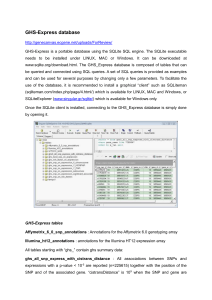
GHS-Express database http://genecanvas.ecgene.net/uploads/Fo
... of base pairs separating the SNP from the gene sequence when the SNP and gene are located on the same chromosome, or -999 when the information is lacking. ghs_best_snp_on_expression : This is a convenience table in which the best SNP for each gene was selected. ghs_risk_factors_on_expression: All as ...
... of base pairs separating the SNP from the gene sequence when the SNP and gene are located on the same chromosome, or -999 when the information is lacking. ghs_best_snp_on_expression : This is a convenience table in which the best SNP for each gene was selected. ghs_risk_factors_on_expression: All as ...
slides
... Most SNPs are outside of the protein coding regions 1 SNP every 600 base pairs More than 5 million common SNPs each with frequency 10-‐50% account for the bulk of human DNA sequence difference I ...
... Most SNPs are outside of the protein coding regions 1 SNP every 600 base pairs More than 5 million common SNPs each with frequency 10-‐50% account for the bulk of human DNA sequence difference I ...
Impact of genomics on dairy cattle breeding - VT Dairy
... dear for most of my career. By the way, SNP stands for “single nucleotide polymorphism”. A SNP is a site on a chromosome where animals in a population have different nucleic acids. If all animals have the same nucleic acid at a site, then there is no variation at that site to associate with performa ...
... dear for most of my career. By the way, SNP stands for “single nucleotide polymorphism”. A SNP is a site on a chromosome where animals in a population have different nucleic acids. If all animals have the same nucleic acid at a site, then there is no variation at that site to associate with performa ...
No Slide Title
... established genetic markers that aid in the identification of loci affecting quantitative traits and/or disease in wide variety of eukaryote species. In addition, SNPs have been used extensively in efforts to study the evolution of microbial populations. Such efforts have largely been confined to mu ...
... established genetic markers that aid in the identification of loci affecting quantitative traits and/or disease in wide variety of eukaryote species. In addition, SNPs have been used extensively in efforts to study the evolution of microbial populations. Such efforts have largely been confined to mu ...
document
... – If not, what does this suggest about the evolution of the phenotype in these populations (idea of convergent evolution) ...
... – If not, what does this suggest about the evolution of the phenotype in these populations (idea of convergent evolution) ...
doc bio 202 2009
... 16. (1 point) An individual who is heterozygous has brittle bone disease disease (in the mutant heterozygote an abnormal copy of the protein wraps around one or two copies of the other and distorts the conformation of the functional trimeric molecule). This type of mutation is called a: a. a leaky m ...
... 16. (1 point) An individual who is heterozygous has brittle bone disease disease (in the mutant heterozygote an abnormal copy of the protein wraps around one or two copies of the other and distorts the conformation of the functional trimeric molecule). This type of mutation is called a: a. a leaky m ...
Which best describes the genetics of the afflicting allele in the
... genotypes are known? (i.e., indicate the genotypes on the figure for all known AA, Aa, and aa individuals) 3. Given the following pedigree, would you expect to find more of in Cleopatra-Berenike III compared with the general population? a. Loci which are heterozygous b. Loci which are homozygous for ...
... genotypes are known? (i.e., indicate the genotypes on the figure for all known AA, Aa, and aa individuals) 3. Given the following pedigree, would you expect to find more of in Cleopatra-Berenike III compared with the general population? a. Loci which are heterozygous b. Loci which are homozygous for ...
Rearrangements in the Human T-Cell-Receptor Â
... we were also able to demonstrate rearrangements in the J« Since the TCR aßlocus is the region where several types of chromosome translocations occur in T-cell tumors, rearrangements of the TCR aß locus in DNA from 5 ATL patients. ABSTRACT ...
... we were also able to demonstrate rearrangements in the J« Since the TCR aßlocus is the region where several types of chromosome translocations occur in T-cell tumors, rearrangements of the TCR aß locus in DNA from 5 ATL patients. ABSTRACT ...
Detection of complex mutations in Swedish FAP familes
... The GeneChip 6.0 platform consists of about 906 600 SNP sequences and about 900 000 nonpolymorphic pobes, which cover the whole genome with an average spacing of 0.7Kb. The exon-arrays include over 40 probes for each gene and four probes (one probeset) for every exon for all well annotated genes. Th ...
... The GeneChip 6.0 platform consists of about 906 600 SNP sequences and about 900 000 nonpolymorphic pobes, which cover the whole genome with an average spacing of 0.7Kb. The exon-arrays include over 40 probes for each gene and four probes (one probeset) for every exon for all well annotated genes. Th ...
deschamp_2009_sequencing
... • Small genomes that are not too complex (repeats, duplications...) • The longer the reads, the better – Targeted Resequencing • Complex genomes (crops) – Reduced representation libraries (methyl-sensitive enzymes) ...
... • Small genomes that are not too complex (repeats, duplications...) • The longer the reads, the better – Targeted Resequencing • Complex genomes (crops) – Reduced representation libraries (methyl-sensitive enzymes) ...
Human Identity Testing
... that carries a “lightbulb.” The lightbulb is an analogy for a radioactive label or fluorescent dye that allows it to be visible. The probe is allowed to bind (aka hybridize) with its complementary section in the medium. Then special procedures are used to wash away any remaining single stranded prob ...
... that carries a “lightbulb.” The lightbulb is an analogy for a radioactive label or fluorescent dye that allows it to be visible. The probe is allowed to bind (aka hybridize) with its complementary section in the medium. Then special procedures are used to wash away any remaining single stranded prob ...
MAPPING GENES TO TRAITS IN DOGS USING SNPs
... Part 1: Read and Answer Questions Distribute the student worksheet to students. Ask students to read the NIH press release, Variants in Three Genes Account for Most Dog Coat Differences, and answer the associated questions before class. (Students will return to this article at the end of the activit ...
... Part 1: Read and Answer Questions Distribute the student worksheet to students. Ask students to read the NIH press release, Variants in Three Genes Account for Most Dog Coat Differences, and answer the associated questions before class. (Students will return to this article at the end of the activit ...
Distortion of quantitative genomic and expression
... regarding reproducibility of these techniques have been raised by cross-validation studies in different laboratories (1–5). Strategies to mitigate variability in the results obtained from replicate studies have focused on standardizing technical factors, such as array production, RNA synthesis, labe ...
... regarding reproducibility of these techniques have been raised by cross-validation studies in different laboratories (1–5). Strategies to mitigate variability in the results obtained from replicate studies have focused on standardizing technical factors, such as array production, RNA synthesis, labe ...
emboj2008205-sup
... et al., 2005). Arrays were analyzed using GenePix pro 4.1 (Axon Instruments) and Gene Spring ...
... et al., 2005). Arrays were analyzed using GenePix pro 4.1 (Axon Instruments) and Gene Spring ...
10. Wang T, Liang ZH, Sun SG, Cao XB, Peng H, Liu HJ, et al
... factors for PD through a genome-wide association study and indicated that four SNPs (rs11931532, rs12645693, rs4698412 and rs4538475) showed strong disease association with almost the same significance levels.14 Among them, rs4538475 was the most strongly associated. Another GWAS study in Japanese s ...
... factors for PD through a genome-wide association study and indicated that four SNPs (rs11931532, rs12645693, rs4698412 and rs4538475) showed strong disease association with almost the same significance levels.14 Among them, rs4538475 was the most strongly associated. Another GWAS study in Japanese s ...
Genomic In Situ Hybridization (GISH) as a Tool to Identify
... boiling water for 10 min and labeled with digoxingenin-11-dUTP using the nick translation method (Roche Applied Science, Nutley, NJ, USA). Genomic DNA of HA 89 was used as blocking DNA after shearing, with ratios of blocking DNA to probe DNA ranging from 35:1 to 120:1. Different washing stringencies ...
... boiling water for 10 min and labeled with digoxingenin-11-dUTP using the nick translation method (Roche Applied Science, Nutley, NJ, USA). Genomic DNA of HA 89 was used as blocking DNA after shearing, with ratios of blocking DNA to probe DNA ranging from 35:1 to 120:1. Different washing stringencies ...
Distinguishing Different DNA Heterozygotes by
... These are not identical because such curves are skewed at low temperatures from heteroduplex contributions. In either case, the Tm is only one point on the melting curve. Use of the complete melting curve, conveniently displayed as difference plots, allows differentiation of most heterozygotes (21 o ...
... These are not identical because such curves are skewed at low temperatures from heteroduplex contributions. In either case, the Tm is only one point on the melting curve. Use of the complete melting curve, conveniently displayed as difference plots, allows differentiation of most heterozygotes (21 o ...
Microarrays
... microarrays represent another alternative (e.g., xMAP technology from Luminex [15,16])). Even though, all these elements are essential building blocks of a good MDM, most challenging and most important task is to enable and demonstrate that the developed tool can perform as required in the scope of ...
... microarrays represent another alternative (e.g., xMAP technology from Luminex [15,16])). Even though, all these elements are essential building blocks of a good MDM, most challenging and most important task is to enable and demonstrate that the developed tool can perform as required in the scope of ...
Molecular Inversion Probe

Molecular Inversion Probe (MIP) belongs to the class of Capture by Circularization molecular techniques for performing genomic partitioning, a process through which one captures and enriches specific regions of the genome. Probes used in this technique are single stranded DNA molecules and, similar to other genomic partitioning techniques, contain sequences that are complementary to the target in the genome; these probes hybridize to and capture the genomic target. MIP stands unique from other genomic partitioning strategies in that MIP probes share the common design of two genomic target complementary segments separated by a linker region. With this design, when the probe hybridizes to the target, it undergoes an inversion in configuration (as suggested by the name of the technique) and circularizes. Specifically, the two target complementary regions at the 5’ and 3’ ends of the probe become adjacent to one another while the internal linker region forms a free hanging loop. The technology has been used extensively in the HapMap project for large-scale SNP genotyping as well as for studying gene copy alterationsand characteristics of specific genomic loci to identify biomarkers for different diseases such as cancer. Key strengths of the MIP technology include its high specificity to the target and its scalability for high-throughput, multiplexed analyses where tens of thousands of genomic loci are assayed simultaneously.























