* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Essential Statistics in Biology: Getting the Numbers Right
Survey
Document related concepts
Transcript
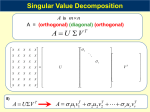
Essential Statistics in Biology: Getting the Numbers Right Raphael Gottardo Clinical Research Institute of Montreal (IRCM) [email protected] http://www.rglab.org Outline • Exploratory Data Analysis • 1-2 sample t-tests, multiple testing • Clustering • SVD/PCA • Frequentists vs. Bayesians Day 1 2 PCA and SVD (Multivariate analysis) Outline • What is SVD? Mathematical definition • Relation to Principal Component Analysis (PCA) • Applications of PCA and SVD • Illustration with gene expression data Day 1 - Section 4 4 SVD Let X be a matrix of size mxn (m≥n) and rank r≤n then we can decompose X as n m X n =m n n x S x V U n T n - U is the matrix of left singular vectors - V is the matrix of right singular vectors - S is a diagonal matrix who’s diagonal are the singular values Day 1 - Section 4 5 SVD Let X be a matrix of size mxn (m≥n) and rank r≤n then we can decompose X as n m Day 1 - Section 4 X n =m n n x S x V U n T n 6 SVD Let X be a matrix of size mxn (m≥n) and rank r≤n then we can decompose X as n m X n =m n n x S x V U n T n Direction Amplitude Day 1 - Section 4 7 Relation to PCA New variables Variance Assume that the rows of X are centered then is (up to a constant) the empirical covariance matrix and SVD is equivalent to PCA The rows of V are the singular vectors or principal components Gene expression: Eigengenes or eigenassays 8 Day 1 - Section 4 Applications of SVD and PCA • Dimension reduction (simplify a dataset) • Clustering • Discriminant analysis • Exploratory data analysis tool • Find the most important signal in data • 2D projections Day 1 - Section 4 9 Toy example set.seed(100) x1<-rnorm(100,0,1) y1<-rnorm(100,1,1) s=(13.47,1.45) var0.5<-matrix(c(1,-.5,-.5,.1),2,2) data1<-t(var0.5%*%t(cbind(x1,y1))) set.seed(100) x2<-rnorm(100,2,1) y2<-rnorm(100,2,1) var0.5<-matrix(c(1,.5,.5,1),2,2) data2<-t(var0.5%*%t(cbind(x2,y2))) data<-rbind(data1,data2) svd1<-svd(data1) plot(data1,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd1$v[2,1]/svd1$v[1,1]),col=2) abline(coef=c(0,svd1$v[2,2]/svd1$v[1,2]),col=3) Day 1 - Section 4 10 Toy example svd2<-svd(data2) plot(data2,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd2$v[2,1]/svd2$v[1,1]),col=2) abline(coef=c(0,svd2$v[2,2]/svd2$v[1,2]),col=3) s=(47.79,13.25) svd<-svd(data) plot(data,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd$v[2,1]/svd$v[1,1]),col=2) abline(coef=c(0,svd$v[2,2]/svd$v[1,2]),col=3) Day 1 - Section 4 11 Toy example ### Projection data.proj<-svd$u%*%diag(svd$d) svd.proj<-svd(data.proj) plot(data.proj,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd.proj$v[2,1]/svd.proj$v[1,1]),col=2) ### svd.proj$v[1,2]=0 abline(v=0,col=3) Day 1 - Section 4 12 Toy example s=(47.17,11.88) New coordinate s Day 1 - Section 4 Projected data 13 Toy example ### New data set.seed(100) x1<-rnorm(100,-1,1) y1<-rnorm(100,1,1) var0.5<-matrix(c(1,-.5,-.5,1),2,2) data1<-t(var0.5%*%t(cbind(x1,y1))) set.seed(100) x2<-rnorm(100,1,1) y2<-rnorm(100,1,1) var0.5<-matrix(c(1,.5,.5,1),2,2) data2<-t(var0.5%*%t(cbind(x2,y2))) data<-rbind(data1,data2) svd1<-svd(data1) plot(data1,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd1$v[2,1]/svd1$v[1,1]),col=2) abline(coef=c(0,svd1$v[2,2]/svd1$v[1,2]),col=3) svd2<-svd(data2) plot(data2,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd2$v[2,1]/svd2$v[1,1]),col=2) abline(coef=c(0,svd2$v[2,2]/svd2$v[1,2]),col=3) svd<-svd(data) plot(data,xlab="x",ylab="y",xlim=c(-6,6),ylim=c(6,6)) abline(coef=c(0,svd$v[2,1]/svd$v[1,1]),col=2) abline(coef=c(0,svd$v[2,2]/svd$v[1,2]),col=3) Day 1 - Section 4 14 Toy example s=(26.48,24.98) Day 1 - Section 4 15 Application to microarrays • Dimension reduction (simplify a dataset) • Clustering (two many samples) • Discriminant analysis (find a group of genes) • Exploratory data analysis tool • Find the most important signal in data • 2D projections (clusters?) Day 1 - Section 4 16 Application to microarrays Cho cell cycle data set384 genes We have standardized the data cho.data<-as.matrix(read.table("logcho_237_4class.txt",skip=1)[,3:19]) cho.mean<-apply(cho.data,1,"mean")cho.sd<-apply(cho.data,1,"sd")cho.data.std<-(cho.data-cho.mean)/cho.sd svd.cho<-svd(cho.data.std) ### Contribution of each PC barplot(svd.cho$d/sum(svd.cho$d),col=heat.colors(17)) ### First three singular vectors (PCA) plot(svd.cho$v[,1],xlab="time",ylab="Expression profile",type="b") plot(svd.cho$v[,2],xlab="time",ylab="Expression profile",type="b") plot(svd.cho$v[,3],xlab="time",ylab="Expression profile",type="b") ### Projection plot(svd.cho$u[,1]*svd.cho$d[1],svd.cho$u[,2]*svd.cho$d[2],xlab="PCA 1 ",ylab="PCA 2") plot(svd.cho$u[,1]*svd.cho$d[1],svd.cho$u[,3]*svd.cho$d[3],xlab="PCA 1 ",ylab="PCA 3") plot(svd.cho$u[,2]*svd.cho$d[2],svd.cho$u[,3]*svd.cho$d[3],xlab="PCA 2 ",ylab="PCA 3") ### Select a cluster ind<-(svd.cho$u[,2]*svd.cho$d[2])^2+(svd.cho$u[,3]*svd.cho$d[3])^2>5 & svd.cho$u[,2]*svd.cho$d[2]>0 & svd.cho$u[,3]*svd.cho$d[3]<0 plot(svd.cho$u[,2]*svd.cho$d[2],svd.cho$u[,3]*svd.cho$d[3],xlab="PCA 2 ",ylab="PCA 3") points(svd.cho$u[ind,2]*svd.cho$d[2],svd.cho$u[ind,3]*svd.cho$d[3],col=2) matplot(t(cho.data.std[ind,]),xlab="time",ylab="Expression profiles",type="l") Day 1 - Section 4 17 Application to microarrays Main contribution Relative contribution Why? Singular values Day 1 - Section 4 18 Application to microarrays PC1 Day 1 - Section 4 19 Application to microarrays PC2 Day 1 - Section 4 20 Application to microarrays PC3 Day 1 - Section 4 21 Application to microarrays Projection onto PC1 PC2 Day 1 - Section 4 22 Application to microarrays Projection onto PC1 PC3 Day 1 - Section 4 23 Application to microarrays Projection onto PC2 PC3 Day 1 - Section 4 24 Application to microarrays Projection onto PC2 PC3 24 genes Day 1 - Section 4 25 Application to microarrays Projection onto PC2 PC3 24 genes Day 1 - Section 4 26 Conclusion • SVD is a powerful tool • Can be very useful in gene expression data • SVD of genes (eigen-genes) • SVD of samples (eigen-assays) • Mostly an EDA tool Day 1 - Section 4 27 Overview of Statistics inference: Bayes vs. Frequentists (If time permits) Introduction • Parametric statistical model • Observation are drawn from a probability distribution where is the parameter vector Likelihood function → (Inverted density) Day 1 - Section 5 29 Introduction • Parametric statistical model • Observation are drawn from a probability distribution where is the parameter vector Likelihood function → (Inverted density) Day 1 - Section 5 30 Introduction Normal distribution Probability distribution for one observation is If independence Day 1 - Section 5 31 Introduction 15 observations N(1,1) Day 1 - Section 5 32 Introduction 15 observations N(1,1) True probability distribution Day 1 - Section 5 33 Inference • The parameters are unknown • “Learn” something about the parameter vector θ from the data • Make inference about θ Day 1 - Section 5 ‣ Estimate θ ‣ Confidence region ‣ Test an hypothesis (θ=0) 34 The frequentist approach • The parameters are fixed but unknown • Inference is based on the relative frequency of occurrence when repeating the experiment • For example, one can look at the variance of an estimator to evaluate its efficiency Day 1 - Section 5 35 The Normal Example: Estimation Normal distribution is the mean and is the variance (Sample mean and sample variance) Numerical example, 15 obs. from N(1,1) Use the theory of repeated samples to evaluate the estimators. Day 1 - Section 5 36 The Normal Example: Estimation In our toy example, the data are normal, and we can derive the sampling distribution of the estimators. For example we know that is normal with mean and variance . The standard deviation of an estimator is called the standard error. What if we can’t derive the sampling distribution? Use the bootstrap! Day 1 - Section 5 37 The Bootstrap - Basic idea is to resample the data we have observed and compute a new value of the statistic/estimator for each resampled data set. - Then one can assess the estimator by looking at the empirical distribution across the resampled data sets. set.seed(100) x<-rnorm(15) mu.hat<-mean(x) sigma.hat<-sd(x) B<-100 mu.hatNew<-rep(0,B) for(i in 1:B) { x.new<-sample(x,replace=TRUE) mu.hatNew[i]<-mean(x.new) } se<-sd(mu.hatNew) set.seed(100) x<-rnorm(15) mu.hat<-mean(x) sigma.hat<-sd(x) B<-100 mu.hatNew<-rep(0,B) for(i in 1:B) { x.new<-sample(x,replace=TRUE) mu.hatNew[i]<-median(x.new) } se<-sd(mu.hatNew) Day 1 - Section 5 38 The Normal Example: CI Confidence interval for the mean : where and usually depends on n but when n is large Numerical example, 15 obs. from N(1,1) set.seed(100) x<-rnorm(15) t.test(x,mean=0) > set.seed(100) > x<-rnorm(15) > t.test(x,mean=0) What does this mean? One Sample t-test data: x t = 0.3487, df = 14, p-value = 0.7325 alternative hypothesis: true mean is not equal to 0 95 percent confidence interval: -0.2294725 0.3185625 sample estimates: mean of x 0.044545 Day 1 - Section 5 39 The Normal Example:Testing Test an hypothesis about the mean: t-test If , t follows a t-distribution with n-1 degrees of freedom p-value Day 1 - Section 5 40 The Bayesian Approach • Parametric statistical model • Observation are drawn from a probability distribution where is the parameter vector ● The parameters are unknown but random ● The uncertainty on the vector parameter is model through a prior distribution Day 1 - Section 5 41 The Bayesian Approach A Bayesian statistical model is made of 1. A parametric statistical model 2. A prior distribution Q: How can we combine the two? A: Bayes Theorem! Day 1 - Section 5 42 The Bayesian Approach Bayes theorem ↔ Inversion of probability If A and E are events such that P(E)≠0 and P(A)≠0 then P(A|E) and P(E|A) are related by Day 1 - Section 5 43 The Bayesian Approach From prior to posterior: Prior information Information on θ contained in the observation y Normalizing constant Day 1 - Section 5 44 The Bayesian Approach Sequential nature of Bayes’ theorem: The posterior is the new prior! Day 1 - Section 5 45 The Bayesian Approach Justifications: Actualization of the information about θ by extracting the information about θ from the data • • Condition upon the observations (Likelihood principle) •Avoids averaging over the unobserved values of y •Provide a complete unified inferential scope Day 1 - Section 5 46 The Bayesian Approach Practical aspect: • Calculation of the normalizing constant can be difficult • Conjugate priors (exact calculation is possible) • Markov chain Monte Carlo Day 1 - Section 5 47 The Bayesian Approach Conjugate priors: Example:Normal mean, one observation → + and Day 1 - Section 5 48 The Bayesian Approach Conjugate priors: Example:Normal mean, n observations → + and Day 1 - Section 5 Shrinkage 49 Introduction Standardized likelihood Day 1 - Section 5 15 observations N(1,1) 50 Introduction Standardized likelihood 15 observations N(1,1) Prior Day 1 - Section 5 51 Introduction Standardized likelihood 15 observations N(1,1) Posterior Prior Day 1 - Section 5 52 Introduction Standardized likelihood 15 observations N(1,1) Prior Day 1 - Section 5 53 Introduction Standardized likelihood 15 observations N(1,1) Posterior Prior Day 1 - Section 5 54 The Bayesian Approach Criticism of the Bayesian choice: • Many! • Subjectivity of the prior (most critical) • The prior distribution is the key to Bayesian inference Day 1 - Section 5 55 The Bayesian Approach Response: Prior information is (almost) always available • • There is no such things as a prior distribution • The prior is a tool summarizing available information as well as uncertainty related with this information • Day 1 - Section 5 The use of your prior is ok as long as you can justify it 56 The Bayesian Approach Bayesian statistics and Bioinformatics • Make the best of available prior information • Unified framework • The prior information can be used to regularize noisy estimates (few replicates) • Computationally demanding? Day 1 - Section 5 57