* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Amgen Bruce Wallace Transformation Labs (2-7)
Chemical biology wikipedia , lookup
Gene therapy wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene prediction wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Expression vector wikipedia , lookup
Point mutation wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Designer baby wikipedia , lookup
Genetic engineering wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genomic library wikipedia , lookup
Molecular cloning wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
DNA vaccination wikipedia , lookup
Genome editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
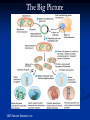
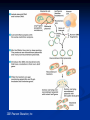
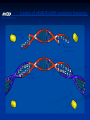
Amgen Bruce Wallace Transformation Labs (27) Timeline Thursday—Lecture Tuesday—Finish Lecture, Quiz, lab 2 Thursday--Lab 3, 4, 5 (Duffy does lab 6) Monday—Lab 7 Part 1 Tuesday—Lab 7 Part 2 Finals Friday—Packet Due Assignments Pre-lab notes worksheets: do these before we do that lab by reading the information and lab procedures Flowcharts—draw on the side or bottom of the procedures page Complete conclusion questions: do at the end of the lab each day Draw your lab 4 gel results at the end of the conclusion (use a ruler, make it nice!) Entire packet will be due 1/27/2012 (day of final)—no late work because this goes in semester grade! Prep. for Labs Week Before Make labels 10% bleach solution Aliquot chart Day before lab Lab 2—water bath set-up for 37 C Lab 3--Water baths set-up 70 C Lab 4--Pour 8 gels (35ml each) for lab 4 Pour .8% gels, add ethidium bromide (200ng/mL final or 1uL of 10mg/mL stock in gel prepared from 50mL), 6 well comb, SB buffer (2.4 grams agarose add up to 300mL TBE buffer) lab 5--ice Lab 5—water bath set-up at 42C Lab 6--Start overnight culture for lab 6 (use update instructions) Lab 7--Container of 10% at front for waste --Set-up chromatography columns Vocab. “transformed cell” – cell has acquired new characteristics “characteristics” – due to the expression of incorporated foreign genetic material Gene expression – process by which the information encoded in a gene is converted into an observable phenotype Gene regulation – control mechanisms that turn genes on or off Inducible proteins – synthesis is regulated depending on the bacterium’s nutritional status Thank you Francois Jacob and Jacques Monod! Prokaryote operon model of gene control Repressors and activators are “trans-acting” – affect expression of their genes no matter on which DNA molecule in the cell these are located. Overview of Labs Lab 2—Restriction Analysis of pARA and pKAN-R Cut the 2 plasmids using restriction enzymes Lab 3—Ligation of pARA/pKAN-R Restriction Fragments Producing a Recombinant Plasmid, pARA-R Insert the gene of interest into the pARA plasmid from the pKAN-R Lab 4—Confirmation of Restriction and Ligation Using Agarose-Gel Electrophoresis Run a gel to confirm the ligation in lab 3 worked (we want to make sure the gene of interest was inserted into that plasmid, if not, there is no reason to transform the plasmid into the bacteria) Lab 5—Transforming Escherichia coli with a Recombinant Plasmid Insert pARA-R (plasmid with our gene of interest) into bacteria using shock treatment, grow bacteria on plates, plasmid will produce proteins from our gene Lab 6--Preparing an Overnight Culture of Escherichia coli Take a colony that has the gene of interest from the plate, put into broth to replicate, now we have tons of bacteria (so tons of our geneprotein!) Lab 7—Purification of mFP from an Overnight Culture Lyse the bacteria cell, isolate the desired protein using chromatography The Big Picture 2005 Pearson Education, Inc. 2005 Pearson Education, Inc Background Concepts 1) What are Plasmids? 2) How can we modify plasmids? Restriction Enzymes 3) Origins of restriction enzymes. 4) A close look at restriction enzymes. 5) Understanding plasmid diagrams. What are Plasmids? In this Lecture… Circular DNA that is used by bacteria to store their genetic information. Modifying plasmids to include extra genes allows for the production of new proteins. How Can We Modify Plasmids? In this Lecture… 1) Restriction Enzymes 2) Restriction Enzyme attached to DNA before cleavage BamHI, HindIII, etc. Where do they come from? How do they work? Different restriction enzymes do different things. DNA Ligase Origins of Restriction Enzymes 1) Bacteria produce restriction enzymes to protect against invading viral DNA/RNA. Origins of Restriction Enzymes 2) The enzymes cut the invading DNA/RNA, rendering it harmless. Restriction Enzyme in Action Sticky Ends 1) DNA strand with EcoRI restriction site highlighted. 2) EcoRI restriction enzyme added (outline of separation about to occur). 3) Restriction fragments separate, with “sticky ends” at each edge. Adding DNA Ligase Sticky Ends DNA ligase bonds sticky ends cut with the same restriction enzyme. Sticky ends cut with different restriction enzymes will not bond together. Why? Because the base pair sequence of the two sticky ends will be different and not match up. Plasmids Can Be Drawn to Show the Genes They Carry Plasmid Name Bp size In this diagram: Blue and Orange are drawn as genes. Triangles are indicating the known restriction sites for a restriction enzyme. (shapes can vary) Plasmid Maps are more complex. Plasmid Maps Indicate Restriction Sites and Genes Application Exercise Make Recombinant DNA Using Restriction Enzymes DNA From Two Sources (Restriction Sites Labeled) Circular DNA Linear DNA Application of Restriction Enzymes Adding DNA Ligase Recombinant DNA Plasmid Many possible recombinant DNA plasmids can be produced, but this was the desired plasmid for the experiment. Many Other Recombinant Possibilities …and many more! Plasmid DNA Insertion DNA plasmids can be inserted into bacteria using a variety of laboratory processes. Transgenic Colony Allowed to Grow How Do We Get the Desired Plasmid? Recombinant plasmids Transformation of bacterial cells through electroporation. Bacteria are then moved to a growth plate, and grown on selective media to “weed out” cells that have not picked up the desired plasmid. Restriction fragments will ligate randomly, producing many plasmid forms. Bacterial insertion would be necessary, then colony growth, and further testing to isolate bacteria with the desired plasmid. Running Digested DNA Through Gel Electrophoresis Goals of this Hands-On Lab Take plasmid DNA that has been previously cut with restriction enzymes and compare that to a plasmid NOT cut with restriction enzymes, by running them through a gel. Look for different banding patterns and understand how to read them. Predict what kind of banding pattern a plasmid will make based on: 1. 2. The restriction enzyme used. The plasmid’s structural shape. Gel Box Loading Techniques Look directly down the axis of the pipette. Loading dye makes the sample heavy, but it can still easily swish out of the well. Squirt down slowly. Take the tip out of the buffer. Then release the plunger. If you don’t do that, you will suck the sample back up. Add DNA samples and ladder to the wells and “run to red!” 10 kb 8 kb 6 kb 5 kb 4 kb 3 kb 2 kb 1 kb .5 kb Sample fragments move toward positive end. Analyzing Your Gel What Makes Up the Banding Pattern in Restricted DNA? 1400 Bp 2000 Bp Lancer Plasmid 6700 Bp 3300 Bp The restriction enzyme cleaves the DNA into fragments of various sizes. Each different size fragment will produce a different band in the gel. Remember that fragments separate into bands based on size. What Makes Up the Banding Pattern After Adding DNA Ligase? Several combinations of plasmids will result from the reaction. The many forms will contribute to different bands. (See following slides for chemical and structural forms) Different Recombinant Forms Adding DNA Ligase does not always make the desired plasmid! Few if any could be what you wanted. Think about the large number of possible combinations. Different Structural Forms circle “multimer” Nicked Circle Linear “nicked-circle” Supercoiled Different structural forms produce different bands. A+ Multimer Nicked Super Coiled 5 Kb Linear Fragment Linear Fragment 10 Kb Ladder 10 Kb Ladder 10 Kb Ladder A- What Are Some Applications of Recombinant DNA Technology? Bacteria, Yeasts, and Plants can all be modified to produce important pharmaceuticals, enriched foods, and industrial products. Biotechnology Lab Program Bruce Wallace pKAN-R/pARA Sequence Laboratory Protocols by: Marty Ikkanda Powerpoint by: Anthony Daulo Pierce College, Woodland Hills, CA V.1.2.4 Restriction analysis of pKAN-R and pARA Bruce Wallace pKAN-R 5408 bp rfp 702 bp pARA 4058 bp 40 bp Restriction analysis of pKAN-R and pARA Bruce Wallace Restriction fragments after digest with Hind III and BamH I BamH I Hind III 4018 bp BamH I Hind III 4706 bp BamH I Hind III 702bp Hind III BamH I 40 bp Restriction analysis of pKAN-R and pARA Bruce Wallace Prediction for restriction gel M K+ K- A+ A- 10000 8000 5000 4000 3000 2000 1500 1000 500 M K+ K- A+ A- Ligation of pKAN-R/pARA restriction fragments Bruce Wallace BamH I sticky end 3’ 5’ 3’ 5’ sticky end 5’ 5’ 3’ 3’ Hind III BamH I sticky end Hind III sticky end 3’ 5’ 3’ 5’ Ligation of pKAN-R/pARA restriction fragments Bruce Wallace Recombinant plasmid of interest pARA-R 4720 bp rfp 702bp Restriction analysis of pKAN-R and pARA Bruce Wallace Confirmation of restriction and ligation M K+ K- A+ A- L 10000 8000 5000 4000 3000 2000 1500 1000 500 M K+ K- A+ A- L Preparing competent cells for transformation Bruce Wallace Lipid bilayer (inner) Adhesion zone Peptidoglycan layer Lipid bilayer (outer) Calcium ions Transforming Escherichia coli with pARA-R Bruce Wallace Competent Cells pARA-R Recombinant Plasmids Transforming Escherichia coli with pARA-R Bruce Wallace Lipid bilayer (inner) Peptidoglycan layer Adhesion zone Lipid bilayer (outer) Calcium ions pARA-R Growth of transformed bacteria on various plates Bruce Wallace P+ plates LB LB/amp P- plates No growth LB LB/amp LB/amp/ara Why don’t we see the red protein in any LB growth media? Cells conserve energy and resources The rfp gene requires a specific substrate (arabinose) to be turned on (expressed) Preparing an overnight culture of E. Coli for RFP expression Bruce Wallace Colony isolation and culture LB/amp/ara broth Many of the red colonies picked from a Lab 5 plate appear to contain cells that are interfering with rfp expression. When there is a mixed culture of red and white (nonexpressing) cells, the white cells will grow faster than those that are using their resources producing mFP. = less mFP produced for purification. RFP expression Bruce Wallace araC gene PBAD Transcription mRNA Translation araC protein rfp gene RFP expression Bruce Wallace araC protein prevents RFP transcription by causing a loop to form in the region of the r fp gene araC gene araC protein PBAD rfp gene RFP expression Bruce Wallace arabinose Arabinose – araC protein complex prevents DNA looping and helps to align RNA polymerase on the promoter site (PBAD). RFP (red fluorescent protein) Translation RNA polymerase arabinose araC protein araC–protein complex mRNA Transcription araC gene PBAD rfp gene Bruce Wallace RFP Purification of RFP from an overnight culture Bruce Wallace Overnight culture Cell pellet with RFP Lysed cells Pellet RFP with cell debris binding buffer Lab Tips/Reminders Add initials or group # to tubes Anything that touches bacteria must go in sterilizer Sterile technique Carefully READ and FOLLOW the lab protocol. Using bacteria Contamination may affect results Be sure lab partners communicate No Food or Drinks Agar Plate tips (Lab 5) Label the bottom of the plates at the edges Note the plate markings: I=LB, II=LB/amp, III=LB/amp/ara Samples go on the agar, not the lid Open like clam shells Agar is like finger jello, firm but not invincible, be gentle Turn the plates upside down (lids down) for incubation Sterile technique tips Always follow the protocol carefully – know what you’re doing Work quickly – less time = less opportunities for contamination Do not leave any container (tube, plate) open any longer than needed Watch what your equipment touches – there is no “5 second rule” here. All tips, tubes and spreaders in the “contaminated waste” container at the end of the lab. Look at labels Clam shell technique You need to aliquot and centrifuge twice to get a sufficient number of cells = product. Be sure the centrifuge has a balanced number of tubes. Be careful not to disturb the resultant pellets. When “wicking” don’t let the towel touch the pellet Supernatant and wicking towel go in disinfectant (10% bleach solution) containing beaker Incubate in 37 C water bath (60 min.) instead of overnight at room temperature. Freeze – ice crystals also help to lyse (break open) Science for LIFE