* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Microbial Ecology
Microorganism wikipedia , lookup
Disinfectant wikipedia , lookup
Horizontal gene transfer wikipedia , lookup
Magnetotactic bacteria wikipedia , lookup
Bacterial cell structure wikipedia , lookup
Triclocarban wikipedia , lookup
Metagenomics wikipedia , lookup
Marine microorganism wikipedia , lookup
Phospholipid-derived fatty acids wikipedia , lookup
Bacterial morphological plasticity wikipedia , lookup
Bacterial taxonomy wikipedia , lookup
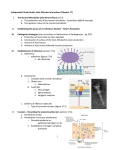
Microb Ecol (2012) 64:555–569 DOI 10.1007/s00248-012-0039-5 Molecular Analysis of Bacterial Microbiota Associated with Oysters (Crassostrea gigas and Crassostrea corteziensis) in Different Growth Phases at Two Cultivation Sites Natalia Trabal & José M. Mazón-Suástegui & Ricardo Vázquez-Juárez & Felipe Asencio-Valle & Enrique Morales-Bojórquez & Jaime Romero Received: 6 December 2011 / Accepted: 6 March 2012 / Published online: 27 March 2012 # Springer Science+Business Media, LLC 2012 Abstract Microbiota presumably plays an essential role in inhibiting pathogen colonization and in the maintenance of health in oysters, but limited data exist concerning their different growth phases and conditions. We analyzed the bacterial microbiota composition of two commercial oysters: Crassostrea gigas and Crassostrea corteziensis. Differences in microbiota were assayed in three growth phases: postlarvae at the hatchery, juvenile, and adult at two grow-out cultivation sites. Variations in the microbiota were assessed by PCR analysis of the 16S rRNA gene in DNA extracted from depurated oysters. Restriction fragment length polymorphism (RFLP) profiles were studied using Dice’s similarity coefficient (Cs) and statistical principal component analysis (PCA). The microbiota composition was determined by sequencing temperature gradient gel electrophoresis (TGGE) bands. The RFLP analysis of post-larvae revealed homology in the microbiota of both oyster species (Cs>88 %). Dice and PCA analyses of C. corteziensis but not C. gigas showed differences in the microbiota according to the cultivation sites. The sequencing analysis revealed low bacterial diversity (primarily β-Proteobacteria, Firmicutes, and Spirochaetes), with N. Trabal : J. M. Mazón-Suástegui : R. Vázquez-Juárez (*) : F. Asencio-Valle : E. Morales-Bojórquez Centro de Investigaciones Biológicas del Noroeste, (CIBNOR), Mar Bermejo 195, Col. Playa Palo de Santa Rita, La Paz, Baja California Sur 23095, Mexico e-mail: [email protected] N. Trabal e-mail: [email protected] J. Romero Laboratorio de Biotecnología, Instituto de Nutrición y Tecnología de los Alimentos, Universidad de Chile, El Líbano 5524, Macul, Santiago, Chile Burkholderia cepacia being the most abundant bacteria in both oyster species. This study provides the first description of the microbiota in C. corteziensis, which was shown to be influenced by cultivation site conditions. During early growth, we observed that B. cepacia colonized and remained strongly associated with the two oysters, probably in a symbiotic host– bacteria relationship. This association was maintained in the three growth phases and was not altered by environmental conditions or the management of the oysters at the grow-out site. Introduction Oysters are the most produced and most valuable bivalve mollusks [23]. However, one of the main problems in the aquaculture of oysters is the repetitive episodes of mortality, which seriously reduces production. These outbreaks of disease affect the larval and post-larval stages in hatcheries and also affect juveniles and adults cultured in the natural environment. In some cases, studies have demonstrated diseases of bacterial etiology, including vibrioses nocardosis and infections by Chlamydia-, Mycoplasma-, and Rickettsia-like organisms [7, 39, 59, 65, 68, 75]. Because oysters may concentrate pathogenic microorganisms, the microbiota of oysters have been examined mainly from the public health perspective, and most of the studies have focused on fecal contamination, enteric pathogens, and pathogenic species of Vibrio [17, 34, 39, 54, 67, 75–77]. The association between aquatic invertebrates and gut microbes is typically attributed to the ingestion of bacteria [31, 62]. Research with aquatic organisms has suggested a variety of beneficial roles for microbiota, including the development of the host gastrointestinal tract, nutrition 556 (providing vitamins, enzymes, and essential fatty acids for the host), immune responses, and disease resistance [3, 19, 26, 31, 46, 56, 62, 64]. An increased susceptibility to infections may be related to the lack of the barrier provided by microbiota, which compete with pathogenic microorganisms for nutrients and space in the intestinal tract [5, 46, 51, 52, 61] or produce substances that inhibit pathogens [26, 28, 61]. In recent years, the natural bacterial microbiota of different oysters (Crassostrea gigas [33], Crassostrea virginica [40], Saccostrea glomerata [29], Ostrea chilensis [69], and Crassostrea iredalei [54]) have been described. However, these studies do not focus on the changes in microbiota that may occur during the growth of the oysters. With the exception of the study by Romero et al. [69], these reports do not distinguish between resident and transient bacteria, which lead to an overestimation of bacterial diversity within the microbiota. Because oysters are filter feeders, their gills are covered with mucus and vibrating cilia that facilitate gas exchange for respiration and simultaneously trap suspended particles, including bacteria and viruses [62]. In oysters, two categories of normal bacterial microbiota can be described: (1) indigenous or resident bacteria, including relatively fixed types of organisms that remain relatively stable over time and do not change with the particles ingested by the host (even if the bacteria are disturbed, they tend to spontaneously recover), and (2) transient or non-indigenous microbiota, which are composed of potentially pathogenic microorganisms that are ingested with the food, survive passage through the gut (possibly proliferating in the gut), and are voided with the feces [6, 31, 52]. During larval development, transient microbiota rapidly become residents of the oyster microbiota [8, 37, 51, 52, 62], although little is known about the dynamics and stability of microbiota in the juvenile and adult growth stages. It is, however, recognized that the physiology of the invertebrate marine organism can influence the gut microbiota [31, 62]. In bivalve mollusks, bacteria have been reported in the style, gland of Deshayes, esophagus, stomach, and intestine [31, 37, 51, 62]; however, many other studies on the microbiota were performed on the whole-oyster homogenate [31, 40, 63, 69]. Colonization by bacteria in the oyster gastrointestinal tract has a particular dependence on the external environment because of the flow of water passing through the digestive tract [26, 31, 62]. Considering that ambient conditions and seasonality have been shown to alter the gut bacteria community in some aquatic invertebrates, it is probable that the habitat can influence the conditions in the gut and may therefore be an important factor in the composition of gut microbiota [26, 31, 62]. However, studies of the changes in microbiota under different environmental conditions are lacking [40, 62, 63]. One of the most critical steps in the cultivation of these oysters is the planting of post-larvae (spat) from production N. Trabal et al. at hatcheries to the field, where large numbers die [7, 11, 18]. Additionally, at high stocking densities in the field, the identification of microbiota is important because the resident microbiota can be disturbed, and transient microorganisms subsequently exploit a temporary advantage [51, 52]. Productivity is related to larval transfer to the field and environmental conditions that can alter the bacterial community associated with oysters. During the past 20 years, oyster cultivation in Mexico has improved; commercially important mollusks, such as the Pacific oyster C. gigas (Thunberg, 1793) and the Cortez oyster C. corteziensis (Hertlein, 1951), have been used. C. gigas is the most cultivated and widely studied oyster worldwide [30] and is the most prevalent species along the Pacific coast; however, massive deaths have led to many studies of the regional species C. corteziensis [11, 13, 47–49]. Presently, the microbiota composition of Cortez oyster is unknown. In this study, using molecular techniques, we analyzed the composition of the bacterial microbiota of C. gigas and C. corteziensis. Variations and differences in the bacterial microbiota composition were assayed in three growth phases: post-larvae at the hatchery, juvenile, and adult at two grow-out cultivation sites in the field. In particular, we studied the variation of the microbiota composition when the post-larvae were planted at different cultivation sites. Methods A synthetic diagram of the experimental design and methodology used in this study is shown in Fig. 1. Oyster Collection and Experimental Design Larvae of C. gigas and C. corteziensis were collected from certified breeding adults following Office International des Epizooties (OIE) standards. Hatchery procedures were followed as described by Mazón-Suástegui et al. [47–49]. To obtain and culture embryos and larvae at Playa Eréndira (PE) (24°14′09″N, 110°18′24″W), broodstock were conditioned for maturation under controlled temperature conditions and food supply in the hatchery prior to being induced to spawn by thermal stimulation (Fig. 1). Swimming larvae were fed with increasing concentrations of cultured Isochrysis galbana and Chaetoceros calcitrans microalgae (25,000–60,000 cells ml–1) while decreasing the culture density (10−1 larvae ml–1). Post-larvae (small and delicate young oyster) were cultivated in upwelling recirculation systems using the same microalgae diet mentioned above; until they reach the optimal post-larvae planting size, these post-larvae are known as spats (mean shell height 3–5 mm; 6 weeks of age after settling). In August 2009, a set of post- Bacterial Microbiota in Two Commercial Oysters 557 Figure 1 Diagram of the experimental design used in this study larvae was transferred (planting) to two grow-out cultivation sites (Fig. 1). An initial culture of spats from the oysters in the cultured site was performed in NestierTM-type trays suspended from a floating long-line. The juvenile oysters (3–5 cm in shell height; age 5 months after planting; Fig. 1) were cultured in cages suspended on rigid structures near the bottom. The adult oysters (6–12 cm in shell height; age 6–10 months after planting, Fig. 1) were cultured exclusively at the bottom of the cages for improved stability and food supply. Two different grow-out cultivation locations were used: Punta Botella (PB; 25°18′14.52″N, 112°04′35″W) at Bahía Magdalena and Bahía de Topolobampo (BT; 25°54′N, 108° 33′W) (Fig. 2). Thirty samples of juveniles of C. gigas and C. corteziensis were collected at both grow-out cultivation areas in December 2009 and February 2010, respectively. The adult oysters (n030 of each species) were collected in April 2010 (Fig. 1). At this time, at both sites, gonad maturation was not detected in the oysters. The samples were transported on ice to the laboratory. On arrival, the surface of the shells was scrubbed with filtered and UV-sterilized seawater for epifauna elimination. Next, the shells were washed with 70 % ethanol and were finally depurated for 72 h in filtered and UV-sterilized seawater with constant aeration according to the Scheduled Depuration Process [24]. Transient microbiota was removed from the oysters following methods previously reported by Son and Fleet [72] and Lee et al. [41]. After cleaning, the juvenile and adult oysters were dissected in a sterile Petri dish using one sterile scalpel blade for each oyster. After discarding the intervalve liquid and adductor muscles, we proceeded to dissect the organisms (juveniles and adults). The tissues of the gastrointestinal tract, gills, and mantle Figure 2 Collection sites for C. gigas and C. corteziensis specimens. p-L0post-larvae; J0juvenile; A0adults. PE0hatchery production of post-larvae at Playa Eréndira; PB0grow-out cultivation site at Punta Botella; BT0grow-out cultivation site at Bahía Topolobampo 558 N. Trabal et al. were separated, and the gastrointestinal tract was frozen at −20°C. Post-larval oysters (n030, for each species) were vigorously rinsed in sterile seawater, and after the intervalve liquid and adductor muscles were eliminated, the organisms were frozen at −20°C. In all cases, at the time of dissection, the oysters were alive, and the appearance of the bodies was normal with no evidence of infection. temperate Pacific environmental conditions that are more suitable for C. gigas, which is a temperate water species, whereas the subtropical conditions in parts of the Gulf of California are more suitable for C. corteziensis, which is a subtropical species [13, 47, 48]. Environmental Conditions of the Grow-Out Cultivation Area The total tissue of the post-larvae was homogenized by lysis using a buffer containing Tris–EDTA–SDS (100 mM NaCl, 50 mM Tris [pH 8], 100 mM EDTA [pH 8], sodium dodecyl sulfate [SDS 1 %]) and 20 μl proteinase K (20 mg ml−1) (Sigma, St. Louis, MO) and was incubated for 1 h at 65°C. The DNA was subsequently extracted with phenol/chloroform and precipitated with ethanol, as previously described [69]. The gastrointestinal tissues (cut into pieces) of juvenile and adult oysters were lysed with 50 ml of Tris–EDTA–SDS buffer (as previously described), and the DNA was extracted using a QIAmp® DNA Mini kit (Qiagen, Valencia, CA) following the manufacturer's instructions. A final purification was performed using the Wizard® SV Gel and PCR Clean-up System (Promega, Madison, WI). The concentration and quality of the DNA were determined at A260 and A280 nm using an ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE). The Punta Botella grow-out cultivation site (PB) is located in Bahía Magdalena, which has one of the most diverse ecosystems in the state of Baja California Sur. This location has low pollution and minor human activity, and it is influenced by oceanic currents of temperate seawater from further south. In Bahía Magdalena, the large lagoon has an irregularly shaped barrier island. There are numerous small estuaries and shallow channels bordered by mangroves that provide organic matter deposited as sediment. Mangroves are the common feeding habitat for communities of many vertebrate and invertebrate species and possess commercial and ecological value [1, 12]. The Bahía Topolobampo grow-out site (BT) is a coastal lagoon complex with relatively large mouths connecting to the Gulf of California that is influenced by oceanic currents. In the last several decades, BT has been supporting shrimp and oyster aquaculture projects. However, the lagoon water is contaminated by agricultural runoff and urban and industrial sewage effluents [57, 58]. The salinity at both sites ranged from 35 to 40 psu with an average of 37 psu. The weekly sea surface temperature (SST) data for 2009– 2010 were derived from the Moderate Resolution Imaging Spectroradiometer-Aqua (MODIS-Aqua) satellite. The SST data were downloaded from the National Aeronautics and Space Administration web site (http://modis.gsfc.nasa.gov/ data/). The monthly SST was estimated along a straight line centered at each one of the study areas, and the corresponding mean was calculated. The following data show the SST during grow-out of juveniles and adults at both sites: Punta Botella (August, 26.6°C; September, 28.0°C; October, 26.2°C; November, 25.5°C; December, 23.4°C; January, 21.7°C; February, 20.6°C; March, 20.4°C; and April, 19.9°C) and Bahía Topolobampo (August, 27.9°C; September, 29.1°C; October, 26.7°C; November, 25.9°C; December, 24.6°C; January, 21.2°C; February, 19.7°C; March, 23.8°C; and April, 25.5°C). Temperate temperatures were found at the PB site, whereas the temperatures at the BT site were subtropical. A temperature drop was also noted in October, which indicated the beginning of autumn at both sites. C. gigas grew faster at PB, and C. corteziensis grew faster at BT. The difference is based on DNA Extraction and Purification PCR Amplification of the 16S rRNA Gene and Analysis of the Products The total DNA was extracted from each oyster (total tissue post-larvae and gastrointestinal tissues of juveniles and adults), diluted in nuclease-free water to obtain a concentration of 50 ng μl−1, and used as a template in PCR. Nested PCR of the V3–V5 region of the 16S rRNA gene was used to obtain profiles of the bacterial communities present in three different growth stages in both oyster species and at the two grow-out cultivation sites. Eubacterial 16S rRNA genes were amplified with the following primers: EUB 27 F (5′-AGAGTTTGATCCTGGCTCAG-3′) and EUB1492R (5′-GTTACCTTGTTACGACTT-3′) [43] in the first amplification and 341 F (5′-GCCTACGGGAGGCAGCAG-3′ with GC clamps at the 5′ end for temperature gradient gel electrophoresis analysis, TGGE) [53] and 939R (5′-CTTGTGCGGGCCCCCGTCAA TTC-3′) [70] for the second phase. The primers used in the specific amplification of the eubacterial 16S rRNA gene variable region were selected from a set of primers available in the literature and aligned in silico using the Sequence Match software with reference sequences (1,921,179 16S rRNAs) from the Ribosomal Database Project II Database (RDP II). The criteria used to choose the best primers were as follows: greater alignment with eubacteria domain sequences, Bacterial Microbiota in Two Commercial Oysters especially with the Proteobacteria group; lack of alignment with Archaea domain sequences; and no cross-amplification with the oyster genome. PCR was performed with a reaction mixture (25 μl) containing 0.2 mM of each deoxynucleoside triphosphate, 0.05 Uml−1 Platinum Taq DNA polymerase (Invitrogen, San Diego, CA), 1× polymerase reaction buffer, 2 mM MgCl2, and 0.25 pmol/ml−1 of each primer. The reaction mixtures were incubated in an Eppendorf Mastercycler (Eppendorf, Hamburg, Germany) with an initial denaturation at 95°C for 10 min followed by 15 cycles at 95°C for 1 min 30 s, 55°C for 1 min 30 s, 72°C for 1 min 30 s, and a final elongation at 72°C for 10 min for the first phase of the nested PCR. Subsequently, 1 μl of the amplified DNA was re-amplified with an initial denaturation at 94°C for 10 min followed by 20 cycles at 97°C for 1 min, 54°C for 1 min, 72°C for 1 min 30 s, and a final elongation at 72°C for 10 min for second phase of the nested PCR. The PCR products were analyzed using polyacrylamide gel electrophoresis and silver nitrate staining [20]. Restriction Fragment Length Polymorphism Analysis of the 16S rRNA Gene By applying restriction fragment length polymorphism (RFLP) analysis to the PCR amplification products of the 16S rRNA gene, we sought to observe variations of the resident microbiota in C. gigas and C. corteziensis in the three growth stages at two different grow-out sites. The products of the 16S rRNA gene amplification (10 μl) from the DNA of post-larval and gastrointestinal tissues of juveniles and adults were digested for 2 h at 37°C with 1.5 U of Alu1 restriction endonuclease (Invitrogen, San Diego, CA). The resulting fragments were subsequently analyzed by polyacrylamide gel electrophoresis and silver nitrate staining [20]. Each gel included BenchTop 100 bp DNA Ladder markers (Promega) and standard bacterial RFLP controls (Vibrio harveyi ATCC 14126, Listonella pelagia ATCC 25916) to validate the comparisons between the gels. Analysis of the Variation of Microbiota Using RFLP Patterns The RFLP profiles were analyzed using software (GelCompar II® 5.2, Applied Maths, Austin, TX) and by applying the Dice similarity index (Cs). The pairwise Cs is a similarity index used to compare microbiotal variation in different samples based on the presence or absence of bands in each lane in the gels [50]. Thus, two identical profiles create a Cs value of 100 %, whereas completely different profiles result in a Cs value of 0 %. Each lane in the gels (samples from individual oysters) can be compared with every other sample; therefore, the mean percent similarities (Cs values) were used to compare the similarity of microbiota profiles in 559 post-larvae, juveniles, and adults in the two grow-out sites [50]. Using this index, microbiota profiles derived from different species (C. gigas vs. C. corteziensis) and from each individual of the same species were compared. The cluster analysis results were displayed as a dendrogram. UPGMA was applied with GelComparII software. The reliability was tested by calculating the co-phenetic correlations of the nodes with tools available in the GelCompar program. Analysis of Microbiota Composition Using 16S rRNA Gene Temperature Gradient Gel Electrophoresis Using TGGE analysis, we determined the composition of the microbiota associated with the oysters. We chose individual samples that showed different RFLP profiles and triplicate samples of organisms having the same pattern from each group analyzed (the two species of post-larvae, juveniles, and adults from Punta Botella and Bahía Topolobampo). Products obtained from the nested PCR of the 16S rRNA gene were separated by TGGE and analyzed using the DCode System (Bio-Rad, Hercules, CA, USA) in 0.8 mm gels composed of 6 % [w/v] acrylamide, 0.1 % bisacrylamide, 8 M urea, 20 % [v/v] formamide, and 2 % [v/v] glycerol with 1× Tris–acetate (Tris 0.04 M, acetate 0.002 M, EDTA 0.001 M, pH 8.5) as the electrophoresis buffer. The polymerization was catalyzed by 110 μl of N-NN′-N′-tetramethylethylenediamine and 80 μl of 10 % ammonium persulfate solution added to 50 ml of the gel solution. The gel was loaded with 6 μl of 341f-939r-GC clamp PCR products and 1 μl of dye solution and run for 18 h 30 min at a fixed voltage of 65 V (±9 mA) with a parallel temperature gradient ranging from 66 to 70°C. After electrophoresis, the gels were stained for 1 h by incubation with SYBR Green at 25±2°C (room temperature). Each gel included standards containing PCR amplicons of known bacterial sequences ( % GC) to validate comparisons between gels [55]. The standard was composed of amplicons from the following bacteria: Pseudomona aeruginosa P1 (%GC: 51.8), V. harveyi ATCC 14126 (%GC: 53.1), Bacillus subtilus B4 (% GC: 54.6) and Citrobacter gillenii (%GC: 56). Sequence Analysis of the 16S rRNA Gene The dominant bands were identified by their corresponding intensity in each TGGE pattern. The bands were excised from the gel and eluted for 12 h at 25°C in 50 μl MilliQ water. After elution, 1 μl of DNA was used for re-amplification, using the same conditions described above. To test for the presence of similar amplicons, bands showing the same migration in different lanes were checked by RFLP using AluI (an enzyme that cuts frequently) [55]. Next, 16S rDNA from the re-amplified bands or from the 560 bacterial isolates was purified using the Wizard® SV Gel and PCR Clean-up System and sequenced (by Macrogen, Rockville, MD) with an automatic sequencer (Applied Biosystems ABI3730XL, Carlsbad, CA). The sequences were initially compared to the available databases using the basic Local Alignment Search Tool network service [2] and aligned with reference sequences using Sequence Match software from the Ribosomal Database Project II (RDP II) website [15] to determine the approximate phylogenetic affiliations. The sequences were deposited in GenBank (accession numbers JF522195–JF522232). Statistical Analysis We analyzed the differences in the bacterial communities associated with C. gigas and C. corteziensis in three growth phases (post-larvae, juvenile, and adult) between two growout cultivation sites using orthogonal empirical functions. The differences were estimated using a multivariate analysis, namely, the principal component analysis (PCA). The vectors computed from PCA are orthogonal and are uncorrelated, which prevents the regression on residuals [36]. The PCA permitted the ranking and simplification of the new variables, called principal components, and determination of the total variation of the data Xj, thereby explaining the variation with a few principal components (factor loadings≥0.7). Only the principal component was considered to be statistically significant for eigenvalues (l)>1.0 [36]. PCA was performed from correlation matrices generated from a binary matrix considering the presence/absence of the RFLP patterns bands, which were expressed as a value of the Dice similarity coefficient [25]. The PCA was conducted using the Statistica 8.0 software package (StatSoft Inc., Tulsa, OK). Results Similarity of Bacterial Microbiota in the Post-larvae of C. gigas and C. corteziensis The similar RFLP profiles obtained from the cluster of postlarvae of C. corteziensis revealed a homology in the microbiota of this oyster with Cs>92 % (Table 1, Fig. 3a). C. gigas, however, exhibited a lower similarity than did C. corteziensis with Cs>72 % (Fig. 3a, Table 1). A comparison of the patterns obtained from the post-larvae of both species showed uniformity in the composition of the microbiota and clustering with high similarity (cluster I, Cs>88 %). However, cluster II was observed with high similarity (Cs> 83 %), which partly corresponded to the post-larvae of C. gigas (Fig. 3a). N. Trabal et al. Table 1 Summary of Dice similarity coefficients (percent) of the RFLP profiles of the bacterial microbiota associated with oysters, depending on the growth stage and two grow-out cultivation sites Species Growth stage Site H C. corteziensis PB BT 87 75 81 55 Juvenile 53 57 Adult Post-larvae 56 68 80 40 55 40 Post-larvae 92 Juvenile Adult C. gigas Both speciesa Post-larvae Juvenile Adult 72 88 H hatchery production of post-larvae at Playa Eréndira, PB grow-out cultivation at Punta Botella, BT grow-out cultivation at Bahía Topolobampo a Comparison of RFLP profiles of C. gigas and C. corteziensis The PCA statistical method was used to compare the bacterial microbiota variation in the three growth phases (post-larvae, juvenile, and adult) of C. gigas and C. corteziensis at different grow-out cultivation sites, and the results are presented as a tridimensional scatter plot in Fig. 4. The results for the microbiota associated with the post-larvae in both species of oyster are shown in Fig. 4a. The observed post-larval cluster of C. gigas and C. corteziensis was explained by three principal components (PC1 042.4 % and l036.9, PC2019.8 % and l017.1, PC3010.2 % and l08.8) with a 72.4 % cumulative variance. However, a separated group of C. gigas post-larvae was also observed, which agrees with the results of the overall cluster analysis of RFLP patterns of post-larvae (Fig. 3a). Variation in the Bacterial Microbiota of Juveniles and Adults in Different Grow-Out Cultivation Sites The microbiota composition in juveniles and adults of both species at the two sites was studied by analyzing the patterns obtained by RFLP (Table 1). The profiles obtained from juvenile C. corteziensis indicated differences in the microbiota composition when these oysters were raised at different sites (Fig. 3b). Two groups were defined on the basis of the RFLP profiles, which correlated with the C. corteziensis grow-out site (Punta Botella cluster and Bahía Topolobampo cluster). For the population at Bahía Topolobampo, the Cs was >82 %, whereas at Punta Botella, the Cs was >87 %, which indicates uniformity of the microbiota associated with each locality with the exception of oyster JC6 (Fig. 3b, Table 1). For the C. corteziensis adults, the similarity between samples was higher at PB (Cs>75 %) than at BT (Cs>55 %) (Table 1). Bacterial Microbiota in Two Commercial Oysters 561 Figure 3 Dendrogram representing the similarity (Dice coefficient) observed in the RFLP profiles of rRNA genes amplified from bacterial microbiota associated with oysters. Only representatives of each cluster are shown. a Comparisons between postlarvae C. gigas and C. corteziensis, and b comparisons between juvenile samples of C. corteziensis at two grow-out cultivation sites. pL0postlarvae; J0juvenile; G0C. gigas; C0C. corteziensis; PB0cultivation at Punta Botella; BT0cultivation at Bahía Topolobampo When this analysis was performed for juveniles and adults of C. gigas, differences in the microbiota profiles of individual samples were observed. In contrast to the results for C. corteziensis, the dendrogram based on the Dice coefficient did not reveal an association between the microbiota profile and the grow-out cultivation location (data not shown). Juveniles and adults of C. gigas had lower Cs values than C. corteziensis, which indicates variation within the microbiota among individual oysters of C. gigas (Table 1).The similarity in the microbiota associated with juvenile or adult C. gigas and C. corteziensis was always higher at PB than at BT (Table 1). The PCA of juveniles and adults of C. corteziensis (Fig. 4b) generated two clusters in relation to the grow-out cultivation sites with a 73.6 % cumulative variance (PC1035.0 % and l030.4, PC2023.2 % l020.1, PC3015.4 % l013.4). These results confirmed the pattern observed in the dendrogram estimated from the same samples (Fig. 3b) and the results found by Dice coefficient analysis (Table 1). In contrast, the PCA for juveniles and adults of C. gigas showed minor 562 N. Trabal et al. Figure 4 A 3D scatter plot of the principal component (PC) analysis based on the RFLP of the rRNA gene fragments of C. gigas and C. corteziensis in three growth stages and two grow-out cultivation sites. a RFLP banding patterns of post-larvae C. corteziensis (filled circle) vs. C. gigas (empty circle). b RFLP banding patterns of juveniles and adults of C. corteziensis in two grow-out cultivation sites. Punta Botella (filled triangle) and Bahia Topolobampo (empty triangle). c RFLP banding patterns of juveniles and adults of C. gigas in two grow-out cultivation sites. Punta Botella (filled triangle) and Bahia Topolobampo (empty triangle). d RFLP banding patterns of juveniles of C. corteziensis (filled triangle) vs. C. gigas from Punta Botella (filled diamond), juveniles of C. corteziensis (empty triangle) vs. C. gigas (empty diamond) from Bahia Topolobampo correlations between the RFLP profiles at the two different grow-out cultivation sites (Fig. 4c). The results were similar to those obtained with the Dice coefficient (Table 1). The PCA for juveniles of C. gigas and C. corteziensis in the same cultivation site (Fig. 4d) showed a clear dispersion of the data, thereby supporting the results obtained with the Dice coefficient analysis (Table 1), which did not suggest any similarities in the microbiota associated with these oyster species. Analysis of the Stability of the Microbiota Associated with the Oysters When the Post-larvae are Planted at the Grow-Out Cultivation Sites Using RFLP profiles, the effect of cultivation site transfer upon microbiota associated with the post-larvae was studied (Table 2). Strong similarities (Cs >80 %) in the RFLP profiles of post-larvae and juveniles of C. corteziensis were Bacterial Microbiota in Two Commercial Oysters 563 Table 2 Summary of Dice similarity coefficients (percent) of RFLP profiles of microbiota associated with post-larvae during planting versus juveniles at two grow-out cultivation sites Site Species Growth stages C. corteziensis Post-larvaea C. gigas Post-larvaea a PB Juvenile BT Juvenile 80 40 54 65 Post-larvae planting grow-out sites larvae at the grow-out site (Fig. 5). The PCA showed significant correlations between the RFLP profiles for postlarvae and the juveniles of C. corteziensis at Punta Botella (PC1037.2 %, l032.4, PC2019.1 % l016.6, PC3016.2 % l014.1); this correlation did not exist in post-larvae planted at Bahía Topolobampo (Fig. 5a). These results confirm the findings from the dendrogram analysis using the Dice coefficient (Table 2). The PCA for C. gigas showed considerable variation between the profiles obtained for the post-larvae and the juveniles at both cultivation sites (Fig. 5b). PB cultivation at Punta Botella, BT cultivation at Bahía Topolobampo Identification of Dominant Bacterial Composition Based on TGGE Band Sequencing found at Punta Botella, which suggests that the microbiota composition was stable with little environmental influence (Table 2). The microbiota of C. corteziensis in the Punta Botella cultivation site remained stable, even throughout the fattening process, as the RFLP profiles between juveniles and adults was also similar (Cs>84 %, data not shown). However, upon introduction into Bahía Topolobampo, the microbiota of C. corteziensis was not observed to be stable (Table 2). Similar Dice values (Cs >40 % at PB, Cs >65 % at BT) showed that the microbiota of C. gigas undergo variations during the planting of the post-larvae (Table 2). Using PCA, we studied the variation in the microbiota associated with the oysters when we introduced the post- The sequences of the bands detected visually by TGGE were identified for both oyster species in different growth stages and at the two cultivation sites. The partial sequences of the 16S rRNA gene (400–600 bp) were compared with sequences available in RDP II and GenBank (Table 3). Our results indicate that the microorganisms represented by the main bands (OTUs) belong to the phyla Proteobacteria and Firmicutes. Specifically, these OTUs were related to gramnegative bacteria from the genera Burkhordelia and Borrelia and to gram-positive bacteria such as Bacillus and Propionibacterium. Unique bands corresponding to genera Ruegeria, Paracoccus, Methylobacterium, and Vibrio were also observed. The most frequent OTU in both species was Figure 5 A 3D scatter plot of principal component (PC) analysis based on the RFLP analysis of rRNA gene fragments of C. gigas and C. corteziensis of post-larvae vs. juveniles in two grow-out cultivation sites. a RFLP banding patterns of C. corteziensis post-larvae (filled circle) vs. juveniles from Punta Botella (filled triangle) and, Bahía Topolobampo (empty triangle). b RFLP banding patterns of C. gigas post-larvae (empty circle) vs. juveniles from Punta Botella (filled diamond) and Bahía Topolobampo (empty diamond) 564 N. Trabal et al. Table 3 Nearest-match identification of 16S rRNA gene sequences obtained by PCR-TGGE from known bacterial sequences in the RDP II and GenBank databases Accession number OTU Class affiliation Identity (%) Closest relative Growth stage N/Total Sample Site Burkholderia cepacia Post-larvae 8/8 H Borrelia sp. Juvenile 4/7 PB Crassostrea corteziensis β-Proteobacteria JF522209 B2 JF522195 JF522210 A1 B1 JF522206 JC13E JF522205 JF522208 JC13F JC6A JF522207 JC13A β-Proteobacteria 100 Burkholderia cepacia Juvenile 7/7 PB JF522232 AC9B β-Proteobacteria 100 Burkholderia cepacia Adult 6/6 PB JF522231 JF522203 AC9F JC24C Actinobacteria 99 Propionibacterium sp. Juvenile 2/7 BT JF522204 JC24 A JF522202 JC31A JF522201 JC33 A β-Proteobacteria 100 Burkholderia cepacia Juvenile 7/7 BT JF522200 JF522229 JF522230 JC33B AC28B AC28C β-Proteobacteria 100 Burkholderia cepacia Adult 7/7 BT JF522228 Crasssotrea gigas JF522197 Spirochaetes 100 85.4 AC29A SG2D Bacilli Bacillus subtilis Post-larvae 3/8 H JF522213 JF522196 SG22B SG30G β-Proteobacteria α-Proteobacteria 100 98 89 Burkholderia cepacia Ruegeria sp. Post-larvae Post-larvae 8/8 1/8 H H JF522199 JF522198 JF522227 JG5C JG6F JG9A Bacilli α-Proteobacteria β-Proteobacteria 96 98 100 Bacillus sp. Paracoccus sp. Burkholderia cepacia Juvenile Juvenile Juvenile 4/7 1/7 7/7 PB PB PB JF522226 JF522223 JF522220 AG9B AG9F AG10B JF522218 JF522217 JF522216 AG12A AG12P AG14P β-Proteobacteria 100 Burkholderia cepacia Adult 6/6 PB JF522225 JF522222 JF522219 AG9D AG9I AG11I α-Proteobacteria 100 Methylobacterium sp. Adult 1/6 PB Bacilli 100 Bacillus subtilis Adult 2/6 PB JF522215 JF522214 JF522211 JF522212 JG22A AG22C AG30B AG26H β-Proteobacteria 100 Burkholderia cepacia Juvenile 6/6 BT β-Proteobacteria γ-Proteobacteria 100 100 Burkholderia cepacia Vibrio sp. Adult Adult 6/6 1/6 BT BT N Number of oysters presenting the closest relative OTU operational taxonomic unit/number total of oyster samples, H hatchery production of post-larvae at Playa Eréndira, PB cultivation at Punta Botella, BT cultivation at Bahía Topolobampo Burkholderia cepacia, which was found during all growth stages and at both sites. A particular bacterium (probably Bacillus subtilis or another in the genus Bacillus) was a component of C. gigas throughout its growth at Punta Botella. In most cases, the bacterial diversity was higher in C. gigas than in C. corteziensis, especially in adults (Table y3). Discussion The composition of microbiota associated with oysters can be influenced by numerous factors, including particular characteristics of the host, diet, and environmental conditions. In this study, bacterial microbiota of depurated C. Bacterial Microbiota in Two Commercial Oysters gigas and C. corteziensis oysters were compared using the RFLP molecular technique to analyze the microbiota variation in three growth stages of the oysters and at two growout cultivation sites. In parallel, the microbiota composition in these two species of oysters was also determined using the TGGE method. At the post-larval stage, the microbiota presented a uniform composition that was independent of the host (C. gigas or C. corteziensis) with similar RFLP profiles and formed a group when analyzed by PCA. During the cultivation of the larvae at the hatchery, the bacteria associated with the food, which influence the composition of the microbiota, were controlled in the laboratory [4, 5, 26, 31, 74]. All postlarvae were produced in the same hatchery, where they were fed a mixture of microalgae. Generally, the microalgae cultivated in shellfish hatcheries are not free of bacteria and are a highly nutritive product, that is, notably useful as oyster food [38, 39]. Thus, when oysters are fed microalgae, the associated bacteria may colonize the gastrointestinal tract and become part of the microbiota of the oyster species. It has been reported that bivalve mollusks, which have similar diets, provide similar gastrointestinal environments for bacterial colonization [4, 27, 31, 66]; this condition may explain the similarity found in the composition of the microbiota in C. gigas and C. corteziensis. There is evidence that certain aquatic invertebrates, particularly oysters, maintain a permanent microbiota in the gastrointestinal tract [31, 33, 40, 62, 69, 74]. Several studies have revealed that the bacterial populations isolated from the gut differ in species composition from those isolated from the habitat [31, 40, 62, 63]. Other studies have indicated that ambient conditions significantly affect the microbiota in marine invertebrates; oysters of the same species caught in the wild and laboratory-reared animals may have different bacterial communities in the gastrointestinal tract [31, 40, 62, 75]. During the cultivation of marine organisms, changes in environmental parameters, such as salinity, temperature [17, 34], nutrients, availability of food, and operational design in grow-out and management methods, may induce stress in the animals [38] and influence the composition of the gastrointestinal microbiota. Our results show that C. corteziensis microbiota composition was influenced by the conditions presented at the cultivation site. The results obtained by RFLP profile analysis and PCA show that the associated microbiota vary depending on where C. corteziensis was fattened (differences were observed between Punta Botella and Bahía Topolobampo). It is noteworthy that the microbiota composition of C. corteziensis remained stable from the post-larvae stage to the growth of the oyster when the oysters were planted at Punta Botella, which was not observed at Bahía Topolobampo. This result was somewhat surprising because the subtropical temperature observed at Bahía Topolobampo promotes better 565 growth in C. corteziensis [13, 47, 48]. There is limited data about the environmental parameters of the field site used in this study. Thus, we opted to report the temperature ranges, but these data are not sufficient to establish a relationship with the microbiota composition. It will be interesting to evaluate the impact of environmental parameters on microbiota composition. The differences found in the stability of the microbiota in C. corteziensis at Punta Botella and Bahía Topolobampo may be the result of the characteristics of each cultivation site. In particular, the environmental conditions and grow-out management at Punta Botella reflect low pollution and little anthropogenic activity, which support the stability of the associated microbiota. In contrast, Bahía Topolobampo is a highly impacted site, where the lagoon water is contaminated by agricultural runoff and urban and industrial sewage effluents [57, 58]. The difference in the filtering capacity of the two species is another factor that may explain the differences in microbiota [31, 62]. C. gigas has a higher filtration capacity than native oysters, and the capacity increases further when food is abundant; thus these oysters are typically larger in size [11, 22]. A difference in filtration capacity might explain the greater diversity of bacteria in C. gigas because this oyster has a greater capacity to ingest bacteria during respiration and feeding. Despite the differences in behavior among microbiota associated with oysters in different grow-out sites, we found a common bacterial component associated with C. corteziensis and C. gigas. This bacterium was a member of the Burkholderia genus, was acquired in the post-larval stage, and remained associated with the gastrointestinal tract in juvenile and adult oysters at both cultivation sites. Presumably, the changes during the different growth stages of the oysters result in changes in the environment of the microbiota associated with the gastrointestinal tract [8, 31, 37, 62]. Changes in the composition of the microbiota were mainly observed in C. gigas in the three growing phases included in this study and at the two cultivation sites. These changes are probably related to a particular feature of the host [31]. However, our results show that at least part of the microbiota remains strongly associated with the gastrointestinal tracts of C. corteziensis and C. gigas, presumably because of attributes of the bacteria (such as adhesion to the gut wall) that prevent expulsion from the gut. In a previous report, B. cepacia was found in the three growth stages and was not influenced by the conditions at the cultivation site, which could indicate a symbiotic association between this bacterium and the host (i.e., the two oyster species). It would be interesting to demonstrate positive relationships between these host oysters and B. cepacia and to determine whether the bacteria play an important role in the physiological processes of C. gigas and C. corteziensis. The genus Burkholderia was originally classified with 566 Pseudomonas [44, 47, 78]. In the past two decades, Burkholderia has been found in a variety of ecological niches, including soil [21], plants [14, 44, 71], and freshwater and marine habitats [16]. Recently, a strain was isolated from C. corteziensis and identified as B. cepacia. This strain had been isolated from the digestive gland of native C. corteziensis from the State of Nayarit, Mexico [9, 10]. Another common bacterium in the post-larvae of C. gigas was Bacillus sp., which was also present in juveniles and adults at Punta Botella. Bacillus spp. have been reported in oysters, especially in Saccrostrea glomerata [29] and Ostrea edulis [63]. Hernández and Olmos [33] reported findings similar to ours, and they explained that Bacillus spp. adheres to the gills of C. gigas at the cultivation site in the west coast of Mexico. It has been reported that some species of Bacillus may have potential as a probiotic in cultivated marine animals because it can colonize marine organisms and have an antagonistic effect on pathogenic microorganisms. In addition, the spores of these bacteria can be easily introduced into the food supply of the oysters [26, 52]. Several OTUs were found in C. corteziensis sampled at the Punta Botella cultivation site and were recognized as bacteria of the family Spirochaetaceae. The biodiversity of spirochetes is high, and they are present in large numbers in the gastrointestinal tract of humans, mammals, insects, and bivalves (especially oysters), as well as in aquatic and marine environments [45, 60, 73]. These bacteria seem to be limited to the crystalline styles (mucoproteinaceous rod of the digestive gland) [45, 73], but the hypothesis that styleassociated spirochetes play a major role in the extracellular enzymatic digestion processes of bivalves has not been tested [35]. Recently, Husmann et al. [35] investigated the phylogeny of spirochete groups present in crystalline styles from select bivalves at different habitats and found that spirochetes are not randomly distributed, which implies an association between oyster species and specific spirochete clusters. However, the spirochetes were present in only a portion of the crystalline styles investigated; therefore, the authors concluded that spirochetes are not essential symbionts for these bivalves. All of the identified spirochetes were grouped into two families: Spirochaetaceae, which included two genera (Cristispira and Spirochaeta), and Brachyspiraceae, with one genus (Brachyspira) [35]. Similarity analyses of the OTU found in this study using databases and GenBank RPDII revealed an 85 % sequence similarity with Borrelia spp. This result is similar to the results of Green & Barnes [29] for microbiota in the digestive gland of the Sydney rock oyster. In this study, we demonstrate that molecular biology techniques may clarify differences in bacterial species associated with an oyster species, as most spirochetes cannot be cultured. The α-, β-, and γ-Proteobacteria, Firmicutes, Spirochaetes, and Actinobacteria species that we identified N. Trabal et al. among the microbiota associated with C. gigas or C. corteziensis have been reported in other oysters using cultureindependent techniques [33, 40, 54, 63, 69]. However, our results showed low bacterial diversity compared with other studies. We used TGGE methodology to analyze the bacterial community composition in the oysters. To rule out the possibility that the observed lack of diversity may be the result of primer bias, the 16S rRNA primers were analyzed using the informatics tool Probe Match in RDP II, which contains almost 2 million sequences that are distributed in 33 phyla. Each primer matched with over 80 % of the total sequences deposited in RDP II, recognizing all phyla. More than 70 % of the sequences assigned in each phylum were recognized by these primers. Therefore, no specific preference during amplification should be expected, and the narrow diversity is therefore not a consequence of primer bias. To reduce the nested PCR bias, the first amplification step was limited to 15 cycles. For TGGE analysis, we used a previously established protocol [55]. This protocol includes the use of known sequences (%GC content) as a control for each run and checks the separation of bands with different GC contents. Regardless of the growth stage or the cultivation site of oysters, we found low bacterial diversity and this may be due to the use of depurated oysters. With the exception of the study by Romero et al. [69], the reports on microbiota associated with other oysters, do not distinguish between resident and transient bacteria. This could lead to an overestimation of bacterial diversity within the microbiota. The Vibrio and Pseudomonas species are commonly isolated from oyster tissues using traditional methods [31, 54, 62, 63, 75, 77] and are recognized as symbionts of diverse marine invertebrates [32]. In general, the Vibrios species have been reported in sick oysters and have been detected using selective culturing [17, 59, 75, 76]. However, we did not find Vibrios in this study (only one OTU in an adult C. gigas). It has been demonstrated that the distribution of Vibrios is typically seasonal with peaks of abundance that depend on environmental conditions, similar to mollusks [17, 32, 75, 76]. However, although the juveniles and adults of C. corteziensis and C. gigas were collected in warm water, which has a positive effect on the abundance of Vibrios, these bacteria were not found. In recent years, some authors have suggested that Vibrio is not necessarily predominant in the microbiota of healthy oysters [29, 42]. We analyzed samples from healthy and depurated oysters, which may explain why we did not find Vibrios to be a component of the microbiota of C. gigas and C. corteziensis. It would be interesting in a future study to use Vibriospecific primers to verify our results. This study provides the first description of the bacterial community in Crassostrea corteziensis. Additionally, this study is the first attempt to describe what happens to the Bacterial Microbiota in Two Commercial Oysters stability of the microbiota when the post-larvae are planted at grow-out cultivation sites. Considering that one of the most critical steps in the cultivation of these oysters is the planting of post-larvae (spat) at the cultivation site in the natural environment in which large numbers of oyster die [7, 11, 18], the studies described here are probably important. We observed that during the early growth stages, certain bacteria (especially B. cepacia) colonize oysters and remain strongly associated with their host, probably in a symbiotic host–bacteria relationship. This association is maintained during the growth phases of the oysters and is not altered by environmental conditions or the management of the organisms at the grow-out site. This finding is important because probiotics have been used in aquaculture to control pathogenic microorganisms by competitive exclusion and may help to reduce and prevent outbreaks in several marine species [26–28, 51, 61]. Resident bacteria with potential probiotic characteristics that strongly associate with the oysters seem to be stable in the host because the community was not affected by the conditions of the grow-out site, which suggests a competitive advantage for the sustainable development of aquaculture. The main findings of the present study were (1) the bacterial communities associated with the oysters, especially C. corteziensis, at each grow-out site were similar, if not identical, and were influenced by the environment. (2) The microbiota associated with the oysters underwent certain changes when the post-larvae were planted at grow-out sites and during their growth from juvenile to adult. (3) However, B. cepacia was established in the post-larval phase and remained associated with the oysters throughout their growth, regardless of the grow-out site. (4) There were certain differences among the microbiota associated with C. gigas and C. corteziensis, even when grown at the same site. (5) C. gigas has more intraspecific differences in its composition of microbiota. 567 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. Acknowledgments We thank Acuícola Robles and Acuícola CuateMachado for providing the oysters used in this research. We also thank Victoria Urzúa (INTA), Alejandra Givovich (INTA), Angel Carrillo García (CIBNOR), and Hever Latisnere-Barragan of Marine Biotechnology Laboratory (CIBNOR) for technical support. Funding was provided by Consejo Nacional de Ciencia y Tecnología of Mexico (SEP-CONACYT grants 129025 and 106887). N. A. is a recipient of a CONACYT doctoral fellowship and also an internship grant at the Instituto Nacional de Tecnología de los Alimentos (Universidad de Chile). 14. 15. 16. 17. References 18. 1. Acosta-Velázquez J, Ruíz-Luna A (2007) Variación en la cobertura, distribución y estructura de los manglares del complejo lagunar Bahía Magdalena-Bahía Almejas (1990–2005). In: FunesRodríguez R, Gómez-Gutiérrez J, Palomares-García R (eds) Estudios ecológicos en Bahía Magdalena, CICIMAR [Centro 19. Interdiciplinario de Ciencias Marinas]-IPN [Instituto Politécnico Nacional], La Paz. B.C.S, Mexico, pp 127–141 Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 17:3389–3402 Austin B (2006) The bacterial microflora of fish, revised. Sci World J 6:931–945 Avendaño R, Riquelme C (1999) Establishment of mixed-culture probiotics and microalgae as food for bivalve larvae. Aquacul Res 30:893–900 Avendaño R, Riquelme C, Escribano R, Reyes N (2001) Postlarval survival and growth of Argopecten purpuratus (Lamarck, 1819) in Bahia Inglesa, Chile: effects of origin, distribution in the bay and larval bacterioflora. Rev Chilena Hist Nat 74:669–679 Berg R (1996) The indigenous gastrointestinal microflora. Trends Microbiol 4:430–435 Bower SM, McGladdery SE (2003) Disease interactions between wild and cultured shellfish. In: Fisheries and Oceans Canada (ed) A scientific review of the potential environmental effects of aquaculture in aquatic ecosystems. Can Tech Rep Fish Aquat Sci 2:1–5 Brown C (1973) The effects of some selected bacteria on embryos and larvae of the American oyster Crassostrea virginica. J Invertebr Pathol 21:215–233 Campa-Córdova AI, González-Ocampo H, Luna-González A, Mazón-Suástegui JM, Ascencio F (2009) Growth, survival, and superoxide dismutase activity in juvenile Crassostrea corteziensis (Hertlein, 1951) treated with probiotics. Hidrobiologia 19:151–157 Campa-Córdova AI, Luna-González A, Mazón-Suástegui JM, Aguirre-Guzmán G, Ascencio-Valle F, González-Ocampo HA (2011) Efecto de bacterias probióticas en el cultivo larvario del ostión de placer Crassostrea corteziensis (Bivalvia: Ostreidae). Int J Trop Biol 59:183–191 Castillo-Durán A, Chávez-Villalba J, Arreola-Lizárraga A, Barraza-Guardado R (2010) Comparative growth, condition, and survival of juvenile Crassostrea gigas and C. corteziensis oysters cultivated in summer and winter. Cienc Mar 36:29–39 Chávez S (2006) El papel de los manglares en la producción de las comunidades acuáticas de Bahía Magdalena, BCS. In: FunesRodríguez R, Gómez-Gutiérrez J, Palomares-García R (eds) Estudios ecológicos en Bahía Magdalena, CICIMAR [Centro Interdiciplinario de Ciencias Marinas]-IPN [Instituto Politécnico Nacional], La Paz. B.C.S, Mexico, pp 127–141 Chávez-Villalba J, López-Tapia M, Mazón-Suástegui JM, RoblesMungaray M (2005) Growth of the oyster Crassostrea corteziensis (Hertlein, 1951) in Sonora, Mexico. Aquacul Res 36:1337–1344 Coenye T, Vandamme P (2003) Diversity and significance of Burkholderia species occupying diverse ecological niches. Environ Microbiol 5:719–729 Cole J, Chai B, Farris R, Wang Q, Kulam S, McGarrell D, Garrity G, Tiedje J (2005) The Ribosomal Database Project (RDP-II): sequences and tools for high-throughput rRNA analysis. Nucleic Acids Res 33:294–296 Compant S, Nowak J, Coenye T, Clément C, Barka EA (2008) Diversity and occurrence of Burkholderia spp. in the natural environment. FEMS Microbiol Rev 32:607–626 DePaola A, Nordstrom JL, Bowers JC, Wells JG, Cook DW (2003) Seasonal abundance of total and pathogenic Vibrio parahaemolyticus in Alabama oysters. Appl Environ Microbiol 69:1521–1526 Eckmaye W (1983) Growth and survival of hatchery-reared American oysters set on three types of cultch and in Bon Secour Bay, Alabama. North Amer J Fish Manage 3:171–175 Erasmus JH, Cook PA, Coyne VE (1997) The role of bacteria in the digestion of seaweed by the abalone Haliotis midae. Aquaculture 155:377–386 568 20. Espejo RT, Escanilla D (1993) Detection of HIV1 DNA by a simple procedure of polymerase chain reaction, using “primerdimer” formation as an internal control of amplification. Res Virol 144:243–246 21. Estrada-de Los Santos P, Bustillos-Cristales R, Caballero-Mellado J (2001) Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Appl Environ Microbiol 67:2790–2798 22. Farías A (2008) Nutrición y alimentación en moluscos bivalvos. In: Lovatelli A, Farías, Uriarte I (eds). Estado actual del cultivo y manejo de moluscos bivalvos y su proyección futura: factores que afectan su sustentabilidad en América Latina, FAO Actas de Pesca y Acuicultura, No. 12., Rome, pp 297–308 23. FAO (2009) El estado mundial de la pesca y la acuicultura 2008. Food and Agriculture Organization, Rome, pp 36–39 24. Food and Drug Administration (1992) Scheduled Depuration Process. National Shellfish Sanitation Program. 25. Fromin N, Hamelin J, Tarnawski S, Roesti D, Jourdain-Miserez K, Forestier N, Teyssier-Cuvelle S, Gillet F, Aragno M, Rossi P (2002) Statistical analysis of denaturing gel electrophoresis (DGE) fingerprinting patterns. Environ Microbiol 4:634–643 26. Gatesoupe FJ (1999) The use of probiotics in aquaculture. Aquaculture 180:147–165 27. Gibson LF, Woodworth J, George AM (1998) Probiotic activity of Aeromonas media on the Pacific oyster, Crassostrea gigas, when challenged with Vibrio tubiashii. Aquaculture 169:111–120 28. Gómez-Gil B, Roque A, Turnbull JF (2000) The use and selection of probiotic bacteria in the larval culture of aquatic organisms. Aquaculture 191:259–270 29. Green TJ, Barnes AC (2010) Bacterial diversity of the digestive gland of Sydney rock oysters, Saccostrea glomerata, infected with the paramyxean parasite, Marteilia Sydney. J Appl Microbiol 109:613–622 30. Guo X, Yongping W, Lingling W, Lee JH (2008) Genome mapping and genomics in oyster. In: Kole C, Kocher TD (eds) Fishes and Aquatic Animals. Springer, Berlin, pp 23–36 31. Harris JM (1993) The presence nature, and role of gut microflora in aquatic invertebrates: a synthesis. Microb Ecol 25:195–231 32. Hazen TH, Pan L, Gu JD, Sobecky PA (2010) The contribution of mobile genetic elements to the evolution and ecology of Vibrios. FEMS Microbiol Ecol 74:485–499 33. Hernández-Zárate G, Olmos-Soto J (2006) Identification of bacterial diversity in the oyster Crassostrea gigas by fluorescent in situ hybridization and polymerase chain reaction. J Appl Microbiol 100:664–667 34. Hofmann E, Ford S, Powell E, Klinck J (2001) Modeling studies of the effect of climate variability on MSX disease in eastern oyster (Crassostrea virginica) populations. Hydrobiologia 460:195–212 35. Husmann G, Gerdts G, Wichels A (2010) Spirochetes in crystalline styles of marine bivalves: group-specific PCR detection and 16 s rRNA sequence analysis. J Shellfish Res 29:1069–1075 36. Krzanowski WJ (2000) Principles of multivariate analysis. A user's perspective. Oxford University Press, Oxford 37. Kueh C, Chan K (1985) Bacteria in bivalve shellfish with special reference to the oysters. J Appl Bacteriol 59:41–47 38. Lacoste A, Jalabert F, Malham SK, Cueff A, Poulet S (2001) Stress and stress-induced neuroendocrine changes increase the susceptibility of juvenile oysters (Crassostrea gigas) to Vibrio splendidus. Appl Environ Microbiol 67:2304–2309 39. Lafferty KD, Porter JW, Ford SE (2004) Are diseases increasing in the ocean? Annu Rev Ecol Evol Syst 35:31–54 40. LaValley KJ, Steve Jones LA, Gómez-Chiarri M, Dealteris J, Rice M (2009) Bacterial community profiling of the eastern oyster (Crassostrea virginica): comparison of culture-dependent and culture-independent outcomes. J Shellfish Res 28:827–835 N. Trabal et al. 41. Lee R, Lovatelli T, Ababouch A (2008). Bivalve depuration: fundamental and practical aspects. FAO, Fisheries Technical Paper, Food and Agriculture Organization, Rome, No 551: pp 11–39 42. Li Q, Zhang Y, Juck D, Fortin N, Greer CW, Tang Q (2010) Phylogenetic analysis of bacterial communities in the shrimp and sea cucumber aquaculture environment in northern China by culturing and PCR–DGGE. Aquacult Int 18:977–990 43. Liesack W, Weyland H, Stackebrandt E (1991) Potential risks of gene amplification by PCR as determined by 16S rDNA analysis of a mixed culture of strict barophilic bacteria. Microb Ecol 21:191–198 44. Mahenthiralingam E, Baldwin A, Dowson CG (2008) Burkholderia cepacia complex bacteria: opportunistic pathogens with important natural biology. J Appl Microbiol 104:1539–1551 45. Margulis L, Nault L, Sieburth J (1991) Cristispira from oyster styles: complex morphology of large symbiotic spirochetes. Symbiosis 11:1–19 46. Marques A, Ollevier F, Verstraete W, Sorgeloos P, Bossier P (2006) Gnotobiotically grown aquatic animals: opportunities to investigate host-microbe interactions. J Appl Microbiol 100:903–918 47. Mazón-Suástegui JM, Parres-Haro MA, Ruíz-Ruíz KM, RodríguezJaramillo MC, Saucedo PE (2009) Influence of hatchery diets on early grow-out of the Cortez oyster Crassostrea corteziensis in Guasave, Sinaloa, Mexico. Aqua Res 40:1908–1914 48. Mazón-Suástegui JM, Ruíz-García MC, Chávez-Villalba J, RodríguezJaramillo C, Saucedo PE (2011) Analysis of growth and first reproduction of hatchery-reared juvenile Cortez oyster (Crassostrea corteziensis) in northwestern Mexico: proposal of a minimal fishing size. Aqua Res 42:1–11 49. Mazón-Suástegui JM, Ruiz-Ruiz KM, Parres-Haro A, Saucedo P (2008) Combined effects of diet and stocking density on growth and biochemical composition of seed of the Cortez oyster Crassostrea corteziensis at the hatchery. Aquaculture 284:98–105 50. McCracken VJ, Simpson JM, Mackle RI, Gaskins HR (2001) Molecular ecological analysis of dietary and antibiotic-induced alterations of the mouse intestinal microbiota. J Nutrition 131:1862–1870 51. Moriarty DJW (1990) Interactions of microorganisms and aquatic animals, particularly the nutritional role of the gut flora. In: Lésel R (ed) Microbiology in Poecilotherms. Elsevier, Paris 52. Moriarty DJW (1997) The role of microorganisms in aquaculture ponds. Aquaculture 151:333–349 53. Muyzer G, Dewaal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for rRNA. Appl Environ Microbiol 59:695–700 54. Najiah M, Nadirah M, Lee KL, Lee SW, Wendy W, Ruhil HH, Nurul FA (2008) Bacteria flora and heavy metals in cultivated oysters Crassostrea iredalei of Setiu Wetland, East Coast Peninsular Malaysia. Vet Res Commun 32:377–381 55. Navarrete P, Espejo RT, Romero J (2009) Molecular analysis of microbiota along the digestive tract of juvenile Atlantic salmon (Salmon salar L.). Microb Ecol 57:550–561 56. Nayak SK (2010) Role of gastrointestinal microbiota in fish. Aquac Res 41:1553–1573 57. Osuna Flores I, Riva MC (2002) Organochlorine pesticide residue concentrations in shrimps, sediments, and surface water from Bay of Ohuira, Topolobampo, Sinaloa, Mexico. Bull Environ Contam Toxicol 68:532–539 58. Páez-Osuna F, Ruiz-Fernández JI, Botello AV, Ponce-Velez G, Osuna-López JI, Frías-Espirucueta MG, López-López G, Zazueta-Padilla HM (2002) Concentrations of selected trace metals (Cu, Pb, Zn), organochlorines (PCBs, HCB) and total PAHs in mangrove oysters from the Pacific coast of Mexico: an overview. Mar Pollut Bull 44:1303–1308 Bacterial Microbiota in Two Commercial Oysters 59. Paillard C, Le Roux F, Borrego JJ (2004) Bacterial disease in marine bivalves, a review of recent studies: trends and evolution. Aquat Living Resour 17:477–498 60. Paster BJ, Dewhirst FE (2000) Phylogenetic foundation of spirochetes. J Mol Microbiol Biotechnol 4:341–344 61. Playne MJ (1995) Probiotic microorganisms. Rec Adv Microbiol 3:215–254 62. Prieur D, Mvel G, Nicolas J-L, Plusquellec A, Vigneulle M (1990) Interactions between bivalve molluscs and bacteria in the marine environment. Oceanogr Mar Biol Annu Rev 28:277–352 63. Pujalte MJ, Ortigosa M, Macián MC, Garay E (1999) Aerobic and facultative anaerobic heterotrophic bacteria associated to Mediterranean oysters and seawater. Int Microbiol 2:259–266 64. Rawls JF, Samuel BS, Gordon JI (2004) Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proc Natl Acad Sci U S A 13:4596–4601 65. Renault T, Cochennec N (1995) Chlamydia-like organisms in ctenidia and mantle cells of the Japanese oyster Crassostrea gigas from the French Atlantic coast. Dis Aquat Org 23:153–159 66. Riquelme CE, Avendaño RE (2003) Microalgae and bacteria interaction in the aquatic environment and their potential use in aquaculture. Rev Chil Hist Nat 76:725–736 67. Rippey SR (1994) Infectious diseases associated with molluscan shellfish consumption. Clin Microbiol Rev 7:419–425 68. Romalde JL, Barja JL (2010) Bacteria in molluscs: good and bad guys. In: Current Research, Technology and education topics in Applied Microbiology and Microbial Biotechnology. Mendez – Vila ed, Spain. pp:136-147 69. Romero J, Garcia-Varela M, Laclette JP, Espejo RT (2002) Bacterial 16S rRNA gene analysis revealed that bacteria related to Arcobacter spp. constitute an abundant and common component of the oyster microbiota (Tiostrea chilensis). Microb Ecol 44:365–371 569 70. Rudi K, Skulberg OM, Larsen F, Jacoksen KS (1997) Strain classification of oxyphotobacteria in clone cultures on the basis of 16S rRNA sequences from variable regions V6, V7 and V8. Appl Environ Microbiol 63:2593–99 71. Salles JF, Van Veen JA, Van Elsas JD (2004) Multivariate analyses of Burkholderia species in soil: effect of crop and land use history. Appl Environ Microbiol 70:4012–4020 72. Son TH, Fleet GH (1980) Behavior of pathogenic bacteria in the oyster, Crassostrea commercialis, during depuration, re-laying, and storage. Appl Environ Microbiol 40:994–1002 73. Tall BD, Nauman RK (1981) Scanning electron microscopy of Cristispira species in Chesapeake Bay oysters. Appl Environ Microbiol 42:336–343 74. Tanaka R, Otsubob M, Sawabec T, Ezurac Y, Tajimac K (2004) Biodiversity and in situ abundance of gut microflora of abalone (Haliotis bulmes) determined by culture-independent techniques. Aquaculture 241:453–463 75. Thompson JR, Marcelino LA, Polz MF (2005) Diversity, sources, and detection of human bacterial pathogens in the marine environment. In: Belkin C (ed) Oceans and health: pathogens in the marine environment. Springer, New York, pp 29–68 76. Thompson JR, Randa MA, Marcelino LA, Tomita-Mitchell A, Lim E, Polz MF (2004) Diversity and dynamics of a north atlantic coastal Vibrio community. Appl Environ Microbiol 70:4103–4110 77. Thompson FL, Tetsuya I, Swings J (2004) Biodiversity of Vibrios. Microbiol Mol Biol Rev 68:403–431 78. Yabuuchi E, Kosako Y, Oyaizu H, Yano I, Hotta H, Hashimoto Y, Ezaki T, Arakawa M (1992) Proposal of Burkholderia gen. nov. and transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. nov. Microbiol Immuno 36:1251–1275