* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download dolphin1
Maximum parsimony (phylogenetics) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Point mutation wikipedia , lookup
Quantitative comparative linguistics wikipedia , lookup
DNA barcoding wikipedia , lookup
Koinophilia wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
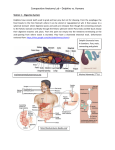
Dolphin Phylogeny Introduction Our research question was to learn how river dolphins are related to ocean dolphins and other cetaceans. Our hypothesis was that biogeography would show a higher correlation to common ancestry than would habitat. A secondary question was to use two different gene trees for the same species to see if similar phylogenies would be generated by each. April Maskiewicz, Dawne Page & Kerry Fulcher Hypothesis: Individual species of river dolphins will be more closely related to geographic neighbors than to each other. Fig. 1 16S rRNA unrooted tree Methods: In order to test our hypothesis, we chose to compare cetacean species with particular biogeographic ranges. Due to the limited amount of sequence available from river dolphins, we could only use sequences from mitochondrial DNA. We chose nucleic acid sequence from the 16S rRNA gene and amino acid sequence from cytochrome b for the species listed in Table 1.. We chose hippo and horse sequences as potential outgroups. Sequences were collected and edited for common length on the Biology Workbench 3.2 via internal (Workbench) or external (NCBI) database searches. Alignments were made and unrooted sequence similarity trees generated using Clustal W and DrawTree respectively. Finally, an evolutionary distance matrix was generated for the Cytochrome B data Common Dolphin using PROTDIST (Fig. 3) Amazon RD Indus RD Fig. 3 Evolutionary Distance Matrix form the multiple sequence comparison of Cytochrom B protein sequences. Data were artificially grouped into Low (green), Medium (Yellow) and High (Red) percent divergence to highlight relationships. Results: Two very different trees were generated. The rRNA gene tree (Fig. 1) showed 3 clusters: horse, Amazon river dolphin with all of the whales, and Asian river dolphins with the ocean dolphins and hippo. These data imply separate origins for the Amazon vs. Asian river dolphins. However, the cytochrome b tree (Fig. 2) did not imply such a simple conclusion. The Beluga and Humpback whales were closely related to the ocean dolphins (93-94%). The Humpback whale was also more closely related to the Indian river dolphins (92%) than to the Amazon & Yangtze river dolphins (88%). In contrast, the Humpback whale was more closely related to the Amazon river dolphin in the 16S rRNA phylogeny. Conclusion: Our tentative conclusion is that the river dolphins in India, the Amazon, and the Yangtze have separate origins, but these origins are unclear from the data, which illustrate the inherent difficulties of constructing phylogenies from several gene trees. Fig. 2 Cyt B unrooted tree Table 1. Species comparison using Cyt B protein sequences and 16S rRNA sequences