* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Laser Light Scattering
Survey
Document related concepts
Molecular evolution wikipedia , lookup
Maurice Wilkins wikipedia , lookup
Gel electrophoresis wikipedia , lookup
Western blot wikipedia , lookup
Community fingerprinting wikipedia , lookup
Non-coding DNA wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Circular dichroism wikipedia , lookup
DNA vaccination wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Agarose gel electrophoresis wikipedia , lookup
Molecular cloning wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Transcript
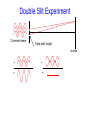
Laser Light Scattering - Basic ideas – what is it? - The experiment – how do you do it? - Some examples systems – why do it? Double Slit Experiment Coherent beam Extra path length screen + = + = Light Scattering Experiment Scatterers in solution (Brownian motion) Scattered light Laser at fo Narrow line incident laser Doppler broadened scattered light Df 0 is way off scale fo Df ~ 1 part in f 1010 - 1015 More Detailed Picture detector q Inter-particle interference Detected intensity Iaverage time How can we analyze the fluctuations in intensity? Data = g(t) = <I(t) I(t + t)>t = intensity autocorrelation function Intensity autocorrelation • g(t) = <I(t) I(t + t)>t t For small t For larger t t g(t) tc t What determines correlation time? • Scatterers are diffusing – undergoing Brownian motion – with a mean square displacement given by <r2> = 6Dtc (Einstein) • The correlation time tc is a measure of the time needed to diffuse a characteristic distance in solution – this distance is defined by the wavelength of light, the scattering angle and the optical properties of the solvent – ranges from 40 to 400 nm in typical systems • Values of tc can range from 0.1 ms (small proteins) to days (glasses, gels) Diffusion • What can we learn from the correlation time? • Knowing the characteristic distance and correlation time, we can find the diffusion coefficient D • According to the Stokes-Einstein equation kBT D 6h R where R is the radius of the equivalent sphere and h is the viscosity of the solvent • So, if h is known we can find R (or if R is known we can find h) Why Laser Light Scattering? 1. 2. 3. 4. 5. Probes all motion Non-perturbing Fast Study complex systems Little sample needed Problems: Dust and best with monodisperse samples Some Examples Superhelical DNA where = Watson-Crick-Franklin double stranded DNA pBR322 = small (3 million molecular weight) plasmid DNA Laser light scattering measurements of D vs q give a length L = 440 nm and a diameter d = 10 nm DNA-drug interactions: intercalating agent PtTS produces a 26o unwinding of DNA/molecule of drug bound Since D ~ 1/size, as more PtTS is added and DNA is “relaxed,” we expect a minimum in D As drug is added DNA first unwinds to open circle and then overwinds with opposite handedness. At minimum in D the DNA is unwound. This told us that there are 34 superhelical turns in native pBR pBR is a major player in cloning – very important to characterize well Antibody molecules • Technique to make 2-dimensional crystals of proteins on an EM grid (with E. Uzgiris at GE R&D) Change pH 60o 120o Conformational change with pH results in a 5% change in D – seen by LLS and modeled as a swinging hinge Aggregating/Gelling Systems Studied at Union College • Proteins: – Actin – monomers to polymers and networks Study monomer size/shape, polymerization kinetics, gel/network structures formed, interactions with other actin-binding proteins Why? Epithelial cell under fluorescent microscope Actin = red, microtubules = green, nucleus = blue Aggregating systems, con’t – BSA (bovine serum b amyloid - insulin – Chaperones what factors cause or promote albumin) aggregation? what is the structure of the aggregates? how can proteins be protected from aggregating? Focus on the onset of gelation – • Polysaccharides: – Agarose – Carageenan what are the mechanisms causing gelation? how can we control them? what leads to the irreversibility of gelation? Collaborators and $$ • • • • • • • • • • • • • • Nate Poulin ’14 & Christine Wong ‘13 Michael Varughese ’11 (med school) Anna Gaudette ‘09 Bilal Mahmood ’08 & Shivani Pathak ’10 (both in med school) Amy Serfis ‘06 & Emily Ulanski ’06 (UNC, Rutgers ) Shaun Kennedy (U Michigan, Ann Arbor in biophysics) Bryan Lincoln (PhD from U Texas Austin, post-doc in Dublin) Jeremy Goverman (medical school) Shirlie Dowd (opthamology school) Ryo Fujimori (U Washington grad school) Tomas Simovic (Prague) Ken Schick, Union College J. Estes, L. Selden, Albany Med Gigi San Biagio, Donatella Bulone, Italy Thanks to NSF, Union College for $$