Protein–Nucleic Acid Interactions
... In the idealized A-form of duplex DNA, the major and minor grooves are more equal in width (Figure 10.1a). The major groove of the A-form is comparatively narrower than the B-form, so that the bases of the Aform are more deeply buried within the body of the duplex and are consequently less accessibl ...
... In the idealized A-form of duplex DNA, the major and minor grooves are more equal in width (Figure 10.1a). The major groove of the A-form is comparatively narrower than the B-form, so that the bases of the Aform are more deeply buried within the body of the duplex and are consequently less accessibl ...
Analysis of Guanine Oxidation Products in Double
... guanine-rich sequences can form quadruplex structures. In a previous study using 6-mer DNA d(TGGGGT), which is the shortest oligomer capable of forming quadruplex structures, we demonstrated that guanine oxidation products of quadruplex DNA differ from those of single-stranded DNA. Therefore, the ho ...
... guanine-rich sequences can form quadruplex structures. In a previous study using 6-mer DNA d(TGGGGT), which is the shortest oligomer capable of forming quadruplex structures, we demonstrated that guanine oxidation products of quadruplex DNA differ from those of single-stranded DNA. Therefore, the ho ...
15. nucleic acids
... with gastric juice (we now know that gastric juice contains an enzyme, pepsin, that digests protein). He observed microscopically that only the shrunken nuclei were left after the pepsin treatment; the remainder of the cell had dissolved. These nuclei were then analyzed and found to have different c ...
... with gastric juice (we now know that gastric juice contains an enzyme, pepsin, that digests protein). He observed microscopically that only the shrunken nuclei were left after the pepsin treatment; the remainder of the cell had dissolved. These nuclei were then analyzed and found to have different c ...
3D DNA Crystals and Nanotechnology
... One of the inherent design principles of DNA nanotechnology is that on short length-scales (well below the persistence length), the DNA duplex behaves as a linear, fairly-rigid rod [30]. Constructing branching structures of sufficient rigidity and uniformity to connect these duplexes is the central ...
... One of the inherent design principles of DNA nanotechnology is that on short length-scales (well below the persistence length), the DNA duplex behaves as a linear, fairly-rigid rod [30]. Constructing branching structures of sufficient rigidity and uniformity to connect these duplexes is the central ...
Evidence that MEK1 positively promotes
... poles is dependent upon the interhomologue joints mediated by crossovers and sister chromatid cohesion. Without crossovers or sister chromatid cohesion homologue segregation becomes randomized, causing first meiotic non-disjunction. In some organisms the mechanisms ensuring that sufficient DSBs are rep ...
... poles is dependent upon the interhomologue joints mediated by crossovers and sister chromatid cohesion. Without crossovers or sister chromatid cohesion homologue segregation becomes randomized, causing first meiotic non-disjunction. In some organisms the mechanisms ensuring that sufficient DSBs are rep ...
Non-destructive DNA extraction methods that yield DNA barcodes in
... species, and voucher specimens (Chen et al., 2010). Thus, by using this mechanism, species, old and new alike, could be barcoded efficiently. DNA extraction of Spiders to Gain Barcodes Since there are about 37,000 identified species of spiders and 40,000 suspected species of spiders, barcoding each ...
... species, and voucher specimens (Chen et al., 2010). Thus, by using this mechanism, species, old and new alike, could be barcoded efficiently. DNA extraction of Spiders to Gain Barcodes Since there are about 37,000 identified species of spiders and 40,000 suspected species of spiders, barcoding each ...
Homologous pairing and the role of pairing centers in meiosis
... initially inferred from the effects of reciprocal translocations – chromosome rearrangements involving the exchange of chromosome segments between two non-homologous chromosomes (Fig. 2A) – on the frequency of recombination. In individuals that are heterozygous for such a translocation, recombinatio ...
... initially inferred from the effects of reciprocal translocations – chromosome rearrangements involving the exchange of chromosome segments between two non-homologous chromosomes (Fig. 2A) – on the frequency of recombination. In individuals that are heterozygous for such a translocation, recombinatio ...
Generalized Transduction by Phage P22 in Salmonella typhimurium. II. Mechanisms of Integration of Transducing DNA.
... in this experiment the density of the his+ particles can be calculated from the known densities of the phage (p = 1.506 g/cm3) and the pur+ transducing particles (p = l-511 g/om3). The average density of the his+ transducing particles in this gradient is calculated to be approximately l-512 g/cm3 (a ...
... in this experiment the density of the his+ particles can be calculated from the known densities of the phage (p = 1.506 g/cm3) and the pur+ transducing particles (p = l-511 g/om3). The average density of the his+ transducing particles in this gradient is calculated to be approximately l-512 g/cm3 (a ...
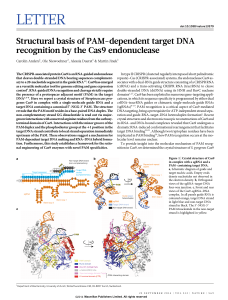
Structural basis of PAM-dependent target DNA recognition by the
... as a versatile molecular tool for genome editing and gene expression control3. RNA-guided DNA recognition and cleavage strictly require the presence of a protospacer adjacent motif (PAM) in the target DNA1,4–6. Here we report a crystal structure of Streptococcus pyogenes Cas9 in complex with a singl ...
... as a versatile molecular tool for genome editing and gene expression control3. RNA-guided DNA recognition and cleavage strictly require the presence of a protospacer adjacent motif (PAM) in the target DNA1,4–6. Here we report a crystal structure of Streptococcus pyogenes Cas9 in complex with a singl ...
AP Bio Chapter 16-20 Practice test
... d. an enzyme that synthesizes RNA as part of the transcription process e. an enzyme that synthesizes RNA primers during DNA replication ____ 43. What are the coding segments of a stretch of eukaryotic DNA called? a. introns b. exons c. codons d. replicons e. transposons ____ 44. Once transcribed, eu ...
... d. an enzyme that synthesizes RNA as part of the transcription process e. an enzyme that synthesizes RNA primers during DNA replication ____ 43. What are the coding segments of a stretch of eukaryotic DNA called? a. introns b. exons c. codons d. replicons e. transposons ____ 44. Once transcribed, eu ...
The consequences of Rad51 overexpression for normal and tumor
... in the genome to repair the breaks. When HR is used for repair, in eukaryotes it is promoted by the recombinase Rad51, which binds to 3’-tailed single strands at the end of DSBs in a helical fashion and promotes pairing with homologous DNA sequences as a prelude to strand invasion and repair of the ...
... in the genome to repair the breaks. When HR is used for repair, in eukaryotes it is promoted by the recombinase Rad51, which binds to 3’-tailed single strands at the end of DSBs in a helical fashion and promotes pairing with homologous DNA sequences as a prelude to strand invasion and repair of the ...
Sequence-specific RNA Photocleavage by Single
... substrate for the sequence-specific photocleavage has the frequency of one repeat in every 4 bp, which reveals the broad generality of this molecular tool with respect to different RNA sequence. The length of two binding arms of the DNA strand is variable and the catalytic efficiency would not be af ...
... substrate for the sequence-specific photocleavage has the frequency of one repeat in every 4 bp, which reveals the broad generality of this molecular tool with respect to different RNA sequence. The length of two binding arms of the DNA strand is variable and the catalytic efficiency would not be af ...
From Gene to Protein—A Historical Perspective - AP Central
... life, and the storage and transfer of this information are necessary for life to continue. In most cases, this information is passed from parent to offspring via deoxyribonucleic acid, or DNA. This double-stranded molecule provides a simple and elegant solution for the transmission of heritable info ...
... life, and the storage and transfer of this information are necessary for life to continue. In most cases, this information is passed from parent to offspring via deoxyribonucleic acid, or DNA. This double-stranded molecule provides a simple and elegant solution for the transmission of heritable info ...
16 System and a 10X Primer Pair Mix Stored in TE
... this change should not affect amplification results. This change is not necessary for the PowerPlex® Y System because TE-4 buffer is already used to prepare the PowerPlex® Y 10X Primer Pair Mix. We will continue to generate more real time stability data, and if the product shelf life is extended, th ...
... this change should not affect amplification results. This change is not necessary for the PowerPlex® Y System because TE-4 buffer is already used to prepare the PowerPlex® Y 10X Primer Pair Mix. We will continue to generate more real time stability data, and if the product shelf life is extended, th ...
Intrinsic sequence specificity of the Cas1 integrase directs
... activity (Arslan et al., 2014). The order of the two integration steps shown in Figure 1 is not yet ...
... activity (Arslan et al., 2014). The order of the two integration steps shown in Figure 1 is not yet ...
Specific inhibition of DNA polymerase (3 by its 14 kDa domain: role
... protease-sensitive site and can bind both single- and doublestranded nucleic acids (29; Prasad and Wilson, unpublished data). These domains, however, do not show any detectable polymerase activity. To prepare large amounts of protein, a plasmid carrying the coding sequence of the N-terminal 14 kDa d ...
... protease-sensitive site and can bind both single- and doublestranded nucleic acids (29; Prasad and Wilson, unpublished data). These domains, however, do not show any detectable polymerase activity. To prepare large amounts of protein, a plasmid carrying the coding sequence of the N-terminal 14 kDa d ...
Inactivation of the p53 Tumor Suppressor Gene
... cells derived from human cancers. In some of these cases, the Alu elements appear to have been participants in homologous recombina tion. Examples are the ALL-l oncogene in acute myeloid leukemia (9) and the ire oncogene in Ewing's sarcoma (10). The possible involvement of Alu-associated events in t ...
... cells derived from human cancers. In some of these cases, the Alu elements appear to have been participants in homologous recombina tion. Examples are the ALL-l oncogene in acute myeloid leukemia (9) and the ire oncogene in Ewing's sarcoma (10). The possible involvement of Alu-associated events in t ...
Education®
... weak bonding force found only between hydrogen and either nitrogen or oxygen atoms. 19. Introns – sections of the mRNA template that are cut out prior to translation into protein and, hence, do not code for amino acids. 20. Lagging Strand – the strand of double-stranded DNA that replicate ...
... weak bonding force found only between hydrogen and either nitrogen or oxygen atoms. 19. Introns – sections of the mRNA template that are cut out prior to translation into protein and, hence, do not code for amino acids. 20. Lagging Strand – the strand of double-stranded DNA that replicate ...
78780 TG DNA Replication and Transcription
... 4. Codon – a sequence of three (3) consecutive nitrogen-containing bases on mRNA that code for an amino acid or a stop signal. Codons are the basic component of the genetic code. 5. Complementary Base Pairs – specific pairs of nitrogen-containing bases that always bond together when double stranded ...
... 4. Codon – a sequence of three (3) consecutive nitrogen-containing bases on mRNA that code for an amino acid or a stop signal. Codons are the basic component of the genetic code. 5. Complementary Base Pairs – specific pairs of nitrogen-containing bases that always bond together when double stranded ...
the pdf of this lesson!
... 4. Codon – a sequence of three (3) consecutive nitrogen-containing bases on mRNA that code for an amino acid or a stop signal. Codons are the basic component of the genetic code. 5. Complementary Base Pairs – specific pairs of nitrogen-containing bases that always bond together when double stranded ...
... 4. Codon – a sequence of three (3) consecutive nitrogen-containing bases on mRNA that code for an amino acid or a stop signal. Codons are the basic component of the genetic code. 5. Complementary Base Pairs – specific pairs of nitrogen-containing bases that always bond together when double stranded ...
Assembly of RecA-like Recombinases
... Accessory Factors Act to Promote Assembly of Recombinase. Recombinases are able to promote strand exchange to form hybrid DNA in vitro without additional proteins. However, accessory factors can stimulate strand exchange. These factors can be divided into two broad classes: those that act before hom ...
... Accessory Factors Act to Promote Assembly of Recombinase. Recombinases are able to promote strand exchange to form hybrid DNA in vitro without additional proteins. However, accessory factors can stimulate strand exchange. These factors can be divided into two broad classes: those that act before hom ...
Chemistry and biology of DNA-binding small
... to DNA bases. This alteration results in the DNA being fragmented by repair enzymes in their attempts to replace the alkylated bases. A second mechanism by which alkylating agents cause DNA damage is the formation of cross-bridges, bonds between atoms in the DNA. In this process, two bases are linke ...
... to DNA bases. This alteration results in the DNA being fragmented by repair enzymes in their attempts to replace the alkylated bases. A second mechanism by which alkylating agents cause DNA damage is the formation of cross-bridges, bonds between atoms in the DNA. In this process, two bases are linke ...
Combing of Molecules in Microchannels
... understood, we were able to vary the shape of the interface in a continuous fashion by altering the rate at which the fluid was withdrawn from the channel. In particular, we were able to obtain a flat interface, shown in Figure 1E, as an intermediate state between the convex interface (Figure 1A) th ...
... understood, we were able to vary the shape of the interface in a continuous fashion by altering the rate at which the fluid was withdrawn from the channel. In particular, we were able to obtain a flat interface, shown in Figure 1E, as an intermediate state between the convex interface (Figure 1A) th ...
DNA Fingerprinting by Restriction Enzyme Patterns
... DNA typing (also called DNA profile analysis or DNA fingerprinting) is the process whereby the genomic DNA of an organism is analyzed by examining several specific, variable DNA sequences located throughout the genome. In humans, DNA fingerprinting is now used routinely for identification purposes. Human ...
... DNA typing (also called DNA profile analysis or DNA fingerprinting) is the process whereby the genomic DNA of an organism is analyzed by examining several specific, variable DNA sequences located throughout the genome. In humans, DNA fingerprinting is now used routinely for identification purposes. Human ...
Identifying the Genetic Material
... into the bacterial cells, while most of the viral proteins remain outside. The injected DNA molecules causes the bacterial cells to produce more viral DNA and proteins. This meant that the DNA, rather than proteins, is the hereditary material, at least in viruses. These important experiments, and ma ...
... into the bacterial cells, while most of the viral proteins remain outside. The injected DNA molecules causes the bacterial cells to produce more viral DNA and proteins. This meant that the DNA, rather than proteins, is the hereditary material, at least in viruses. These important experiments, and ma ...
Homologous recombination
Homologous recombination is a type of genetic recombination in which nucleotide sequences are exchanged between two similar or identical molecules of DNA. It is most widely used by cells to accurately repair harmful breaks that occur on both strands of DNA, known as double-strand breaks. Homologous recombination also produces new combinations of DNA sequences during meiosis, the process by which eukaryotes make gamete cells, like sperm and egg cells in animals. These new combinations of DNA represent genetic variation in offspring, which in turn enables populations to adapt during the course of evolution. Homologous recombination is also used in horizontal gene transfer to exchange genetic material between different strains and species of bacteria and viruses.Although homologous recombination varies widely among different organisms and cell types, most forms involve the same basic steps. After a double-strand break occurs, sections of DNA around the 5' ends of the break are cut away in a process called resection. In the strand invasion step that follows, an overhanging 3' end of the broken DNA molecule then ""invades"" a similar or identical DNA molecule that is not broken. After strand invasion, the further sequence of events may follow either of two main pathways discussed below (see Models); the DSBR (double-strand break repair) pathway or the SDSA (synthesis-dependent strand annealing) pathway. Homologous recombination that occurs during DNA repair tends to result in non-crossover products, in effect restoring the damaged DNA molecule as it existed before the double-strand break.Homologous recombination is conserved across all three domains of life as well as viruses, suggesting that it is a nearly universal biological mechanism. The discovery of genes for homologous recombination in protists—a diverse group of eukaryotic microorganisms—has been interpreted as evidence that meiosis emerged early in the evolution of eukaryotes. Since their dysfunction has been strongly associated with increased susceptibility to several types of cancer, the proteins that facilitate homologous recombination are topics of active research. Homologous recombination is also used in gene targeting, a technique for introducing genetic changes into target organisms. For their development of this technique, Mario Capecchi, Martin Evans and Oliver Smithies were awarded the 2007 Nobel Prize for Physiology or Medicine.