* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download TWO GENES ENCODING FUNCTIONAL PECTIN
Survey
Document related concepts
Homology modeling wikipedia , lookup
Protein folding wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Protein domain wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein purification wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Alpha helix wikipedia , lookup
Protein structure prediction wikipedia , lookup
Protein moonlighting wikipedia , lookup
Western blot wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
List of types of proteins wikipedia , lookup
Transcript
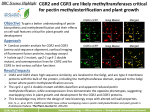
AtPMEI-1 AND AtPMEI-2: TWO GENES ENCODING FUNCTIONAL PECTIN METHYLESTERASE INHIBITORS OF Arabidopsis thaliana Alessandro Raiola1, Laura Camardella2, Alfonso Giovane3, Benedetta Mattei1 , Giulia De Lorenzo1, Felice Cervone1, Daniela Bellincampi1 1 Dipartimento di Biologia Vegetale, Università di Roma “La Sapienza”, Piazzale Aldo Moro 5, 00185 Roma, Italy 2 Institute of Protein Biochemistry, CNR, Via Marconi 10, I-80125 Napoli, Italy 3 Department of Biochemistry and Biophysics, 2nd University of Naples, Via Costantinopoli 16, I80138 Napoli, Italy A proteinaceous inhibitor of pectin methylesterase (PMEI) has been reported in kiwi but to date no other proteins acting as PMEI have been found in plants. Two sequences closely related to PMEI from kiwi were identified in Arabidopsis thaliana. The corresponding cDNAs encode cell wall proteins of 173 and 176 amino acid residues respectively. The sequences corresponding to the mature proteins were cloned and expressed in a heterologous system. The purified proteins were characterized and shown to be functional PMEIs. The inhibitory activity occurs through the formation of a 1:1 complex with tomato PME. Both proteins share with kiwi PMEI a predicted alpha-helix structure and the conservation of four Cys residues involved in the formation of two disulfide bridges as well as a fifth Cys residue bearing a free thiol group. The interaction parameters of the two new inhibitors with PME purified from tomato fruits have been studied . We gratefully acknowledge funding from the Commission of the European Communities contract QLK1-2000-00811 “Solving the Problem of Glycosidase Inhibitors in Food Processing” and MIUR-COFIN.