* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download LIVING WITHOUT OXYGEN
Survey
Document related concepts
Hedgehog signaling pathway wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Magnesium transporter wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Signal transduction wikipedia , lookup
Phosphorylation wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein moonlighting wikipedia , lookup
Gene regulatory network wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Transcript
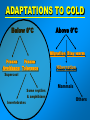
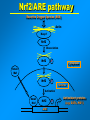
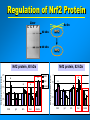
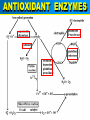
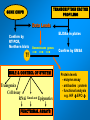
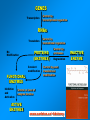
VERTEBRATE FREEZE TOLERANCE ADAPTATIONS TO COLD Below 0°C Above 0°C Migration Stay warm Freeze Freeze Avoidance Tolerance Hibernation Supercool Some reptiles & amphibians Invertebrates Mammals Others WOOD FROG Rana sylvatica TO SURVIVE FREEZING • Alter metabolism to synthesize cryoprotectants (polyols, sugars) • Defend against intracellular desiccation • Suppress metabolic rate ACCOMPLISHED BY: • Activate signaling enzymes in every cell - ‘SAP’ kinases - Role: reversible controls on cell processes Up-regulate selected genes SURVIVING FREEZING • Extracellular freezing only • Up to 70% of body water frozen • High ‘polyols’ • Acclimation required • Glucose • Glycerol • Sorbitol [ i + e Factors] Nucleus mRNAs GENES ON/OFF [Trans.F] CHO PROTEINS PATHWAYS ATP K AA ? SAPK P Ca+2 PROT SMW FAT ADP ATP KINASES (2nd) GENES Na MITO ETC WOOD FROG CRYOPROTECTANTS • Blood glucose rises from ~5 mM to 200-400 mM • Glucose triggered by ice formation • Made from liver glycogen (180 mg/g) • Glucose distribution via Blood: Liver > Core organs > Periphery 250 Glucose, mM • Liver is ~12% of body mass Blood 200 Liver 150 Heart 100 Kidney 50 Muscle 0 0 1 2 Days frozen 3 1 3 5 7 9 Days thawed 11 GLYCOGEN PHOSPHORYLASE Glycogen + Pi kinase Phos a Phos b phosphatase Glucose-1-P + glycogen (n-1) Liver Phosphorylase a Activity, U/g 0 2 5 30 60 2 3 1 2 min hours days TIME OF FREEZING 3 13 1 3 4 min hours TIME OF THAW PROTEIN KINASES PROTEIN PROTEIN-(P)n nATP nADP • Covalent modification by phosphorylation • Families of protein kinases: PKA (cAMP), PKG (cGMP), CaMK (Ca2+), PKC (Ca2+,PL,DG) • SAPKs : daisy chain phosphorylations • Regulation is via interconversion of active vs subactive forms of protein substrates [ i + e Factors] Nucleus mRNAs GENES ON/OFF [Trans.F] CHO PROTEINS PATHWAYS ATP K AA ? SAPK P Ca+2 PROT SMW FAT ADP ATP KINASES (2nd) GENES Na MITO ETC FREEZE INDUCED CHANGES • • • • • Protein Synthesis slows to 1% Pumps & channels closed Energy Production slows to 5% Energy Utilization slows to 2% Few ‘SAP’ kinases activated • Gene ‘inactivation’ ( mRNA ) • Few Genes activated (1-2%) p38 Pathway Signaling Path activated by ?? FREEZE INDUCED CHANGES • • • • • • • Protein Synthesis slows to 1% Pumps & channels closed Energy Production slows to 5% Energy Utilization slows to 2% Few ‘SAP’ kinases activated Gene ‘inactivation’ ( mRNA) Few Genes activated ROLE OF TRANSCRIPTION • Global rate of mRNA synthesis depressed. Method: nuclear run-on • Are selected genes up-regulated ? • TO ASSESS GENE UPREGULATION: What new mRNAs are created - cDNA library, Gene Chip TURNING OFF GENES: Role of Epigenetics Epigenetics: - Stable changes in gene activity that do not involve changes in DNA sequence Common mechanisms: - DNA methylation - Histone modification / histone variants - Regulatory non-coding RNAs cDNA Arrays - Methods - Materials - Sources - Publications FREEZE-INDUCED GENES: WOOD FROGS cDNA Library / Gene Chip • • • • Transcription Factors Mito ETC; Transporters AOE & Shock proteins The Unknowns: Fr10, Li16, FR47 Storey KB 2004. Cryobiology 48, 134-145 TRANSCRIPTION FACTORS • • • • • ATF (Glucose Regulated Proteins) HIF (O2), HSF (Hsp) NFkB (IkB-P), Nrf-2 (GST), NRF-1 PPAR, PGC, RXR, chREBP, CREB-P STAT, SMAD, p53-P, HNF, AP (1,2) • Methods: EMSA, PCR CONTROL REGION OF A TYPICAL EUKARYOTIC GENE Epigenetics: • microRNA • RNA Polymerase-P • Histones modified • HDAC / HAT changes FREEZE-INDUCED GENES: WOOD FROGS cDNA Library / Gene Chip • Transcription Factors NRF -2 • AOE Storey KB 2004. Cryobiology 48, 134-145 NRF-2 • Increased NFR-2 protein • Increased NFR-2 in the Nucleus • Increased levels of co-Tf: MafG • Downstream gene activation: • GST, HO-1, HO-2, Peroxiredoxin Nrf2/ARE pathway Reactive Oxygen Species (ROS) Actin Keap1 Nrf2 Dissociation Nrf2 P Nrf2 P Cytoplasm Small Maf Nucleus Activation Small Maf Nrf2 ARE P Antioxidant proteins (e.g. GSTs, HO1) Regulation of Nrf2 Protein Liver C C F F Actin Nrf2 82 kDa 40 kDa Nrf2 Nrf2 protein expression (~82KDa) Nrf2 protein, 82 kDa 2.5 2 a a control frozen a 1.5 1 a recovery a 0.5 0 brain gut skin liver muscle Relative protein levels Relative protein levels Nrf2 protein expression(~40KDa) Nrf2 protein, 40 kDa 2 1.5 a a 1 a a 0.5 0 brain gut skin liver muscle Regulation of MafG Small Maf Nrf2 Antioxidant proteins (e.g. GSTs, HO1) P ARE mafG protein expression MafG protein a Relative protein level 2.5 2 control a a frozen a recovery a 1.5 a 1 0.5 0 brain muscle gut liver skin Glutathione S-Transferase Pi isozyme in Wood Frogs GST Pi mRNA expression GST Pi mRNA a * control 2.5 frozen recovery 2 mRNA increases - transcriptional control a 1.5 1 0.5 GST Pi protein expression 0 brain gut liver kidney heart GST Pi protein muscle 12 Protein increases - translational control Relative protein level Relative mRNA levels 3 control a 10 frozen recovery 8 6 a a a 4 a a a 2 a 0 brain gut liver kidney heart muscle Conclusions Activation of the Nrf2 pathway: Activated in early-late torpor, along with downstream gene protein products Increased GST protein and activity Result: Detoxification of H2O2, intracellular signaling control ANTIOXIDANT ENZYMES TRANSCRIPTION FACTORS • • • • • ATF (Glucose Regulated Proteins) HIF (O2), HSF (Hsp) NFkB (IkB-P), Nrf-2 (GST), NRF-1 PPAR, PGC, RXR, chREBP, CREB-P STAT, SMAD, p53-P, HNF, AP (1,2) • Methods: EMSA, PCR TRANSCRIPTION FACTOR PROFILING GENE CHIPS Data Leads Confirm by RT-PCR, Northern blots ELISAs in plates Downstream genes Confirm by EMSA Tf ROLE & CONTROL OF SYSTEM Transgenics Cell Assay RNAi Knock out Epigenetics FUNCTIONAL ASSAYS Protein levels - enzyme assay - antibodies : protein - functional analysis e.g. HIF EPO Unique Animal Stress Model Vertebrate whole-body freeze tolerance Tissue cryopreservation Tolerance of extreme ischemia and hyperglycemia GENES Transcription Control by transcriptional regulation RNAs Translation No Modification FUNCTIONAL ENZYMES Inhibition and Activation Control by translational regulation PROTEINS (ENZYMES) Covalent modification Control by proteases Degradation Control by posttranslational modification Control at level of enzyme function ACTIVE ENZYMES www.carleton.ca/~kbstorey INACTIVE ENZYME