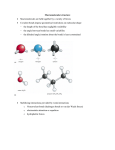
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download One 2-fold axis of symmetry
Western blot wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Structural alignment wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Homology modeling wikipedia , lookup
Proteolysis wikipedia , lookup
Biomolecular Graphics T Using the computer for displaying proteins and ligands to study their structure and function he application of X-ray crystallography to determining protein structures has caused a revolution in understanding protein function. As more structures have been solved, it has been possible to identify common themes of structural organization, and address how they fold. Initially laborious model building was required to transform the data specifying the position in space of different atoms (x, y and z co-ordinates) into a visual form. This was eliminated by the development of computer programs allowing the user to obtain any desired view of the protein on screen and manipulate it interactively in three-dimensional space. In this class you use PCs in conjunction with the freeware program RasMol to examine the structure of proteins, the co-ordinates of which have been downloaded from a Brookhaven Protein Data Bank web site. You use computer graphics to: • • • • • • Identify basic secondary structure elements in portions of a polypeptide chain. Identify loops and turns linking different secondary structure elements. Identify supersecondary structures. Measure inter-atomic distances to determine enzyme–substrate interactions. Determine contact residues in the interface between protein subunits. Examine residues involved in protein–nucleic acid interaction. Biomolecular Graphics 1 LEARNING RASMOL The use you have already made of RasMol in the Internet Data Retrieval unit only involved the graphic interface, whereas much of the power of RasMol resides in typing commands on the command-line in the text window. In addition, it takes a while to get used to the mouse-key combinations needed to manipulate molecules on the screen. A summary of the main features of RasMol is provided in the Appendix at the end of this manual, but this is more meaningful in the context of the following guided tour. Complete it before proceeding to the exercises. Starting Up Download the PDB file ‘3cln.pdb’, which contains the coordinates for calmodulin. You see two windows — the graphic window (Main Window) you used previously, and a ‘Command Line’ window. Adjust the relative positions of the windows so that the graphic window is in the foreground, but the current text in the command-line window is visible. (You can use the keyboard to type instructions onto the command line, even though it is the background window.) Calmodulin is a convenient protein to view while covering some main aspects of RasMol. Further instructions are in the Appendix, which you should read. 2 Biochemistry-3H and Medical Biochemistry-3H y x z Rotation To rotate about y axis, drag with the LH mouse button horizontally. To rotate about x axis, drag with the LH mouse vertically. To rotate about the z axis, press the shift key and drag the LH mouse button horizontally. Zooming Zooming in allows an image to be viewed close-up. To zoom out, hold down the Shift key and drag up vertically with the mouse (LH). To zoom in, hold down the Shift key and drag down vertically with the mouse (LH). Translation Having zoomed in you can move the picture to any part of the graphics window. This is called translation. To translate, drag the RH mouse button in the desired direction. Identifying Atoms Zoom out and find how to identify atoms. To do this: Click the cursor (LH) on the atom, and read the information in the command-line window. This is the -sulphur atom of a methionine residue (Met-109). Consult the appendix on amino acid structures for information on atom nomenclature in amino acids. It is often useful to identify the residues by number, especially when in close-up. This can done with the command ‘label on’. However, by default, this causes all atoms to be identified. To restrict labelling to residue numbers: Select Label in the Options menu. Before proceeding to the next section turn the labelling off by deselecting Label. Biomolecular Graphics 3 Selecting and restricting It is possible to select individual parts of a molecule (polypeptide chain, backbone, ligand, metal ions) and then display or operate on them in a different way from the rest of the molecule. In that case the rest of the molecule is still visible. Alternatively, it is possible to restrict display to individual parts of the molecule. Proceed directly from the previous view: Type ‘select Ca’. Select Spacefill from the Display menu. Two of the calcium ions should now be visible. Type ‘restrict Ca’. Now only the calcium ions are visible. As selecting and restricting is so central to working with RasMol we continue with another example starting from the structure reset to its original position. Select Spacefill from the Display menu. (Type Select all first, if continuing directly.) Because, by default, everything is selected, a view including some red atoms floating in space is seen. These are the oxygen atoms of water molecules in the crystal. To suppress these : Type ‘restrict !water’. The ! has a negating effect and the water molecular are no longer visible. N.B. You can make everything invisible with the command ‘restrict 0’. This allows you to start from scratch with ‘restrict all’ or ‘restrict protein’ etc. You must then make a selection from the Display menu before you see anything. Slabbing Slabbing is a means of causing the front of the molecule to be deleted so that you can view a transverse section of the back half. To illustrate it, start from the spacefilling view above: Select Slab Mode from the Options menu. You may see that the nearer portion of the molecule has been deleted. You can control the amount deleted by slabbing: Hold down the Control key and drag down vertically with the mouse. Repeating Previous Commands: Use the up arrow on the keyboard. This can save retyping. 4 Biochemistry-3H and Medical Biochemistry-3H SECONDARY STRUCTURE OF PROTEINS Introduction Dihedral Angles As the diagram below, left, illustrates, the peptide bond is planar, so there is no rotation about the C–N bond, whereas rotation is possible about the other two bonds in the protein backbone: the N–C and C–C bonds. The angles of rotation about these bonds are referred to as dihedral or torsion angles, and are designated by the Greek letters (phi) and (psi), respectively. 180Þ C 90Þ L 0Þ R –90Þ Because of steric hindrance, only certain pairs of dihedral angles for one amino acid are thermodynamically favoured, and these are indicated on the Ramachandran plot as dark grey areas (for L-Ala) or light grey areas (for Gly). In secondary structures, stable dihedral angles that allow strong hydrogen bonds between peptide bonds repeat in succession. The values of dihedral angles for various secondary structures are shown on the Ramachandran plot, although it must be emphasized it is only repeating values that indicate a particular secondary structure. The diagram opposite shows how dihedral angles are measured relative to the three planar atoms and the one rotating. RasMol can be used to measure dihedral angles by typing ‘set picking torsion’ and clicking on the relevant atoms. –180Þ –180Þ –90Þ 0Þ 90Þ 180Þ (AB to the front) Biomolecular Graphics 5 Aspects of -sheets Although the dihedral angles in -helices and -sheets are sufficiently different to allow easy identification, those of parallel and anti-parallel -sheets are not well differentiated. When examining the -sheet secondary structure elements of proteins, the different patterns of hydrogen bonding (illustrated below) are helpful for distinguishing purposes. Antiparallel -sheet Parallel -sheet Sometimes the regular nature of the anti-parallel -sheet is disturbed by the oxygen atom on one chain missing a corresponding NH on the opposite chain and making a hydrogen bond with the following one (indicated by a grey circle in the diagram below). Likewise, the adjacent NH misses a corresponding O. This feature, which you may not have encountered previously, is called a -bulge, and causes a bend in the sheet. -Hairpins and -Turns Where the strand of an anti-parallel -sheet turns back on itself, the resulting motif of two strands and a non-hydrogen-bonded looped-out region is called a -hairpin. The number of unpaired residues in such hairpins can vary. This type of structure should not be confused with the -turn, in which the CO group of residue n of a polypeptide is hydrogen-bonded to the NH of residue n+3. Despite its name, the -turn is not confined to -sheets. 6 Biochemistry-3H and Medical Biochemistry-3H Individual Secondary Structures Objectives The three structures in this section are small portions of proteins illustrating different types of secondary structure. The main objective is to identify which secondary structure each represents. Procedure Examine each of the structures, p110117, t7986 and pabparl, in the Sticks or Wireframe Display mode, consulting pp. 3–7 and the RasMol instructions in the Appendix, as necessary. On the diagrams, colour the oxygen atoms red and the nitrogen atoms blue, and mark or list, as appropriate: • The position of the N- and C-termini. • The amino-acid sequence (single-letter code). • The positions of the hydrogen bonds, with the relative residue numbers of the amino acid residues (e.g. n, n+3 would mean that the CO of residue n bonds to the NH of residue n+3). • The main-chain dihedral angles for each residue except the N- and C- terminal ones (p110117 and t7986 only). Relate each pair of values to a region of the Ramachandran plot. From this: • Deduce the secondary structure present, marking this on the diagram, with the position of the axis of any helix and the number of helical turns. Indicate any other hydrogen-bonded features such as -hairpins or -turns. t7986 (Thermolysin: 79–86) Biomolecular Graphics 7 p110117 (Prealbumin: 110–117) pabparl (Prealbumin: 13–17, 106–110) 8 Biochemistry-3H and Medical Biochemistry-3H ASPECTS OF PROTEIN STRUCTURE Introduction Marking General Features Each of the proteins to be examined in this section illustrates a different aspect of protein structure. However, before focusing on specific features it is useful to orient yourself in relation to the general features of the protein. Thus, in every case except ldm, mark the following features on the diagrams: • The position of the N- and C-termini. • The -helices. Label these A, B, C etc., in the order they occur along the chain. • The -sheets and strands within them. Use different colours to indicate different -sheets. Label strands 1, 2, 3 etc., in the order they occur along the chain. Use arrows to show the direction and whether strands are parallel or anti-parallel. Consult a demonstrator before you start to colour the diagrams. • The supersecondary structure and other features such as -hairpins. (Most of the proteins here can be categorized as all-, all- or /.) • The position of any active sites or ligand-binding sites, if known. (This may be more convenient after examining the specific features of the protein.) • A representative sample of hydrogen bonds (up to, say, 8 of them). Using RasMol with larger structures The Wireframe or Sticks views you used for small segments of polypeptide chain often gives too complex a picture with whole proteins. The following can make things clearer: • Change to Backbone in the Display menu. (This is actually an -carbon plot.) You can add hydrogen bonds and ligands to this. • Select Group from the Colour menu to see the directionality of the chain. • Select Structure from the Colour menu to display -helices in magenta and -sheets in yellow. • Most of the protein diagrams you have to annotate are smoothed -carbon plots to make them easier to follow. They differ somewhat from your computer images. Immunoglobulins Objective A main objective is to compare the corresponding portions of the heavy and light chains of immunoglobulins to see to what extent a similar tertiary structure requires a similar primary structure, and to identify the antigenbinding regions. Here the antigen is the protein lysozyme. Background The structure of immunoglobulins is familiar from second-year science. The diagram opposite reiterates this and, in addition, indicates two features not covered previously. One is the hypervariable segments of the variable regions, especially involved in binding antigen. The other is the so-called Fab fragment, which is generated proteolytically and is simpler to study than the whole antibody. (See Stryer (5th ed.) Ch. 33 for further details.) General Features Start by examining the file hfll, which consists of a single light chain domain: the variable region. Identify the hydrogen bonds and -strands (as instructed on p. 10), locate the disulphide bonds and hypervariable regions and mark them on the diagram. Biomolecular Graphics 9 To orient yourself in relation to the main chain: Select Backbone from the Display menu. This displays only the -carbon backbone. Select Group from the Colour menu. This colours rainbow-like from blue to red, corresponding to N- to C-terminus. • Follow the backbone on screen, and mark the directionality with arrows on the diagram. To display the -sheet hydrogen bonds: Type ‘restrict sheet’. This restricts the display to the -sheet. Type ‘set hbonds backbone’. This makes the hydrogen bonds connect to the backbone -carbons we are displaying. Type ‘hbonds on’. The hydrogen bonds are displayed. Type ‘select all’. Select Backbone from the Display menu. The rest of the backbone should now be restored. Mark the features requested on p. 10. To highlight the disulphide bonds: To display the hypervariable regions: Type ‘select cys’. Type ‘select 27-32L,49-52L,90-96L’. Type ‘colour red’. Type ‘colour yellow’. Select Spacefill from the Display menu. hfll This section compares hfll, the light-chain variable domain you have just examined, with hflh, the corresponding heavy-chain variable domain. 10 Biochemistry-3H and Medical Biochemistry-3H The diagram provides an alignment of the primary structures of these two domains (H=heavy, L=light), generated by the sort of computer program used in the DNA Cloning laboratory class. Identities are indicated by vertical bars, and residues of greater and lesser chemical similarity are indicated by ‘:’ and ‘.’, respectively. H L H L H L . 1 EVQLQQSGAEL.MKPGASVKISCKASGYTFSDYWIEWVKQRPGHGLEWIG 49 ::.|.||.| : .||..|.:.|.||: . ||.: .. ... : : . 1 DIVLTQSPAIMSASPGEKVTMTCSASSSVNYMYWYQQKSGTSPKRWIYDT 50 . . . . . 50 EILPGSGSTNYHERFKGKATFTADTSSSTAY.MQLNSLTSEDSGVYYCLH 98 . |:::....: .:.:|:|.| : :.|:..||.: ||| : 51 SKLASGVPVRF...........SGSGSGTSYSLTISSMETEDAAEYYCQQ 89 . 99 GNYDFDGWGQGTTLTVSS 116 . : ..:|.||.|.:.. 90 WGRN.PTFGGGTKLEIK 105 . . . . On the basis of this alignment of primary structures, complete the table below. No. of residues inchainH No. of residues inchainL No. of identities Identities as apercentage of residues inchainH Identities as apercentage of residues inchainL No. of unaligned residues inchainH No. of unaligned residues inchainL On the Primary structure alignment mark the position of the hypervariable regions. The Interaction of the Antigen with the Immunoglobulin Biomolecular Graphics 11 In this section you are to examine hfl, the complex of the Fab fragment with lysozyme, the antigen. Because there are three different polypeptide chains it is necessary to be able to distinguish them. This can be done (after selecting Backbone display) by • Select Chain from the Colour menu. This presents each chain in a different colour. Identify the chains by clicking on their atoms and reading the command line code: L = light chain, H = heavy chain, Y = lysozyme • Make a very simple diagram below of the immunoglobulin domains and the antigen. (Do not try to trace the -carbon backbone — just draw irregular spheroids.). • Label the different domains and subunits; point to the domain that is the same as hfll. • Type ‘select 27–32L, 49–52L, 90–96L, 26–31H, 53–56H, 98–105H’. • Indicate the position of the six hypervariable regions on the light (l) and heavy (h) chains. • Relate your diagram to that given in the introduction. (This is reproduced below, but without the labelling, which you should add yourself to both diagrams. Note also that the disulphide bonds are omitted as they are not present in hfl) 12 Biochemistry-3H and Medical Biochemistry-3H Haemoglobin Objective One of the main objectives here is to study the subunit contacts in the quaternary structure of haemoglobin. Background It is expected you are familiar with molecular physiology of haemoglobin. If, necessary consult Chapter 10 of Stryer (5th ed.). (You may be unfamiliar with either the second (distal) histidine residue (E7) in the haem pocket, or the 2,3,-diphosphoglyerate (BPG) molecule between the subunits; BPG is described in Stryer 5th Edn, Chapter 10) In studying subunit interaction in proteins with quaternary structure it is useful to consider symmetry. As you may not have encountered this topic previously, the following is an introduction. There are two types of symmetry, reflection and rotational symmetry. Only rotational symmetry occurs in proteins. Rotational symmetry of multimeric proteins occurs when an identical structure is obtained by rotating the molecule by less than 360˚ about an axis through its centre (the axis of symmetry). The angle through which the molecule needs to be rotated defines the type of symmetry: . If a molecule is rotated through 360˚/n to produce a structure that is indistinguishable from the original, it is said to have an n-fold axis of symmetry The simplest situation is the dimer, where rotation through 180˚ gives an indistinguishable structure. Thus, there is a 2-fold axis of symmetry, also called a dyad. In haemoglobin we are concerned with a tetramer. (Although there are two different subunits, and , we regard them as practically identical and qualify our statements by referring to pseudosymmetry.) One possible type of symmetry for a tetramer is a single 4-fold axis, but in haemoglobin the subunits are in a tetrahedron and have three mutually perpendicular 2-fold axes of symmetry (222 symmetry). These axes are not marked on the diagram below. Do this on your own diagrams after studying the structure. Dimer One 2-fold axis of symmetry Note that in this type of tetramer there are potentially three different types of subunit–subunit contact, each corresponding to one of the dyads. Tetramer (tetrahedral) Three 2-fold axes of symmetry Tetramer (planar) One 4-fold axis of symmetry General Features Biomolecular Graphics 13 Start by examining the file, hhba, which consists of a single -subunit of human deoxyhaemoglobin. Identify the hydrogen bonds and -helices (as instructed on p. 10), then focus on the haem group and its neighbouring histidine residues. Some hints: • Select Backbone from the Display menu. This simplifies the display to the -carbon backbone and allows you to identify the helices. As there is a general naming convention for globin helices, label the helices A, B, C, E, F, G and H (there is no Dhelix in human haemoglobin). Note the C-helix is very short here. When you are ready to look at the haem pocket: • Type ‘select hem’ to select the haem group. • Select Ball & Stick from the Display menu. This allows you to see the haem and its iron atom. • • • • Type ‘select his’ Type ‘centre hem’ and then ‘colour cpk”. Select Wireframe from the Display menu. Zoom in on the haem. • • • Mark the ‘proximal’ histidine (res.87) and the ‘distal’ histidine (res.58) on the diagram. Mark the position at which oxygen binds. Comment on the position of the iron atom with respect to the plane of the haem ring. hhba (-subunit of human deoxyhaemoglobin) 14 Biochemistry-3H and Medical Biochemistry-3H Symmetry Now examine the file hhb, the human deoxyhaemoglobin tetramer: • The Data Bank file designates the four globin subunits: 1 is a, 1 is b, 2 is c, and 2 is d. • Spacefill Display is useful here, as is Chain Colour. • Use the command ‘restrict !water’ to remove unwanted (red) water molecules. For this part draw your own diagrams: • Make a diagram showing the subunits as spheres and label them. • Draw the positions of the part of the haems that are visible. Type ‘set axes on’. • Label the axes (x, y, z) in your diagram. Indicate whether they are true or pseudo axes. • The molecule 2,3,-diphosphoglycerate normally binds in the space between the two -subunits. In hhb you see two phosphate ions at this position. Indicate their position (and hence that of the ligand) in your diagram. • Make a separate simple drawing with subunits as spheres to show the three possible types of subunit–subunit contact that occur in tetramers with 222 symmetry. Illustrate them by views of the tetramers in which one subunit is to the front and the other three lie to the back. Show where the axes of symmetry are. Biomolecular Graphics 15 Subunit–Subunit contacts Having obtained a general view of the three subunit–subunit contacts, we examine them to investigate their strengths, i.e. the number of atoms in contact. Select Ball & Stick from the Display menu. Select Chain from the Colours menu. Type ‘restrict within (4.0,*a)’. This restricts display to subunit a (1) and all atoms within 4Å of it. The value of 4Å is generally used as it encompasses hydrogen bonds and most Van der Waals interactions. Type ‘select *a’. This selects the subunit alone, which can now be distinguished by displaying in backbone mode: Select Backbone from the Display menu. Now only the atoms from the other subunits are displayed in Ball & Stick mode. Count the numbers of atoms in the three colours to identify the contacts of 1 with each of the other subunits. • Hence deduce the relative strength of each contact. • When haemoglobin changes conformation from the R-state to the T-state, the 1-1 dimer rotates with respect to the 2-2 dimer. Is this consistent with the strengths of the contacts you have determined? 16 Biochemistry-3H and Medical Biochemistry-3H Triose-Phosphate Isomerase Objective The objective here is to delineate an extensive and regular supersecondary structure: the -barrel . Procedure The file, tim, contains the co-ordinates of the triose-phosphate isomerase dimer. To restrict viewing to one chain: • Type ‘restrict *a’. Then, to ensure that rotation is about the centre of this subunit (rather than that of the dimer): • Type ‘centre *a’. • Annotate the diagram below according to the points on p. 10. • State the number of strand–loop–helix motifs that you observe. • Also, draw a sample of the main-chain/main-chain hydrogen bond arrangement of the -barrel. A second file that contains only the -barrel portion of the subunit, timbrl, is provided to facilitate this. Slabbing can help to show a suitable portion. • • • • . In addition, examine the middle of the barrel to see what type of residues it contains and how much space there is in it. Select Spacefill from the Display menu. Type ‘select hydrophobic’. Type ‘colour green’. Then ‘slab’ through the molecule. Describe briefly the occupancy of the middle of the barrel. tim (Triose-phosphate isomerase) Biomolecular Graphics 17 Lactate Dehydrogenase Objective The main objective here is to examine the active site of lactate dehydrogenase, and, from the proximity of amino acid residues and substrate, assess the proposed mechanism of action of the enzyme. Background Lactate dehydrogenase (LDH) is a tetramer that catalyses the reaction: O + H3 C C COOH + NADH + H Pyruvate It is not possible to crystallize the enzyme with its substrate without it reacting, so that oxamate — an analogue of pyruvate in which the methyl group is replaced by an amino group — is used and appears in the structures you examine OH + H3 C C COOH + NAD H Lactate O H2 N C COOH Oxamate A feature of LDH and many other nucleotide-binding proteins is a six-stranded domain called the Rossman fold, which you examine here. Despite this tertiary structure similarity, there is little primary structure homology in the Rossman fold, the only conserved feature being the motif, GxGxxG, as shown in the table below. Dihydrolipoate dehydrogenase (E coli) Alcohol dehydrogenase (Drosophila) Lactate dehydrogenase (Mouse) Glyceraldehyde-phosphate dehydrogenase (Pig) GGGILGL GLGGIGL GVGDVGM GFGRIGR You recall the role of a histidine residue in the proposed mechanism of action of LDH. You examine that histidine here, but, because the structure is for dogfish LDH, its numbering may differ from other LDHs. General Position of the Substrates Start by examining the file, ldm, which contains the co-ordinates of one subunit of dogfish LDH. This is too complex to indicate the features listed on p. 10, but is useful for observing some features which you should indicate on the diagram overleaf: There are two large domains and one smaller N-terminal domain (residues 1–18), which reaches across to the other subunits in the tetramer. The large N-terminal domain (19–161) can be seen by selection and differential colouring. • Mark the three domains. NADH and oxamate are in a groove between the two domains. Examine the extent to which the substrates are buried by selecting Spacefill display mode for all atoms and selecting and colouring the ligands differentially (select nad and select oxm are the commands). Then slab through the molecule, as described on p.5 and in the Appendix. • Mark the positions of the substrates, and indicate to what degree, and which parts of, the two substrates are buried. 18 Biochemistry-3H and Medical Biochemistry-3H ldm (Single subunit of LDH bound to NADH) ldmd1 (N-terminal domain —residues 19–161— of LDH subunit bound to NADH) Biomolecular Graphics 19 20 Biochemistry-3H and Medical Biochemistry-3H