration to regions of low concentration, thus destroying the proton gradient. Electron transport chain goes merrily on and on, but no gradient is formed and no ATP is prod ...
Functional characterization of polypeptide release factor 1b in the
... linear amino acids, Pro-Ala-Thr (PAT), whereas RF2 recognizes UAA/UGA through Ser-Pro-Phe (SPF) [4,5]. In higher eukaryotes, there is only one RF-I, eRF1 (eukaryotic release factor 1), which can decode all three stop codons [6–8]. ...
... linear amino acids, Pro-Ala-Thr (PAT), whereas RF2 recognizes UAA/UGA through Ser-Pro-Phe (SPF) [4,5]. In higher eukaryotes, there is only one RF-I, eRF1 (eukaryotic release factor 1), which can decode all three stop codons [6–8]. ...
Understanding The Function And Regulation Of Eukaryotic Release
... Nascent polypeptides are synthesized from a messenger RNA (mRNA) template through a process called translation. Eukaryotic translation is composed of four main steps: initiation, elongation, termination and recycling [7]. Through this process essentially all necessary peptides and proteins are gener ...
... Nascent polypeptides are synthesized from a messenger RNA (mRNA) template through a process called translation. Eukaryotic translation is composed of four main steps: initiation, elongation, termination and recycling [7]. Through this process essentially all necessary peptides and proteins are gener ...
Introduction to the BLAST Suite and BLASTN
... unique identifiers. Accession numbers are unique to a sequence record, but should that record be revised, then the accession number is given a new version number. When this sequence was submitted to GenBank, it was given an accession number of DD148865, and the version was DD148865.1. If this record ...
... unique identifiers. Accession numbers are unique to a sequence record, but should that record be revised, then the accession number is given a new version number. When this sequence was submitted to GenBank, it was given an accession number of DD148865, and the version was DD148865.1. If this record ...
The PRT protein family Sangita C Sinha* and Janet L Smith
... between layers of α helices, α1 and α2 on one face of the β sheet and α3 on the other face (Figure 1). The core fold is expanded by between two and five additional secondary structures, which vary among PRT family members. At the C-terminal edge of the core β sheet, a second domain or subdomain, kno ...
... between layers of α helices, α1 and α2 on one face of the β sheet and α3 on the other face (Figure 1). The core fold is expanded by between two and five additional secondary structures, which vary among PRT family members. At the C-terminal edge of the core β sheet, a second domain or subdomain, kno ...
Stimulation Do Not Alter TTP Function Protein Kinase and
... received the following: 1 g of pGL3 luciferase vector and varying concentrations of the specified pcDNA 3.1 His C-TTP vector, with the difference in DNA transfected made up with the pcDNA3.1 His-C parental for a total of 1 g in 100 l of total, serum free RPMI 1640 medium. The DNA mixture was mixe ...
... received the following: 1 g of pGL3 luciferase vector and varying concentrations of the specified pcDNA 3.1 His C-TTP vector, with the difference in DNA transfected made up with the pcDNA3.1 His-C parental for a total of 1 g in 100 l of total, serum free RPMI 1640 medium. The DNA mixture was mixe ...
Beverage Treatment Products Technical Data Sheet Pure
... Safety For the prior addition of SIHA Fermentation Salt yeast nutrient fermentation aid and/or SIHA Vitamin B 1 yeast nutrient fermentation aid, the legal limit values must also be adhered to. When used and handled correctly, there are no known unfavorable effects associated with SIHA PROFERM Plus c ...
... Safety For the prior addition of SIHA Fermentation Salt yeast nutrient fermentation aid and/or SIHA Vitamin B 1 yeast nutrient fermentation aid, the legal limit values must also be adhered to. When used and handled correctly, there are no known unfavorable effects associated with SIHA PROFERM Plus c ...
ppt for
... Population dynamics of piRNA and TE(transposable element) in Drosophila • Jian Lu and Andrew G. Clark, PNAS 2010 • Activities of a large number of retrotransposons are severely silenced by piRNAs. 1) quantified expression levels of 32 TE families 2) in ovaries of one wild-type and three piRNA ...
... Population dynamics of piRNA and TE(transposable element) in Drosophila • Jian Lu and Andrew G. Clark, PNAS 2010 • Activities of a large number of retrotransposons are severely silenced by piRNAs. 1) quantified expression levels of 32 TE families 2) in ovaries of one wild-type and three piRNA ...
Formate Dehydrogenase, Molecular Modeling and Docking with
... interaction toward NAD+ with grid score of -60.23, which is considered as an appropriate score for binding. After surfacing the 3D-structure of enzyme regions interacting with NAD+, we observed that FDH forms a pocket interaction shape with NAD+. The shape of FDH active site comprised with Pro 68, A ...
... interaction toward NAD+ with grid score of -60.23, which is considered as an appropriate score for binding. After surfacing the 3D-structure of enzyme regions interacting with NAD+, we observed that FDH forms a pocket interaction shape with NAD+. The shape of FDH active site comprised with Pro 68, A ...
Structure and function of haemoglobin: II. Some
... Residues are defined here as invariant when they occur at structurally identical sites in all the normal myoglobins and haemoglobins so far investigated. Abnormal haemoglobins have been excluded, because some of their abnormalities interfere with the oxygen-combining function, so that the protein ca ...
... Residues are defined here as invariant when they occur at structurally identical sites in all the normal myoglobins and haemoglobins so far investigated. Abnormal haemoglobins have been excluded, because some of their abnormalities interfere with the oxygen-combining function, so that the protein ca ...
Cardiovascular proteomics
... the posttranslational modifications “That is soon going to change…. Proteomics is going to have to have the functional verification, with time.” Van Eyk ...
... the posttranslational modifications “That is soon going to change…. Proteomics is going to have to have the functional verification, with time.” Van Eyk ...
Metal ion reconstitution studies of yeast copper
... tyrosine residue, and one disulfide bond per subunit, resulting in a calculated ε280 of 3230 M –1 cm –1/dimer. Bovine apo CuZnSOD has identical numbers of tryptophan, tyrosine, and cystine residues [7], and therefore it is expected to have the same extinction coefficient at 280 nm, i.e., 3230 M –1 c ...
... tyrosine residue, and one disulfide bond per subunit, resulting in a calculated ε280 of 3230 M –1 cm –1/dimer. Bovine apo CuZnSOD has identical numbers of tryptophan, tyrosine, and cystine residues [7], and therefore it is expected to have the same extinction coefficient at 280 nm, i.e., 3230 M –1 c ...
Invariant amino acids essential for decoding function of polypeptide
... As follows from the structural analysis (Figure 3A) positions 55 and 125 are close in space and can form a hydrogen bond between the hydrogen atom of OH group of Tyr125 and the oxygen atom of the COO group of Glu55. The length of the bond is equal to 1.73 s (Figure 3B). This suggestion is consistent ...
... As follows from the structural analysis (Figure 3A) positions 55 and 125 are close in space and can form a hydrogen bond between the hydrogen atom of OH group of Tyr125 and the oxygen atom of the COO group of Glu55. The length of the bond is equal to 1.73 s (Figure 3B). This suggestion is consistent ...
mitochondrial biogenesis during
... The incorporation system was similar to that used by Reid and Parsons (1971) but omitting maleate and pyruvate . The concentration of [3 H]UTP was 0.2 pM with a specific radioactivity of 14.8 Ci/mmol . Slightly better results were obtained when the assay was conducted in the presence of 0 .25 M sucr ...
... The incorporation system was similar to that used by Reid and Parsons (1971) but omitting maleate and pyruvate . The concentration of [3 H]UTP was 0.2 pM with a specific radioactivity of 14.8 Ci/mmol . Slightly better results were obtained when the assay was conducted in the presence of 0 .25 M sucr ...
Xenobiotic-metabolizing cytochrome P450 enzymes in human lung
... xenobiotics, including most of the therapeutic drugs and environmental pollutants (Nelson et al. 1996, Bertz & Granneman 1997). The first report on the existence of a CYP enzyme or a “microsomal carbon monoxide-binding pigment”, as it was called at that time, was published in 1958 by Klingenberg et ...
... xenobiotics, including most of the therapeutic drugs and environmental pollutants (Nelson et al. 1996, Bertz & Granneman 1997). The first report on the existence of a CYP enzyme or a “microsomal carbon monoxide-binding pigment”, as it was called at that time, was published in 1958 by Klingenberg et ...
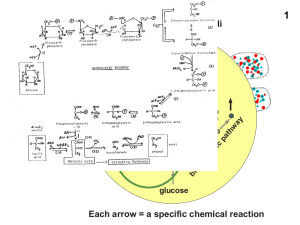
Pathways of genetic code evolution in ancient and modern organisms
... group does not imply that all species in the group are reassigned. Multiple reassignments of the same codon may occur independently within a group ...
... group does not imply that all species in the group are reassigned. Multiple reassignments of the same codon may occur independently within a group ...
Structural elements defining elongation factor Tu mediated
... and Glu-tRNAGln are unable to bind in vitro elongation factor Tu (EF-Tu), which delivers aa-tRNAs to the ribosome A-site during peptide elongation (5,6). Nevertheless, poor binding of the mischarged AsptRNAAsn and Glu-ARNtGln was reported in vivo when the mischarged aaRS is overexpressed (7,8). It i ...
... and Glu-tRNAGln are unable to bind in vitro elongation factor Tu (EF-Tu), which delivers aa-tRNAs to the ribosome A-site during peptide elongation (5,6). Nevertheless, poor binding of the mischarged AsptRNAAsn and Glu-ARNtGln was reported in vivo when the mischarged aaRS is overexpressed (7,8). It i ...
Evolving genetic code
... (by reading the mRNA) the amino acid sequences of so many highly evolved protein molecules that any change to these would be highly disadvantageous unless to correct the ‘‘mistakes’’ produced by altering the code. This accounts for the fact that the code does not change. To account for it being the ...
... (by reading the mRNA) the amino acid sequences of so many highly evolved protein molecules that any change to these would be highly disadvantageous unless to correct the ‘‘mistakes’’ produced by altering the code. This accounts for the fact that the code does not change. To account for it being the ...
eg1
... multiple forms (Knowles et al. 1987, Kubicek 1992). V. volvacea produces a cellulolytic system that includes multiple forms of all three classes when grown on crystalline cellulose (Cai et al. 1994, 1999). In addition to EG1, four other CMC-hydrolysing proteins were separated in lower yields from cu ...
... multiple forms (Knowles et al. 1987, Kubicek 1992). V. volvacea produces a cellulolytic system that includes multiple forms of all three classes when grown on crystalline cellulose (Cai et al. 1994, 1999). In addition to EG1, four other CMC-hydrolysing proteins were separated in lower yields from cu ...
Evolving genetic code - J
... (by reading the mRNA) the amino acid sequences of so many highly evolved protein molecules that any change to these would be highly disadvantageous unless to correct the ‘‘mistakes’’ produced by altering the code. This accounts for the fact that the code does not change. To account for it being the ...
... (by reading the mRNA) the amino acid sequences of so many highly evolved protein molecules that any change to these would be highly disadvantageous unless to correct the ‘‘mistakes’’ produced by altering the code. This accounts for the fact that the code does not change. To account for it being the ...