DNA - VanityWolveriine
... both parents. The offspring is a perfect combination of each parent. The dad is fat, the mom is skinny, the offspring is normal. The dad is brown and white, the mom is white so the offspring is a light brown. ...
... both parents. The offspring is a perfect combination of each parent. The dad is fat, the mom is skinny, the offspring is normal. The dad is brown and white, the mom is white so the offspring is a light brown. ...
DNA: Sample Storage - Sacramento County District Attorney
... Amplified DNA from casework will be retained in frozen storage until the case has been technically and administratively reviewed. After the review process has been completed, the amplified DNA may be destroyed. NOTE: Exceptions to this process are when ...
... Amplified DNA from casework will be retained in frozen storage until the case has been technically and administratively reviewed. After the review process has been completed, the amplified DNA may be destroyed. NOTE: Exceptions to this process are when ...
Researchers ACT on DNA Storage
... Shakespeare's sonnets and an audio recording of Martin Luther King. The work is published in the journal Nature. [Nick Goldman et al, Towards practical, high-capacity, low-maintenance information storage in synthesized DNA] Unlike many forms of information storage, DNA is extremely long-lasting and ...
... Shakespeare's sonnets and an audio recording of Martin Luther King. The work is published in the journal Nature. [Nick Goldman et al, Towards practical, high-capacity, low-maintenance information storage in synthesized DNA] Unlike many forms of information storage, DNA is extremely long-lasting and ...
Whole genome shotgun sequencing
... 1. Select random BAC clones and sequence ends of inserts. Each of these sequences = Sequence-Tagged Site (STS). 2. Design PCR primers from within these STS’s & test other BAC clones for inclusion of this DNA sequence by PCR. ...
... 1. Select random BAC clones and sequence ends of inserts. Each of these sequences = Sequence-Tagged Site (STS). 2. Design PCR primers from within these STS’s & test other BAC clones for inclusion of this DNA sequence by PCR. ...
Genomics wordsearch
... nucleotides in a DNA/RNA molecule which codes for an amino acid Cytosine – A nucleotide component of DNA/RNA ...
... nucleotides in a DNA/RNA molecule which codes for an amino acid Cytosine – A nucleotide component of DNA/RNA ...
2.2 Sequencing learning grid File
... 2.2.8 Studying whole genomes When was the structure of DNA discovered? Understanding and manipulating DNA ...
... 2.2.8 Studying whole genomes When was the structure of DNA discovered? Understanding and manipulating DNA ...
CALL FOR PROPOSALS 2008
... axeny, specific information on genome size (bibliographic references or techniques for estimation of size), G+C content, information on ploidy, polymorphism level (details and methods of estimation), repeat structure with details about how these are known, etc. ...
... axeny, specific information on genome size (bibliographic references or techniques for estimation of size), G+C content, information on ploidy, polymorphism level (details and methods of estimation), repeat structure with details about how these are known, etc. ...
BIOCHEMISTRY 4.1 HOMEWORK
... b. Draw the structures resulting from the reaction of this end sequence DNA polymerase I and four deoxynucloside triphosphases (see Fig 8-36 in the book!) ...
... b. Draw the structures resulting from the reaction of this end sequence DNA polymerase I and four deoxynucloside triphosphases (see Fig 8-36 in the book!) ...
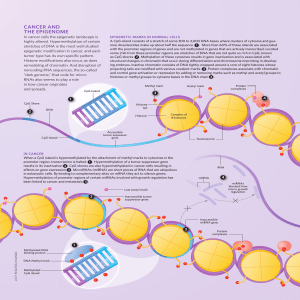
CaNCer aND THe ePIGeNOMe
... stretches of DNA is the most well-studied epigenetic modification in cancer, and each tumor type has its own specific pattern. Histone modifications also occur, as does remodeling of chromatin. And disruption of noncoding RNA sequences, the so-called “dark genome,” that code for microRNAs also seems ...
... stretches of DNA is the most well-studied epigenetic modification in cancer, and each tumor type has its own specific pattern. Histone modifications also occur, as does remodeling of chromatin. And disruption of noncoding RNA sequences, the so-called “dark genome,” that code for microRNAs also seems ...
DNA_LAdders_files/StoS 100bp DNA Ladder flyer new
... A unique combination of PCR products and a number of proprietary plasmids digested with appropriate restriction enzymes to yield 11 fragments suitable for use as molecular weight standards for agarose gel electrophoresis. The DNA includes fragments ranging from 100-1,500 bp. The 500 and 1,500 bp ban ...
... A unique combination of PCR products and a number of proprietary plasmids digested with appropriate restriction enzymes to yield 11 fragments suitable for use as molecular weight standards for agarose gel electrophoresis. The DNA includes fragments ranging from 100-1,500 bp. The 500 and 1,500 bp ban ...
a10c Biotechnology
... 5. What is DNA fingerprinting? What molecular and laboratory tools are used to produce a "fingerprint"? What is the basis of concluding that different individual organisms should have different fingerprints? Is this always true? 6. What is the polymerase chain reaction (PCR)? How has this process be ...
... 5. What is DNA fingerprinting? What molecular and laboratory tools are used to produce a "fingerprint"? What is the basis of concluding that different individual organisms should have different fingerprints? Is this always true? 6. What is the polymerase chain reaction (PCR)? How has this process be ...
DNA Test Review
... 3. If a DNA molecule has the sequence TACGAACCC, what would be the complimentary mRNA sequence? 4. The process by which a DNA molecule is copied is called _____. 5. What is a codon? 6. What are the types of RNA? 7. Messenger RNA is formed in the process of _____. 8. What happens during translation a ...
... 3. If a DNA molecule has the sequence TACGAACCC, what would be the complimentary mRNA sequence? 4. The process by which a DNA molecule is copied is called _____. 5. What is a codon? 6. What are the types of RNA? 7. Messenger RNA is formed in the process of _____. 8. What happens during translation a ...
What are the three steps in PCR?
... It is often used in DNA fingerprinting It requires gel electrophoresis which separates DNA by size ...
... It is often used in DNA fingerprinting It requires gel electrophoresis which separates DNA by size ...
High resolution melting for methylation analysis
... • Visceromegaly (Abnormal enlargement of the large internal organs) • Embryonal tumours (e.g. Wilms tumour) • Hemihyperplasia (asymmetric overgrowth of region(s) of the body) • Neonatal hypoglycaemia • Ear creases/pits ...
... • Visceromegaly (Abnormal enlargement of the large internal organs) • Embryonal tumours (e.g. Wilms tumour) • Hemihyperplasia (asymmetric overgrowth of region(s) of the body) • Neonatal hypoglycaemia • Ear creases/pits ...
Chapter 11 Concept Check Questions
... 1. How did Griffith’s experiments indicate the presence of a “transforming factor” in bacteria? ...
... 1. How did Griffith’s experiments indicate the presence of a “transforming factor” in bacteria? ...
Using microsatellites as molecular markers
... Use PCR primers that are complementary to single copy sequences flanking microsatellites to amplify microsatellite-containing region. Depending on number of microsatellite repeats, will get different lengths PCR products (many different possible alleles, not just two) ...
... Use PCR primers that are complementary to single copy sequences flanking microsatellites to amplify microsatellite-containing region. Depending on number of microsatellite repeats, will get different lengths PCR products (many different possible alleles, not just two) ...
Introduction
... Plasma was separated from the blood cells by centrifugation at 1500 g for 10 minutes. The supernatant was then transferred to fresh tubes ensuring that the buffy coat remained intact. The plasma was then centrifuged at 16000 g for 10 minutes to remove any remaining cells, transferred into 2 ml Lo-Bi ...
... Plasma was separated from the blood cells by centrifugation at 1500 g for 10 minutes. The supernatant was then transferred to fresh tubes ensuring that the buffy coat remained intact. The plasma was then centrifuged at 16000 g for 10 minutes to remove any remaining cells, transferred into 2 ml Lo-Bi ...
2.5.4. DNA Revision Qs
... (b) the production of an enzyme _____________________________________ (c) the ability to play a musical instrument _____________________________________ (d) the ability to form a blood clot _____________________________________ (e) the ability to read _____________________________________ ...
... (b) the production of an enzyme _____________________________________ (c) the ability to play a musical instrument _____________________________________ (d) the ability to form a blood clot _____________________________________ (e) the ability to read _____________________________________ ...
GENETIC ANALYZER We have a 3130xl Genetic Analyzer from
... The applied biosystems 3130xl is capable of performing sequencing and fragment analysis of applications like microsatellite or short Tandem Repeats (STR), AFLP, LOH, SNP, rapid sequencing, standard sequencing de novo sequencing and resequencing. The sequencer is typically set up for rapid sequencing ...
... The applied biosystems 3130xl is capable of performing sequencing and fragment analysis of applications like microsatellite or short Tandem Repeats (STR), AFLP, LOH, SNP, rapid sequencing, standard sequencing de novo sequencing and resequencing. The sequencer is typically set up for rapid sequencing ...
Genomics
... • Steps in Microarray Analysis • Arrange DNA or cDNA in an ordered array on a glass slide • Incubate array with fluorescently-labeled probe • Detect probe hybridization using a laser • Record and store data with computers ...
... • Steps in Microarray Analysis • Arrange DNA or cDNA in an ordered array on a glass slide • Incubate array with fluorescently-labeled probe • Detect probe hybridization using a laser • Record and store data with computers ...
BACKGROUND: UvrC is a DNA repair enzyme found in all
... C. Generate a table like the one below for each combination. D. Build a cladogram (phylogenetic tree) based on the data. QUESTIONS: 1. The Data Table (for all combinations) Organism Accession % Identity Number ...
... C. Generate a table like the one below for each combination. D. Build a cladogram (phylogenetic tree) based on the data. QUESTIONS: 1. The Data Table (for all combinations) Organism Accession % Identity Number ...
Bisulfite sequencing

Bisulphite sequencing (also known as bisulfite sequencing) is the use of bisulphite treatment of DNA to determine its pattern of methylation. DNA methylation was the first discovered epigenetic mark, and remains the most studied. In animals it predominantly involves the addition of a methyl group to the carbon-5 position of cytosine residues of the dinucleotide CpG, and is implicated in repression of transcriptional activity.Treatment of DNA with bisulphite converts cytosine residues to uracil, but leaves 5-methylcytosine residues unaffected. Thus, bisulphite treatment introduces specific changes in the DNA sequence that depend on the methylation status of individual cytosine residues, yielding single- nucleotide resolution information about the methylation status of a segment of DNA. Various analyses can be performed on the altered sequence to retrieve this information. The objective of this analysis is therefore reduced to differentiating between single nucleotide polymorphisms (cytosines and thymidine) resulting from bisulphite conversion (Figure 1).