* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download CBS2005_AMJ - Center for Biological Sequence Analysis
Survey
Document related concepts
Transcript
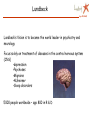
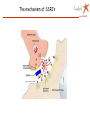
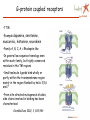
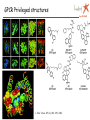
Apr 05/AMJ Computational decision support for drug design Profiling of small molecule compound libraries Anne Marie Munk Jørgensen Lundbeck Apr 05/AMJ Lundbeck’s Vision is to become the world leader in psychiatry and neurology Focus solely on treatment of diseases in the central nervous system (CNS) •depression •Psychoses •Migraine •Alzheimer •Sleep disorders 5000 people worldwide – app 800 in R & D Outline Apr 05/AMJ o What is a small molecule drug? o How can computational methods help during the drug discovery phase? • Library profiling: overall characterisation of a large pool of structures. • Prediction of more specific characteristics like biological activity and ADME properties • Privileged structures…. A small molecule drug Apr 05/AMJ … is a compound (ligand) which binds to a protein, often a receptor and in this way either initiates a process (agonists) or inhibits the natural signal transmitters in binding (antagonists) The structure/conformation of the ligand is complementary to the space defined by the proteins active site The binding is caused by favourable interactions between the ligand and the side chains of the amino acids in the active site. (Electrostatic interactions, hydrogen bonds, hydrophobic contacts…) The ligand binds in a low energy conformation < 3 kcal/mol Binding site complementarity Apr 05/AMJ HIV-Portease inhibitor JACS,V.16,pp847 (1994) H-bond donating H-bond accepting Hydrophobic Flo98, Colin McMartin. J.Comp-Aided Mol. Design, V.11, pp 333-44 (1997) Example of ligand binding Apr 05/AMJ 1UVT, Trombin Inhibitor No vacancy! Apr 05/AMJ Molecular factors Apr 05/AMJ Conformatio n Intramolecular interactions Ionization Intermolecula r forces Electronic distribution Solubility, Partitioning Carrupt P-A., Testa B., Gailard P. Boyd D.B., Lipkowitz K.B., Reviews in Computational Chemistry, Vol. 11, 1997, pp. 241-304. Compound library profiling Apr 05/AMJ Analyze a pool of structures to find out how attractive they are to us….. • 10 years ago: Diversity + HTS • Now: very high focus on how biologically relevant the screening collection is. • Computational methods to predict drug likeness, CNS likeness…. High throughput is not enough … to get high output….. Compound analysis Apr 05/AMJ Ideal 50.000 Structures: Chemical intuition Choosing the right descriptors is difficult Wolfgang Sauer, SMI 2004 Apr 05/AMJ How we describe the structures in the computer Apr 05/AMJ o Calculate a number of phys chem descriptors, like molecular weight, nhba, nhbd, logP, SASA….. o Describe the structures by keys…. Lipinski statistics Apr 05/AMJ Drug Like 1 CNS Like, present work, 90% limit. MW < 500 149.4 – 446.6 # hydrogen acceptors < 10 1-5 # hydrogen donors <5 0-3 logP <5 -0.3 – 4.9 # rotatable bonds NR 0 – 8.4 Rule of 5 References (1) Lipinski, C. A.; Lombardo, F.; Dominy, B. W.; Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev JID - 8710523 1997, 23, 3-25. Diversity and "Chemical Space" Apr 05/AMJ PCA Chemical space navigator Apr 05/AMJ Global Positioning System (GPS) Chem GPS (Oprea & Gottfries, J. Comb. Chem 2001) We want to define the CNS ”world” – the space which is biologically relevant when considering CNS drugs CNS model Apr 05/AMJ PCA 12 descriptors 3 components, R2X=0.71 Blue dots define:: CNS drug space CNS ”World” CNS ”world” sub classes Apr 05/AMJ O O O O N O Chiral O O Br N N N H N O O N N O O O H N O Model used to predict CNS-likeness Apr 05/AMJ N O N O F O I O O N O O N I I N O O O O Chiral N H N O N N N N N O H O O O N N N N H H O N N S N O O O O O O O O S N O O O N O O O N Structural clustering based on keys Apr 05/AMJ C-N 1 0.349 3 6 13 19 26 31 38 1 O N …01000100110001…. C=O O N Cl Cl C=C O N O N Cl Similarity by Tanimoto: O N Cl Tc= Bc/(B1 + B2 – Bc) clust_benzo (order) O N O Structural analysis Apr 05/AMJ o Clustering o Virtual screening – looking for structural similar compounds in a large pool of structures….. Apr 05/AMJ I have talked about overall profiling of a large number of compounds…… in terms of CNS-likeness … now I will turn to talk about prediction of more specific characteristics like biological activity and ADME properties….. Quantitative Structure Activity Relationship or Quantitative Structure Property Relationship In house QSAR study Apr 05/AMJ 2,5 SigmaP/pi 2 O 1,5 1 pi 0,5 N N S 0 -0,5 O sigmaP 0 1000 2000 3000 4000 IC50 Correlation between Glyt-1 inhibitor activity and pi (lipophilicity) and SigmaP (electronic characteristics) for the R substituent R ADME property predictions Apr 05/AMJ Oral absorption …depends heavily on permeability and Solubility… high interest in predicting these things in silico… Other things: Blood-brain Barrier penetration, clearance, Metabolism, tox….. Aqueous Solubility Apr 05/AMJ QSRP model n=775,R2=0.84, Q2=0.83 8 2D descriptors, Cerius2 Most important descriptors: logP, hba*hbd, hba, hbd Drugs: –6 < logS < 0; If error of 1 log unit is OK model predicts 60-80% of the compounds correctly Journal of Medicinal Chemistry, 2003, Vol. 46, No. 17 Permeability Apr 05/AMJ QSRP N= 13 R2=0.93 Q2= 0.83 Key descriptors: PSA> Odbl >N-H > ..NPSA >SA Polar descriptors important and …. size matters…. Simple Rule: PSA < 120 Å2 Journal of Medicinal Chemistry, 2003, Vol. 46, No. 4 Pharmacophore modelling Apr 05/AMJ ….. Another method of biological activity prediction… Observations that modification of some parts of a ligand results in minor changes of activity, whereas modifications of other parts of the ligand result in large change of activity. Pharmacophore element: Atom or functional group essential for biological activity 3D Pharmacophore mode: Collection of pharmacophore elements including their relative position in space From TCAs to SSRIs and Beyond Selective Serotonin Reuptake Inhibitors (SSRIs) CH3 N Apr 05/AMJ NHCH3 CN CH3 O N N CH3 NHCH3 CH3 O F3C F Br Cl zimelidine citalopram cipramil/celexa fluoxetine prozac/fontex 28.04.1971 14.1.1976 10.1.1974 First synt. Aug 1972 First synt. May 1972 Cl sertraline zoloft 1.11.1979 F3C F NH NH N O NH2 O N H indalpine 12.12.1975 O paroxetine paxil/seroxat 30.1.1973 O O fluvoxamine fevarin 20.3.1975 The mechanism of SSRI’s Apr 05/AMJ Pharmacophore modelling example Apr 05/AMJ Fluoxetine Paroxetine Citalopram Sertraline Chapter 13. Pharmacophore Modeling by Automated Methods: Possibilities and Limitations M.Langgård, B.Bjørnholm, K.Gundertofte In "Pharmacophore Perception, Development, and use in Drug Design". Edited by Osman F. Güne International University Line (2000) Privileged structures Apr 05/AMJ ……. are ligand substructures that can provide high-affinity ligands for more than one target….. Privileged structures Apr 05/AMJ ”A single ring system, the 5-phenyl-1,4-benzodiazepine ring, provides ligands for a surprisingly diverse collection of receptors…..” Evans et al., J. Med.Chem 1988, 31, 2235-2246 G-protein coupled receptors Apr 05/AMJ •7 TM •Example:dopamine, serotonine, muscarinic, histamine, neurokinin •Family A, B, C, A = Rhodopsin like •In general low sequence homology even within each family, but highly conserved residues in the TM regions •Small molecule ligands bind wholly or partly within the transmembrane region mainly in the region flanked by helix 3,5,6 and 7 •From site-directed mutagenesis studies, side chains involved in binding has been characterised ChemBioChem 2002, 3, 928-944 GPCR Privileged structures type of receptor Apr 05/AMJ J. Med. Chem., 47 (4), 888 -899, 2004 Amino acid ”hot spots” Apr 05/AMJ Priviledged sub structure for target T1 and T2 Align T1 and T2 Look for these in other GPCR’s Examine which amino acids are conserved in binding pocket for T1 and T2 Amino acid ”HOT SPOTS” Linking target and ligand side….. Didier Rognan at the 5ht international workshop in New Approaches In drug design & discovery, Marburg 21-24 marts 2005 Fluoxetine scaffold common for SERT and GLYT-1 N O Apr 05/AMJ COOH Gibson et al, Biorg. Med. Chem Letters 2001 (11), 2007-2009 CF3 N O F COOH Atkinson et al, Mol. Pharm. 2001 (60), 1414-1420 Comparison between SERT and GLYT-1 Apr 05/AMJ Y102 F288 Y310 GLYT1 sequence; RED: conserved residues GREY: conservative mutations SERT model From Na+/H+ antiporter, J. Pharmacol & Exp Therapeutics, 307, 34-41 Resume Apr 05/AMJ Computational methods for o Compound library profiling, Chem GPS o activity QSAR prediction and pharmacophore modelling o Solubility and permeability QSPR prediction o Privileged structures of GPCR’s ”Hit finding” Apr 05/AMJ Drug discovery ~ Looking for a needle in a haystack Filtering of compounds ~ remove some of the hay Serendipity Apr 05/AMJ “To look for the needle in the haystack and coming out with the farmer’s daughter” Arvid Carlsson