* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download CSCE590/822 Data Mining Principles and Applications
Survey
Document related concepts
Transcript
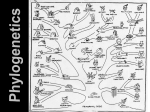
CSCE555 Bioinformatics Lecture 12 Phylogenetics I HAPPY CHINESE NEW YEAR Meeting: MW 4:00PM-5:15PM SWGN2A21 Instructor: Dr. Jianjun Hu Course page: http://www.scigen.org/csce555 University of South Carolina Department of Computer Science and Engineering 2008 www.cse.sc.edu. Outline Introduction to Evolution What is phylogeny and phylogenetics Application of phylogenetics Algorithms for phylogenetic inference 5/24/2017 2 How did life evolve on earth? An international effort to understand how life evolved on earth Biomedical applications: drug design, protein structure and function prediction, biodiversity. Courtesy of the Tree of Life project Evolution Evolution of new organisms is driven by Mutations ◦ The DNA sequence can be changed due to single base changes, deletion/insertion of DNA segments, etc. Selection bias Theory of Evolution Basic idea ◦ speciation events lead to creation of different species. ◦ Speciation caused by physical separation into groups where different genetic variants become dominant Any two species share a (possibly distant) common ancestor Primate evolution A phylogeny is a tree that describes the sequence of speciation events that lead to the forming of a set of current day species; also called a phylogenetic tree. DNA Sequence Evolution -3 mil yrs AAGACTT AAGGCCT AGGGCAT AGGGCAT TAGCCCT TAGCCCA -2 mil yrs TGGACTT TAGACTT AGCACTT AGCACAA AGCGCTT -1 mil yrs today Morphological vs. Molecular Classical phylogenetic analysis: morphological features: number of legs, lengths of legs, etc. Modern biological methods allow to use molecular features ◦ Gene sequences ◦ Protein sequences ◦ Whole genome sequences. E.g. rearrangements Morphological topology (Based on Mc Kenna and Bell, 1997) Bonobo Chimpanzee Man Gorilla Sumatran orangutan Bornean orangutan Common gibbon Barbary ape Baboon White-fronted capuchin Slow loris Tree shrew Japanese pipistrelle Long-tailed bat Jamaican fruit-eating bat Horseshoe bat Little red flying fox Ryukyu flying fox Mouse Rat Vole Cane-rat Guinea pig Squirrel Dormouse Rabbit Pika Pig Hippopotamus Sheep Cow Alpaca Blue whale Fin whale Sperm whale Donkey Horse Indian rhino White rhino Elephant Aardvark Grey seal Harbor seal Dog Cat Asiatic shrew Long-clawed shrew Small Madagascar hedgehog Hedgehog Gymnure Mole Armadillo Bandicoot Wallaroo Opossum Platypus Archonta Glires Ungulata Carnivora Insectivora Xenarthra From sequences to a phylogenetic tree Rat QEPGGLVVPPTDA Rabbit QEPGGMVVPPTDA Gorilla QEPGGLVVPPTDA Cat REPGGLVVPPTEG There are many possible types of sequences to use (e.g. Mitochondrial vs Nuclear proteins). Mitochondrial topology (Based on Pupko et al.,) Donkey Horse Indian rhino White rhino Grey seal Harbor seal Dog Cat Blue whale Fin whale Sperm whale Hippopotamus Sheep Cow Alpaca Pig Little red flying fox Ryukyu flying fox Horseshoe bat Japanese pipistrelle Long-tailed bat Jamaican fruit-eating bat Asiatic shrew Long-clawed shrew Mole Small Madagascar hedgehog Aardvark Elephant Armadillo Rabbit Pika Tree shrew Bonobo Chimpanzee Man Gorilla Sumatran orangutan Bornean orangutan Common gibbon Barbary ape Baboon White-fronted capuchin Slow loris Squirrel Dormouse Cane-rat Guinea pig Mouse Rat Vole Hedgehog Gymnure Bandicoot Wallaroo Opossum Platypus Perissodactyla Carnivora Cetartiodactyla Chiroptera Moles+Shrews Afrotheria Xenarthra Lagomorpha + Scandentia Primates Rodentia 1 Rodentia 2 Hedgehogs Phylogenenetic trees Aardvark Bison Chimp Dog Elephant Leaves - current day species (or taxa – plural of taxon) Internal vertices - hypothetical common ancestors Edges length - “time” from one speciation to the next Types of Trees A natural model to consider is that of rooted trees Common Ancestor Types of trees Unrooted tree represents the same phylogeny without the root node Depending on the model, data from current day species does not distinguish between different placements of the root. Rooted versus unrooted trees Tree a Tree b Tree c b a c Represents the three rooted trees What is phylogenetics? Phylogenetics is the study of evolutionary relationships among and within species. ◦ Inference of trees from data ◦ Interpreting the evolutionary tree ◦ Application of evolutionary trees birds rodents crocodiles marsupials snakes primates lizards What is phylogenetics? crocodiles birds lizards snakes rodents primates marsupials This is an example of a phylogenetic tree. Applications of phylogenetics • Forensics: Did a patient’s HIV infection result from an invasive dental procedure performed by an HIV+ dentist? • Conservation: How much gene flow is there among local populations of island foxes off the coast of California? • Medicine: What are the evolutionary relationships among the various prion-related diseases? HIV case Applications of phylogenetics 1. Forensics Did a patient’s HIV infection result from an invasive dental procedure performed by an HIV+ dentist? Phylogenetic analysis So what do the results mean? • 2 of 3 patients closer to dentist than to local controls. Statistical significance? More powerful analyses? • Do we have enough data to be confident in our conclusions? What additional data would help? • If we determine that the dentist’s virus is linked to those of patients E and G, what are possible interpretations of this pattern? How could we test between them? Applications of phylogenetics 2. Conservation How much gene flow is there among local populations of island foxes off the coast of California? http://bioquest.org/bedrock/ Wayne, K. R, Morin, P.A. 2004 Conservation Genetics in the New Molecular Age, Frontiers in Ecology and the Environment. 2: 89-97. (ESA publication) Applications of phylogenetics 3. Medicine What are the evolutionary relationships among the various prion-related diseases? Inferring Phylogenies Trees can be inferred: ◦ Morphology of the organisms ◦ Sequence comparison Example: Orc: ACAGTGACGCCCCAAACGT Elf: ACAGTGACGCTACAAACGT Dwarf: CCTGTGACGTAACAAACGA Hobbit: CCTGTGACGTAGCAAACGA Human: CCTGTGACGTAGCAAACGA How Many Trees? (assuming bifurcation only) Unrooted trees # # pairwise sequences distances 3 4 5 6 10 30 N # trees # branches /tree Rooted trees # trees # branches /tree How Many Trees? Unrooted trees Rooted trees # sequence s # pairwise distance s 3 3 1 3 3 4 4 6 3 5 15 6 5 10 15 7 105 8 6 15 105 9 945 10 10 45 2,027,025 17 34,459,425 18 30 435 8.69 1036 57 4.95 1038 58 N N (N - 1) 2 # branches /tree # trees (2N - 5)! 2N - 3 (N - 3)! 2N - 3 # branches /tree # trees (2N - 3)! 2N - 2 (N - 2)! 2N - 2 Phylogenetic Methods Many different procedures exist. Three of the most popular: Neighbor-joining • Minimizes distance between nearest neighbors Maximum parsimony • Minimizes total evolutionary change Maximum likelihood • Maximizes likelihood of observed data Comparison of Methods Neighbor-joining Maximum parsimony Maximum likelihood Very fast Slow Very slow Easily trapped in local optima Assumptions fail when Highly dependent on evolution is rapid assumed evolution model Good for generating tentative tree, or choosing among multiple trees Best option when tractable (<30 taxa, strong conservation) Good for very small data sets and for testing trees built using other methods Distance based tree Construction Distance- A weighted tree that realizes the distances between the objects. Given a set of species (leaves in a supposed tree), and distances between them – construct a phylogeny which best “fits” the distances. Distance Matrix Given n species, we can compute the n x n distance matrix Dij Dij may be defined as the edit distance between a gene in species i and species j, where the gene of interest is sequenced for all n species. Distances in Trees Edges may have weights reflecting: ◦ Number of mutations on evolutionary path from one species to another ◦ Time estimate for evolution of one species into another In a tree T, we often compute dij(T) - the length of a path between leaves i and j Distance in Trees: an Exampe j i d1,4 = 12 + 13 + 14 + 17 + 12 = 68 Fitting Distance Matrix Given n species, we can compute the n x n distance matrix Dij Evolution of these genes is described by a tree that we don’t know. We need an algorithm to construct a tree that best fits the distance matrix Dij Summary Evolution and Phylogeny Concepts of Phylogenetics Application of Phylogenetics Category of phylogenetic inference algorithms Next lecture: Detailed algorithms for phylogenetic inference Acknowledgement Anonymous authors